Detection of
de Novo
Insertion of the Medaka Fish
Transposable Element
Tol2
Akihiko Koga and Hiroshi Hori
Division of Biological Sciences, Graduate School of Science, Nagoya University, Nagoya 464-8602, Japan
Manuscript received April 24, 2000 Accepted for publication July 10, 2000
ABSTRACT
Tol2is a terminal-inverted-repeat transposable element of the medaka fishOryzias latipes.It is a member of the hAT (hobo/Activator/Tam3) transposable element family that is distributed in a wide range of organisms. We here document direct evidence forde novoinsertion of this element. ATol2clone marked with the bacterialtetracycline-resistancegene was microinjected into fertilized eggs together with a target plasmid, and the plasmid was recovered from embryos. The screening of plasmid molecules after transfor-mation intoEscherichia colidemonstrated transposition oftetinto the plasmid and, by inference, precise insertion ofTol2in medaka fish cells.De novoexcision ofTol2has previously been demonstrated. The present study provides direct evidence that theTol2element has the entire activity necessary for cut-and-paste transposition. Some elements of themariner/Tc1 family, another widespread group, have already been applied to development of gene tagging systems in vertebrates. TheTol2element of thehATfamily, having different features from mariner/Tc1 family elements, also has potential as an alternative gene tagging tool in vertebrates.
S
EVERAL transposable elements of the terminal- sion has been demonstrated (Kawakami and Shima 1999). The same enzyme would also be expected to inverted-repeat class are known to exist inverte-brate genomes. However, only a few of them have been catalyze insertion, as is usually the case with terminal-demonstrated to be active. The first example for which inverted-repeat elements. The reasonde novoinsertion a transposition event was observed is the Tzf element has not been detected earlier is that it requires, in con-of zebrafish (Lam et al.1996). Other examples are the trast to the excision case, a suitable experimental system synthetic elementSleeping Beauty(Ivicset al.1997) and to allow detection of the inserted element at what would exogenous elements such asTc3ofCaenorhabditis elegans normally be random locations. One strategy to over-(Razet al.1997) andmarinerof Drosophila (Fadoolet come this difficulty is to trace a genetically marked ele-al.1998). All these elements are members of themari- ment. In this study, we markedTol2 with the bacterial ner/Tc1 transposable element family. There is one other tetracycline-resistance (TetR) gene and screened target
element for which part of the transposition reaction has DNA fragments in Escherichia coli transformants for been demonstrated. The element isTol2, found in the transposition products conferring tetracycline resis-medaka fish (Koga et al. 1996), andde novo excision tance. Precise insertion, withTol2flanked by 8-bp target has also been detected in zebrafish (Kawakami et al. site duplications, was detected. In addition, the transpo-1998). However,de novoinsertion has hitherto not been sition was proven to be catalyzed by the transposase that observed. This element is a member of thehoto/Activa- is encoded by a sequence withinTol2.
tor/Tam3(hAT) element family found in various organ-isms, even across kingdoms (Calviet al.1991;Atkinson
et al. 1993;Koga et al. 2000). Such widespread occur- MATERIALS AND METHODS rence is an important feature for developing a gene
Fish:Oryzias latipes(medaka fish, also called Japanese me-tagging system because it implies relative independence
daka fish) andO. melastigma(Indian medaka fish) were used. of transposition mechanisms from species-specific host O. latipeswas purchased from a pet shop in Nagoya and the factors and, thus, applicability to a relatively wide range fish containedⵑ20Tol2copies per diploid genome according
of organisms. to genomic Southern blots (KogaandHori1999).O.
melas-tigmawas obtained from the World Medaka Aquarium of the The purpose of this study was to detectde novo
inser-Nagoya City Higashiyama Zoological Garden. This species tion ofTol2.A putative transposase ofTol2has already
does not harborTol2in its genome (Kogaet al.2000). been suggested (Kogaet al.1999) and catalysis of exci- Construction of plasmids:Tol2was first identified as a
4.7-kb insertion sequence in thetyrosinasegene of an albino mu-tant fish (Koga et al. 1996). This particular Tol2 copy (Tol2-tyr, GenBank accession no. D84375) together with its Corresponding author:Hiroshi Hori, Division of Biological Sciences,
8-bp target site duplication sequences was amplified by PCR.
Graduate School of Science, Nagoya University, Nagoya 464-8602,
Japan. E-mail: hori@bio.nagoya-u.ac.jp The primers used were 5⬘-AAGGATCCTCAAGAACCAGAG
Figure1.—Organization of plasmids. pDon01 is a donor plasmid. It contains the chloramphenicol-resistance(CamR) gene and aTol2element carrying thetetracycline-resistance (TetR) gene. pTar01 is a target plasmid containing thekanamycin-resistance
(KanR) gene.
GTGTAAAGT-3⬘ and 5⬘-CCTCTAGAGTTCTTGACAGAGG chosen and checked for their sizes by agarose gel elec-TGTAAAAA-3⬘. The third to eighth nucleotides are restric- trophoresis (Figure 3). This test revealed the majority tion enzyme sites (BamHI andXbaI, respectively) for cloning,
of the plasmids to be of two sizes: 7.7 and 10 kb. Plasmids the next eight nucleotides are target site duplications (part
of 7.7 kb were observed only in experiments B and D, of thetyrosinasegene), and the remaining sequences to 3⬘are
the ends ofTol2(the left and right ends, respectively). The in which mRNA was microinjected together with the amplified fragment was cloned into pHSG399 (Takeshitaet plasmid DNAs. If theTol2portion (4274 bp) of pDon01 al. 1987) at its BamHI and XbaI sites in the lacZ gene. An (6517 bp) is integrated precisely into pTar01 (3446 bp), internal region of Tol2between its two PstI sites (positions
the expected size of a resultant plasmid is 7.7 kb. There-2401 and 4272 on the Tol2-tyr sequence) was removed and
fore, these plasmids are likely to be products of precise replaced with the TetRgene of pBR322 amplified with PCR
(nucleotides 4227–1375, including nucleotide 1, of accession Tol2 insertion. The fact that all these plasmids were no. J01749). This Tol2-carrying donor plasmid was denoted found to carry singleSacI cutting sites also supports this pDon01 (Figure 1). The target plasmid pTar01 (Figure 1) was inference because pDon01 and pTar01 both have single prepared by inserting a 0.8-kb DNA fragment (nucleotides
SacI sites in their vector portions (see Figure 1). 2679–747, including nucleotide 1, of accession no. M12787
for pHSG664;Hashimoto-Gotohet al.1986) into pHSG299 (Takeshitaet al.1987). The fragment contains the streptomy-cin-sensitive(StrS) gene whose disruption, in association with a specific bacterial strain, confers the streptomycin-resistance phenotype. It was prepared with the long-range plan of screen-ing for insertions in this gene. In the present study, the 0.8-kb fragment was employed simply to enlarge the target plasmid. Transposition assay:The scheme of the transposition assay is illustrated in Figure 2. The pDon01 and pTar01 plasmids, 150 ng/l each in 10 mmTris-HCl (pH 8.0), were microin-jected into fertilized eggs at the one-cell stage, with or without mRNA for the putativeTol2transposase (Kogaet al.1999), prepared by in vitrotranscription using the RiboMAX large scale RNA production system SP6 (Promega, Madison, WI). A mammalian-cap-like sequence was added to its 5⬘end and the mRNA was included in the DNA solution for injection at a final concentration of 150 ng/l. After incubation at 25⬚ for 24 hr, plasmid molecules were recovered from embryos by the method ofHirt(1967) and then introduced into the JM109 strain ofE. coliby electroporation. The bacteria were screened for plasmids showing both the KanRand TetR
pheno-types, which can be expected to include products of transposi-tion. The drug concentrations were 50g/ml for kanamycin and 6g/ml for tetracycline.
Sequencing analysis:Candidate transposition products were further selected by restriction mapping and sequenced for theirTol2terminal and flanking regions, as described (Koga
et al.1995). The sequencing primers were nucleotides 424–401 of theTol2-tyrsequence for the left terminus and nucleotides 4316–4339 for the right terminus.
Figure 2.—Transposition assay using plasmids. Plasmid RESULTS
molecules recovered from embryos were introduced into bac-teria by electroporation. Most of the bacbac-teria were plated on Transposition assay:We carried out five experiments
kanamycin- and tetracycline-containing media to screen for as listed in Table 1. In experiments A–D, which included
Tol2insertion into the pTar01 plasmid. Small aliquots were microinjection of plasmid DNAs, plasmids showing both plated on media containing only kanamycin to estimate the the KanRand TetRphenotypes were found. Twenty
plas-number of pTar01 molecules both with and withoutTol2 inser-tion.
Numbers of colonies and distribution of plasmid sizes
Druga Plasmid sizeb
Experiment Fish mRNA Kan⫹Tet Kan 7.7 kb 10 kb ⬎12 kb
A O. latipes No 156 3.9⫻105 0 19 1
B O. latipes Yes 136 2.7⫻105 8 11 1
C O. melastigma No 101 4.1⫻105 0 18 2
D O. melastigma Yes 90 3.0⫻105 9 10 1
E — No 0 9.3⫻105 — — —
aAfter electroporation, most of the bacterial mixture was plated on medium containing both kanamycin
and tetracycline, “Kan ⫹ Tet” indicating the number of colonies then observed. Small aliquots were also plated on medium containing only kanamycin. With the number of colonies on these media and the ratio of the mixture volumes, the total number of KanRbacteria included was estimated as given in “Kan.”
bTwenty plasmids each from experiments A–D were checked for their sizes.
If the pDon01 and pTar01 plasmids were combined and were flanked by sequences of pTar01 (Figure 4). into one molecule, the resultant plasmid would be ex- In addition, target site duplications of 8 bp were ob-pected to be 10 kb in size, like the plasmids of this size served in all cases (Figure 4). The insertion breakpoints observed in experiments A–D. They all demonstrated according to theTol2-flanking sequences are illustrated twoSacI sites (examples shown in Figure 3) consistent in Figure 5. There was no apparent region where inser-with the inferred link. The fact that restriction patterns tion breakpoints were concentrated.
forSacI differed among these 10-kb plasmids suggested that the integrations, on the assumption of this
infer-ence, were independent events. DISCUSSION
There were also a few plasmids sized ⬎12 kb with
Detection of de novoinsertions: Our results provide threeSacI sites, as shown in Figure 3.
evidence forde novoinsertion ofTol2.Because excision
Sequencing analysis:Of the 7.7-kb plasmids, five each
has already been detected, it can now be concluded from experiments B and D were randomly chosen and
thatTol2has all the activity necessary for cut-and-paste sequenced for theirTol2terminal and flanking regions.
transposition. With all plasmids and for bothTol2 termini, the Tol2
We have not conducted further analysis of the 10-sequences were observed as far as the last nucleotides
kb plasmids and the few larger plasmids because the purpose of our study was to detect insertion of Tol2, attained with the 7.7-kb plasmids. The 10-kb plasmids appear to have resulted from simple combination of pDon01 and pTar01 into one plasmid. If this is true, a possible mechanism for this event is homologous recom-bination, because the two vector plasmids (pHSG399 and pHSG299) share regions inherited from their com-mon origin. Irrespective of the mechanisms involved, the recombination events occurred not in bacteria but in medaka fish cells because KanR TetR plasmids were not observed in experiment E. Precise insertion reaction leading to 7.7-kb plasmids also occurred in medaka fish cells.
Evidence for transposase function:O. melastigmadoes not containTol2 in its genome. Therefore, the differ-ence in results between experiments C and D implies that the mRNA is essential for transposition ofTol2.It is thus evident that the mRNA encodes a transposase
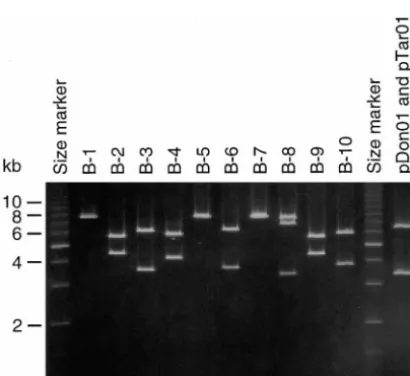
Figure3.—Electrophoresis of plasmid clones. Single
plas-mids isolated from experiments A–D were digested withSacI ofTol2that also functions as such inO. melastigma. and electrophoresed on 0.6% agarose gels. In the example O. latipescontainsⵑ20Tol2 copies and endogenous shown from experiment B, three clones (B-1, -5, and -7)
exhi-mRNA molecules are present in cells of this species bit single fragments of 7.7 kb, six clones (B-2, -3, -4, -6, -9,
(Kogaet al.1999). We infer from the results of experi-and -10) have a total length of 10 kb, experi-and one clone (B-8) is
Figure 4.—DNA sequences ofTol2ends and their flanking regions. Five clones each from ex-periments B and D were sequenced. The regions outsideTol2were all confirmed to be parts of the pTar01 plasmid. Target site duplications of 8 bp, in boldface type, were observed in all the cases.
Figure5.—Locations ofTol2insertions on the pTar01 plasmid. The stippled arrows indicate in-dividualTol2copies, in accordance with the direc-tion of thetransposasegene in the element. Clones from experiment B are shown above and those from experiment D below the plasmid.
the supply of exogenous mRNA by microinjection was gene. Thus, theTol2element of thehATfamily can be expected to provide a gene tagging tool for molecular responsible for invoking transposition ofTol2.
Development of a monitoring system:Our detection techniques as alternatives to those using mariner/Tc1 family elements.
system, if applied on a larger scale, would be useful for
determining transposition frequency and, thus, finding We are grateful to H. Hashikawa, M. Sato, and S. Susaki for providing factors that affect the transposition. However, there is theO. melastigmasamples. We also thank H. Ohtsubo and Y. Sekine for helpful discussions. This work was partly supported by grant no.
a problem that remains to be solved and that is the
10216025 to A.K. and no. 09554053 to H.H. from the Ministry of
presence of 10-kb plasmids. They can be eliminated by
Education, Science, Sports, and Culture of Japan, and also by the
restriction enzyme digestion and agarose gel
electro-Takeda Science Foundation to A.K.
phoresis but a more efficient device is desirable. One possible approach is to use the StrS gene already
con-tained in the pTar01 plasmid (see materials and
LITERATURE CITED
methods). Screening with streptomycin, in addition to
kanamycin and tetracycline, would be useful for collect- Atkinson, P. W., W. D. WarrenandD. A. O’Brochta,1993 The
hobotransposable element ofDrosophilacan be cross-mobilized
ing only insertions in theStrSgene.
in houseflies and excises like theAcelement of maize. Proc. Natl. Potential as a gene tagging tool: Some elements of Acad. Sci. USA90:9693–9697.
the mariner/Tc1 family have already been applied for Calvi, B. R., T. J. Hong, S. D. FindleyandW. M. Gelbart,1991 Evidence for a common evolutionary origin of inverted repeat
developing a gene tagging system in vertebrates.
How-transposons inDrosophilaand plants:hobo,Activator, andTam3. ever,hATfamily elements have different features from Cell66:465–471.
those of the mariner/Tc1 family, for example, being Dowe, M. F. Jr., G. W. RomanandA. S. Klein,1990 Excision and transposition of twoDs transposons from thebronze mutable 4 larger in size. This might be an advantage for carrying
derivative 6856 allele ofZea maizeL. Mol. Gen. Genet.221:475–
large DNA fragments. One element,Activatorof maize,
485.
has been shown to transpose preferentially within the Fadool, J. M., D. L. HartlandJ. E. Dowling,1998 Transposition of themarinerelement fromDrosophila mauritianain zebrafish.
same chromosome (Greenblatt 1984; Dowe et al.
Proc. Natl. Acad. Sci. USA95:5182–5186.
1990;Morenoet al.1992) and, moreover, in a systematic
Greenblatt, I. M.,1984 A chromosomal replication pattern
de-further analysis, preferentially to nearby regions (Mach- duced from pericarp phenotypes resulting from movements of
the transposable element,Modulator, in maize. Genetics108:471–
idaet al.1997). Such a transposition would be beneficial
485.
to target nearby genes or regions. This might lead to
Hashimoto-Gotoh, T., A. Kume, W. Masahashi, S. Takeshita
creation of chromosomal deletions and inversions and andA. Fukuda,1986 Improved vector, pHSG664, for direct
streptomycin-resistance selection: cDNA cloning with G:C-tailing
2000 Evidence for recent invasion of the medaka fish genome mouse cell cultures. J. Mol. Biol.26:365–369.
Ivics, Z., P. B. Hackett, R. H. PlasterkandZ. Izsva´k,1997 Molec- by theTol2transposable element. Genetics155:273–281. Lam, W. L., T. S. LeeandW. Gilbert,1996 Active transposition in ular reconstruction ofSleeping Beauty, aTc1-like transposon from
fish, and its transposition in human cells. Cell91:501–510. zebrafish. Proc. Natl. Acad. Sci. USA93:10870–10875. Machida, C., H. Onouchi, J. Koizumi, S. Hamada, E. Semiartiet
Kawakami, K.,andA. Shima,1999 Identification of theTol2
transpo-sase of the medaka fishOryzias latipesthat catalyzes excision of a al., 1997 Characterization of the transposition pattern of the
Acelement inArabidopsis thalianausing endonuclease I-SceI. Proc. nonautonomousTol2element in zebrafishDanio rerio.Gene240:
239–244. Natl. Acad. Sci. USA94:8675–8680.
Moreno, M. A., J. Chen, I. GreenblattandS. L. Dellaporta,1992 Kawakami, K., A. Koga, H. HoriandA. Shima,1998 Excision of
theTol2transposable element of the medaka fish,Oryzias latipes, Reconstitutional mutagenesis of the maizePgene by short-range
in zebrafish,Danio rerio.Gene225:17–22. Actranspositions. Genetics131:939–956.
Koga, A.,andH. Hori,1999 Homogeneity in the structure of the Raz, E., H. G. van Luenen, R. Schaerringer, R. H. A. Plasterkand medaka fish transposable elementTol2.Genet. Res.73:7–14. W. Driever,1997 Transposition of the nematodeCaenorhabditis
Koga, A., H. Inagaki, Y. BesshoandH. Hori,1995 Insertion of a elegans Tc3element in the zebrafish Danio rerio.Curr. Biol.8:
novel transposable element in the tyrosinase gene is responsible 82–88.
for an albino mutation in the medaka fish,Oryzias latipes.Mol. Takeshita, S., M. Sato, M. Toba, W. MasahashiandT.
Hashimoto-Gen. Genet.249:400–405. Gotoh,1987 High-copy-number and low-copy-number plasmid
Koga, A., M. Suzuki, H. Inagaki, Y. Bessho and H. Hori, vectors forlacZ␣-complementation and chloramphenicol- or ka-1996 Transposable element in fish. Nature383:30. namycin-resistance selection. Gene61:63–74.
Koga, A., M. Suzuki, Y. Maruyama, M. TsutsumiandH. Hori,1999