Copyright 0 1996 by the Genetics Society of America
Differences
in
Crossover Frequency
and
Distribution
Among Three Sibling Species of Drosophila
John R.
True, John M. Mercer
and
Cathy C. Laurie
Department of Zoology, Duke University, Durham, North Carolina 27708 Manuscript received August 7, 1995
Accepted for publication November 10, 1995
ABSTRACT
Comparisons of the genetic and cytogenetic maps of three sibling species of Drosophila reveal marked differences in the frequency and cumulative distribution of crossovers during meiosis. The maps for two
of these species, Drosophila melanogaster and D. simulans, have previously been described, while this report
presents new map data for D. maun'tiana, obtained using a set of P element markers. A genetic map covering nearly the entire genome was constructed by estimating the recombination fraction for each pair of adjacent inserts. The P-based genetic map of maun'tiana is "1.8 times longer than the standard
mlanogastm map. It appears that mauritiana has higher recombination along the entire length of each chromosome, but the difference is greatest in centromere-proximal regions of the autosomes. The
maun'tiana autosomes show little or no centromeric recombinational suppression, a characteristic that
is prominent in melanogaster. D. simulans appears to be intermediate both in terms of total map length and intensity of the autosomal centromeric effect. These interspecific differences in recombination
have important evolutionary implications for DNA sequence organization and variability. In particular,
maun'tiana is expected to differ from mlanogaster in patterns and amounts of sequence variation and transposon insertions.
M
EIOTIC recombination is a very widespread and important component of the hereditary machin- ery of sexually reproducing eukaryotic organisms. The- oretical studies have suggested both advantages and dis- advantages to the existence of recombination, but the evolutionary forces behind the maintenance of recom- bination in natural populations are still poorly under- stood (MICHOD and LEVIN 1988). Although there is a wealth of theoretical work in this area, there is relatively little empirical data on how recombinational systems vary within and among species. Such data could provide important clues about the evolutionary forces that af- fect recombination.Modifiers of recombination are subject to evolution- ary forces such as selection, but recombination is also itself an important evolutionary force that can influence patterns of genome organization and sequence variabil- ity. For example, unequal crossing over can affect the accumulation of tandemly repeated sequences and ec- topic recombination between dispersed transposable el- ements can control their numbers and distribution in the genome (CHARLESWORTH et al. 1994). In addition, recombination plays a vital role in processes such as hitchhiking and background selection that affect levels and patterns of sequence polymorphism (AQUADRO et al. 1994). Thus, it is clear that an understanding of the growing body of data on genome-wide patterns of
Corresponding autho~; Cathy C. Laurie, DCMB/Zoology, Box 91000,
Duke University, Durham, NC 27708. E-mail: cclaurie@acpub.duke.edu
Genetics 142: 507-523 (February, 1996)
sequence organization will require information about the amount and distribution of recombination. Associa- tions between interspecific differences in recombina- tion and sequence organization or variability are likely to provide important insights into processes of genome evolution.
The rate and distribution of recombination within a genome are difficult traits to measure, so there is rather little information about how they vary within and among species. The genus Drosophila provides some important technical advantages in this area and, there- fore, has been the subject of many recombination stud- ies (CARPENTER 1988; HAWLEY et al. 1993). One of the great advantages of Drosophila is its intricately banded polytene chromosomes, which have provided a means of physically mapping genetic markers, first through cytological analysis of chromosomal rearrangements and later through in situ hybridization with cloned genes as probes. Although polytene bands vary cytologi- cally in width and spacing, their number is very highly correlated with DNA content (based on data from HEINO et al. 1994), so they provide a good measure of physical distance over ranges of 2 2 0 bands.
508 J. R. True, J. M. Mercer, and C . C . Laurie
centromeric suppression because the CE is very low in euchromatin proximal to the centromere, but rises gradually as distance from the centromere increases. This feature is typical of many different kinds of organ- isms, based on cytological observations of chiasma dis- tribution (JONES 1987). In addition, the Xchromosome of D. melanogaster shows a strong telomeric suppression of crossing over, which may also be present in reduced form on the autosomes. There are also many local varia- tions in recombination rate, but it is not clear whether those variations are characteristic of the species or whether they may be due to measurement errors or to intraspecific variation in recombination rate.
Genetic control of the pattern of recombination in D. melanogaster is not well understood, but appears to be due to the combined action of local cisacting factors that affect exchange frequency, transacting factors that control both the level and distribution of exchanges and chromosomal structures such as the centromere and telomere that seem to exert a polar influence over exchange frequency (CARPENTER 1988; HAWLEY et al. 1993). A clear example of cisacting control is the lack of recombination in heterochromatin regardless of its chromosomal position (BAKER 1958; ROBERTS 1965). In contrast, the polar effect of the centromere on euchro- matic recombination is not due to the intrinsic proper- ties of sequences located close to the centromere. Stud- ies of translocation and inversion homozygotes have shown that recombination in an interval changes sig- nificantly when its distance to a centromere is altered (BEADLE 1932; SZAUTER 1984). In addition, several mei- otic mutants have been described that alter the distribu- tion of exchange by making recombination rate more nearly proportional to physical distance along the chro- mosome and thereby reducing the centromere effect
(BAKER and CARPENTER 1972).
Although the standard genetic map of D. melanogaster (LINDSLEY and ZIMM 1992) is often regarded as charac- teristic of this species, considerable intraspecific varia- tion in recombination rates has been documented (re- viewed by BROOKS 1988). For example, BROOKS and MARKS (1986) studied variation due to second chromo- some modifiers of recombination. They found that the total amount of crossing over on the second chromo- some varied by 12- 14%, with larger variations for spe- cific intervals. The distribution of exchange was also affected, with intervals in the proximal euchromatin showing more variation than other regions. In addition, artificial selection for changes in recombination in spe- cific intervals has produced significant responses, pro- viding further evidence for variation of recombination modifiers in natural populations (e.g., CHARLESWORTH and CHARLESWORTH 1985).
Drosophila is one of the few genera in which several species are genetically well mapped so that interspecific comparisons of some of the properties of the recombi- nation system can be made. Large differences in total genetic map length are observed between distantly re-
lated species in the genus. For example, the genetic map of D. uirilis (-975 cm) (GUBENKO and EVGEN'EV 1984) is approximately twice as long as that of D. pseudo- obscura (-450 cm) (ANDERSON 1993), which itself is nearly twice as long as that of D. melanogaster (-280 cm) (LINDSLEY and ZIMM 1992). These differences have evolved over very long time periods and may be associ- ated with major changes in karyotype and in genome size and organization. D. uirilis diverged from pseudoobsc-
ura and melanogaster 30-60 mya and its euchromatic genome is -150 Mb compared with
-
110 Mb for mela-nogaster (HARTL and LOZOVSIWYA 1995, p. 142-143).
The karyotypes of all three species (melanogaster, uirilis,
pseudoobscura) differ in the numbers of metacentric and acrocentric chromosomes (PATTERSON and STONE 1952). Because few genetically mapped markers have been cytologically localized in uirilis or pseudoobscura, comparisons of the distribution of recombination along the chromosome have not been made.
This report deals with divergence in recombination frequency and distribution among three sibling species of the melanoguster subgroup: the cosmopolitan species, D. melanogasterand D. simulans, and an island endemic, D. mauritiana. It is estimated that melanogaster diverged from the ancestor of simulans and mcmritiana -2.5-3.4 mya, whereas simulans and mauritiana diverged from one another about 0.6-0.9 mya (HEY and KLIW
1993). All three species readily hybridize with one an- other, but all hybrids with melanogaster are sterile, whereas mauritiana/simulans hybrids are male sterile but female fertile (ASHBURNER 1989). The karyotypes of these species are the same and the polytene chromo- some banding patterns are nearly identical except for a large paracentric inversion on ?R and a few other very small rearrangements that distinguish melanogaster from the other two species (LEMEUNIER and ASHBURNER
1976). The recombination system of melanogasler is, of course, well studied, but there has been relatively little published information about recombination in its sib- ling species. Some comparisons between melanogaster and simulans are possible because a fairly large number of mutants with homology to corresponding melunogas- ter genes have been isolated and mapped in simulans (STURTEVANT 1929; OHNISHI and VOELKER 1979). Here we report a fine-scale genetic and cytogenetic map of L). mauritiana, which provides a basis for detailed com- parison with melanogaster in the amount and distribu- tion of recombination.
The original motivation for producing a genetic map of D. mauritiana was to provide a set of genetic markers that could be used for analyzing the genetic basis of morphological divergence between mauritiana and its sibling species. Because we wished to used these mark- ers for introgression and for localization of quantitative trait loci, we required that they be easily scored and well distributed across the genome. For this purpose, we produced a set of P-element insertions into a white-
Recombination in Drosophila Species 509
white' eye color marker. These insertions were first lo-
calized by in situ hybridization to the polytene chromo- somes. Then, by estimating the recombination fraction for each pair of adjacent inserts, we produced a genetic map covering nearly the entire genome. A comparison with melanogaster reveals striking differences in overall genetic map length and in the distribution of recombi- nation inferred from the relationship between the ge- netic and physical maps.
MATERIALS AND METHODS
Drosophila culture conditions: Flies were raised on stan- dard cornmeal molasses agar medium at room temperature, except where noted otherwise.
Drosophila strains: D. melanogaster: Wild-type isofemale lines BWA27 (Benin, W. Africa) provided by R. SINGH and Raleigh144 (Raleigh, NC 1982) collected by P. BARNES. y w
provided by G. RUBIN. fprovided by Mid-America Drosophila Stock Center. v and g provided by Bloomington Drosophila Stock Center. b cn bred from b AdhnU248 cn b u stock pro- vided by M. ASHBURNER (bw' allele from Raleigh 144 isofem- ale line).
D. simulans: white (No. 13) provided by J. COYNE. Wild- type isofemale lines S . France provided by R. SINCH and P. I. Australia provided by J. BARKER. y w bred from y w mfstock provided by C-I. Wu (m+
f
from S. France isofemale line). v, g, a n d j provided by Bloomington Drosophila Stock Center.b cn bred from b and cn stocks provided by Bloomington Drosophila Stock Center.
D. mauritiana: white provided by J. A. COYNE. Wild-type iso- female lines Petite Reviere provided by D. HICmYand Robert- son provided by H. ROBERTSON. y w derived from spontaneous
yellow mutation that arose in the w stock. v, g, and j provided by Bloomington Drosophila Stock Center. b n bred from
b and cn stocks provided by Bloomington Drosophila Stock Center. Interspecific homology for all mutant loci was con- firmed by failure to complement in crosses with melanogaster
(and/or other simulans clade mutant alleles of known homol- ogy to melanogaster) performed in our laboratory and/or by J. A. COWE (personal communication).
Polytene chromosome band index: The band index used in this paper is the number of polytene chromosome bands between a cytologically localized genetic marker and the cen- tromere. It provides a good measure of physical distance for regions of 2 2 0 bands, because the correlation between DNA content and band number for each chromosome arm is very high ( T > 0.99 using data from HEINO et al. 1994). The band
indices used here comply with the revised maps of C. B. and P. N. BRIDGES (in LINDsLEY and ZIMM 1992). The band index for a given gene or Pelement insert is estimated as the mid- point of the band range to which it has been mapped cytologi- cally. For mauritiana and simuluns, band indices within the large 3R inversion (relative to melanogaster) are renumbered to obtain increasing distance from the centromere.
P-element transformation and characterization of trans- formant lines: Host embryos of D. mauritiana white- were in- jected with P[ lac-w'] (BIER et al. 1989) and p~r25.7WC (KARESS
and RUBIN 1984). Single insert lines were selected for further work by Southern hybridization, using the 4.25-kb Hind111 fragment of P[lac-w']; and digesting transformant line geno- mic DNA with XbaI and BglII.
These P[lac-zo'] insertions appear to be very stable, as ex- pected because rnauritiana has no endogenous P elements to provide transposase activity (BROOKFIELD et al. 1984). Many insertions have a characteristic eye color pattern and intensity that did not change over months of culture.
In Situ hybridizations: Each mauritiam insert was localized cytologically by in situ hybridization to the polytene chromo- somes. The photographic maps of melanogaster (LEFEVRE 1976) were used for mapping, because the banding patterns of these two species are nearly identical. However, the poly- tene chromosomes of mauritiana are smaller than melanogaster,
and it is much more difficult to obtain high-resolution prepa- rations. Therefore, inserts generally were localized only to
the nearest lettered division (or border between two lettered divisions). Larvae for cytology were raised on the medium of WHEELER and CLAWON (1965) and in situ hybridization was performed according to LIM (1993).
Construction of a D. mauritiana genetic map using 41ac- w'] inserts: All of the inserts mapping to the same cytological position were pooled and a single representative of each pool was used for constructing a genetic map. The recombination fractions between adjacent pairs of insert locations were esti- mated as follows. Two single-insert homozygous lines were crossed and the doubly heterozygous female progeny were then crossed to w- males. In many cases, it was difficult to distinguish P[ lac-w+] heterozygotes from homozygotes, so the recombination fraction was estimated as twice the frequency of w- recombinants.
Using this procedure, the expected variance of the recombi- nation fraction estimate is 4[x( 1 - x ) ] / n , where x is the fre- quency of w- recombinants and n is the total number of progeny. The average sample size was 3458 progeny.
For crosses to estimate recombination fraction, 515 vials were set up with 4-5 virgin F, females (5-6 days old) and 4- 5 w- males (of varying age). These parents were transferred to new vials after 5 days and the transfer set was cleared after another 5 days. Progeny of both sets of vials were scored on the 13th, 15th, and 17th days of culture. Due to the large scope of this experiment, nine separate sets of crosses were performed during the course of 1 yr. Room temperature var- ied somewhat during these experiments; ranges for each set were: ( I ) 21-24", (2) 21-24", (3) 22-26", (4) 23-26", (5) 21-
22", (6) 21-22", (7) 20-22", (8) 20-22", (9) 25" (incubator). Nevertheless, crosses repeated in more than one experiment gave very similar results. For example, one interval (19AB/ 19BC) gave independent estimates of 6.1 (21.4) and 6.4 (?1.3) and another interval (24CD/26B) gave estimates of 11.5 ( ? 1.7) and 11.2 ( ? 1.7), where the ? figures in parenthe- ses are 95% confidence intervals.
Recombination fractions were converted to map distance using the mapping function of FOSS et al. (1993; with nt = 4),
which accounts very well for the pattern of interference on the X chromosome of melanogaster. This mapping function has very little effect on recombination fractions less than
-
15 cM (changing only the second decimal place). Because nearly all of the recombination fractions used in the mauritiana map construction are < I 5 cM, the map is influenced very little by this conversion.Map distances from each autosomal centromere to the nearest flanking Pelement markers were estimated by inter- polation. A coefficient of exchange (centimorgans per band) was calculated from the map distance between those flanking markers and then multiplied by the number of bands between each marker and the centromere. For the X chromosome, the centimorgan between the centromere and nearest marker at 19EF was estimated in a similar manner using the coeffi- cient of exchange for the interval between markers at 19BC and 19EF.
D. melanogaster map data: The melanogaster data were pooled from two sources: FlyBase (1994), which provided ge- netic map and cytological positions for a set of loci mapped genetically to within 0.1 cM and cytologically to an interval less than six bands (identified by E. KINDAHL and C. AQUADRO,
Chromosome 3
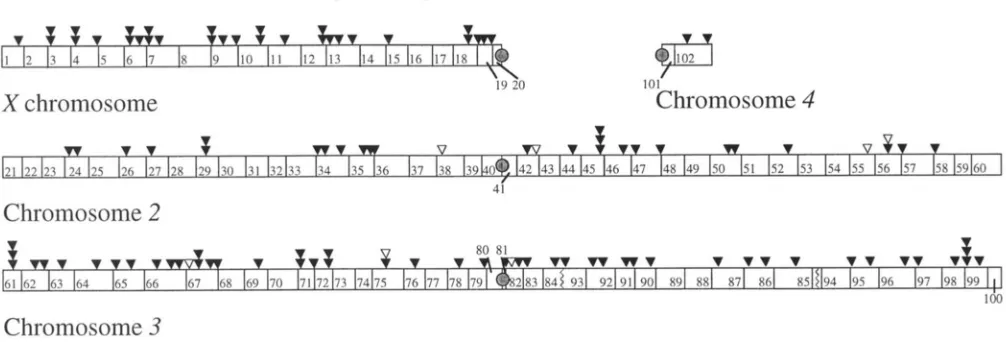
FIGURE 1 .-Cytological positions of P [ Innu'] inserts in D. mnuritinna. Each symbol represents a separate line bearing a single Pinsertion that is homozygous viable. Black symbols represent homozygous fertile inserts; white symbols represent homozygous sterile inserts. Stacked symbols represent inserts at approximately the same position. Jagged lines on the right arm of chromosome
3 represent the breakpoints of a fixed inversion difference between D. mrln,nogmter and D. mauritiana.
for a set of transposable element markers cytologically local- ized and genetically mapped in relation to standard reference loci (summarized in Figure 11.1 of A~HRURNER 1989). The two data sets agree substantially.
The major features of the standard wlanognsto-genetic map were established by the MORGAN group between about 1913 and 192.5 (MORGAN rt al. 1925), using a set of reference loci. The current genetic map positions of these reference loci
(LINI)SI.EY and ZIMM 1992) are the same as those given by MORGAN rt al. (1925) and the current map lengths of each chromosome are only slightly longer than the 1925 maps. Over the past 70-80 years, the mrlanogmter map has been built up by mapping new loci with respect to the reference loci, either directly or indirectly. Map distances between refer- ence loci were calculated by MORGAN et nl. (1925) from a
system of empirical curves based on recombination fractions for closely spaced markers. In essence, the map distance be- tween two points on the wlnnogmtPrmap is the sum of recom- bination fractions for intervals small enough that the probabil- ity of double crossing over is negligible.
D. simulans map data: Genetic map data for simuluns and demonstrations o f interspecific homology with wlanogmtPr
are mainly from the compilations of STURTEVANT (1929) and OHNISHI and VOELKER (1979), with additional data from OH- SISHI and VOELKER (1981) and P U R 0 (1971). In most cases, the cytological map positions of the simuluns genes have not been determined directly, but are assumed to be the same as
the rnrlnnngnskr genes to which they are homologous. Genetic map positions of simulnns mutants were established originally by STTIIRTEVANT (1929) using a system of reference loci similar to those used in constructing the melanogmkr
maps. However, because of the relative paucity of genetic markers in simulnns, the map distances between reference loci were given as observed recombination fractions, uncorrected for possible multiple crossovers. This procedure can result in underestimates of map length. Thus, in calculating total map lengths for each chromosome, we have used the mapping f h c t i o n of FOSS rt nl. (1993; m = 4) to convert observed recombination fractions reported by STURTEVANT (1929) for each pair of reference loci. This conversion increases the map distance estimates by only a small amount compared with using the sum of recombination fractions (ir., 0.2% for the X , 8.9% for the second chromosome and 2.3% for the third chromosome). For analysis of the relationship between ge- netic and physical map distances, only X and third chromo- some loci were used because there are few second chromo- some genes with demonstrated homology to mrlunogmter
genes. In this analysis, map positions given in the literature were used without conversion.
Relationship between genetic and physical map distances:
The incomplete beta function, B(z, a, 0 ) . was chosen to pro- vide a flexible but simple phenomenological representation of the relationship between cumulative genetic and physical map distances from the centromere. The observed physical map distance (band index) was rescaled to the interval 0,l to correspond with the domain of definition of B(z, a, b ) . The Levenburg-Marquardt algorithm with equal weighting was used to minimize the sum of squared residuals to obtain esti- mates of the parameters a, b and c for the distribution func- tion c B(z, a, b ) . This algorithm was implemented using Math- ematica 2.2 (WOLFRAM 1991) supplemented with 2.3 prerelease versions of the packages RegressionCommon.m and Non1inearFit.m. In addition, the following derivatives are required by the algorithm and must be entered explicitly.
dB(z, a, b ) / d a = -i'SF,(a, U, 1
-
6: 1+
a;z)/d+
In (z) B(z, a, b) d B(z, a, b ) / d b = (1 - ~)"3F2(b, b, 1-
U ; 1+
b, 1+
6:1
-
z ) / #-
In (1-
z) B ( l - z, b, a)+
B(u, 6) [ W ( b )-
@ ( a+
b ) ]where 3F2 is a generalized hypergeometric function, 9 is the psi or digamma function, and B ( a , b) is the beta function.
The coefficient of exchange (CE, centimorgans per band) is the slope of the curve describing the relationship between genetic and physical map distances from the centromere. Thus, the derivative of the incomplete beta function that de- scribes this relationship, which is proportional to the beta density function, provides a description of how the coefficient of exchange varies along a chromosome arm. An alternative approach to estimation of the beta function parameters is to fit the beta density function to observed values of CE corre- sponding to discrete marker intervals (as in MORTON d al.
1976) rather than fitting the incomplete beta function to cumulative map distance. We chose the latter approach be- cause estimates of CE for small intervals are subject to consid- erable measurement error. Furthermore, map distances be- tween adjacent markers in the standard melunogmter map are often not estimated directly, but rather each marker may have been positioned in relation to reference loci at some distance from the interval.
Recomhination in Drosophila Species .51 1
chromosome cytological length Fl(;lxb. 2.-Differences i n overall genetic map
range (cM) length hetween D. tnPlm/og(~.s/rr antl 11. nmrri-
X
2
3
I (1~1.2, 19F.7-pI) 65.9 /innn. Cytological rangcs for n w / r r i / i m u refer to
-
( 2 4 A 1 - 3 3 s 8 B ) 92.n those markers on the standard map w i t h posi-=
(61,,l-2,99D) lol,n BASE 1994). The following nw/anogm/rr loci were( I EF. I9EF) 111.1 the most distal P [ / n n o ' ] markrrs o n rach chro- mosome; those for 11. melanogaster refer to
(24A. 5RA) 14n.5 tions closest to the nrcr?o-i/innn endpoints (F1.y-
(61CD,99D) 2 1 1,3 used as these endpoints: . s ? r ( n m ) and o n c on the Xchromosome, oddaantl m ~ i S 3 3 2 o n second chro-
0 D. nw/unopsrer 258.9 mosome, and cnfc andjnnA on the third chromo-
=
D. rnuuririunu ToTAL 462.9 sOllle.Direct comparisons o f recombination fractions were made for The average recombination fraction is ().().',, Ivith a maxi-
four i n t e n d s lnarked hy mutants at loci holnologous i n the mum of 0.20, which pro\,ides a fairly del,se map of the
chromosome). These fractionswcre estimated simultaneously entire genome, except for the termina1 regions Of the
three species ( y r u , rkgand g-/m the X a n d b c n o n the second
for ?nr/crnogu.s!m, n/rtztn'/irrnrr, . s i m r / m . s and nrtrun'/icrnn/simzL/nn.~ second chromosome (see Figure 1 ).
Ilyhrids. Possible biases in recombination fraction estimates:
Crosses t o determine recombination fractions for the inter- The estimation o ~ r e c o l n ~ ~ n a t ~ o l l fi-actions can be biased
vals y r o and IFcn were performed using clouhle-mutant tester
stocks. Tester stock females were mated with wilcl-type males by unequal viabilities of parental and recombinant geno- from cac~l of tlvo isofemale lines anti the F, females were types. In this case, the estimation invokes comparing hackcrossed to tester stock males. No douhle mutant tester one recomhinant genotype with no inserts to three other
were mated with
/
males, and then F, females were matedfirst generation 71 females were mated with g males, gfemales lines llave heen easy to mainmin in tIle laboratoy, illdi-
w i t h wild-type males. Progeny were scored using the same cating generally good viability and fertility. In addition, design and schedlllc as tile backcrosses wit11 p[/nc-7[J'] mark- the viabilities of six randomly chosen inserts were esti-
ers. Recar~sc all parental and recomhinant genotypes could mated quantitatively by crossing a single insert line to
he scored unamhiguously in these crosses, the variance of the ?(,- and then backcrossing to
7(r
again. F ~ ) ~ eac]l insert, recombination fraction estimate is [ x ( 1 - x ) ] / n where x isthe frequency of recombinants and n is the total numher of 1.500-1900 progeny were scored and there were no sig- progeIly. The averagc ,,a~l,e of n in this experiment ,,,as 1336. nificant departures from the expected 1:1 ratio. Thus,
Tests for Xchromosome nondisjunction: Six randomly cho- we have no evidence for a bias in the estimates of recom-
stocks 101- the v g a n d g-/intends were availahle. TIllls, in the genotypes having one or two inserts. The IT/(/rqa+] insert
sen X-linked ['I IUf-r/l' ] insert lines and l%'O wild-type lines (PC- bination fractions due to differential viabilihr.
tite Reviere antl Robertson) o f mauritiana were tested for evi-
dence of X chromosome nondisjunction. Four-day-old virgin A potential bias involving Xchromosome recombina- females (wild tvpe o r homozygous for an X-linked insert) were tion estimates is the occurrence Of
patroclinous
malesmated intlivitluaIIy t o three 7(1 males and the vials were trans- due to XX nondisjunction or chromosome loss in fe-
ferred, cleared an-d scored according to the same scI1edule as for crosses to estimate recombination fractions. An average of
24.5 single females per line were examined and an average of 3462 progeny per line were scored. Putative X 0 males ( r o - )
were mated t o 2-3 ru- females to test fertility. In addition, females homozygous for one second o r third chromosome insert were mated t o Z ~ P - males as a control. These crosses were mass matings o f 4-5 pairs i n each of 15 vials.
RESULTS
The cytological
and
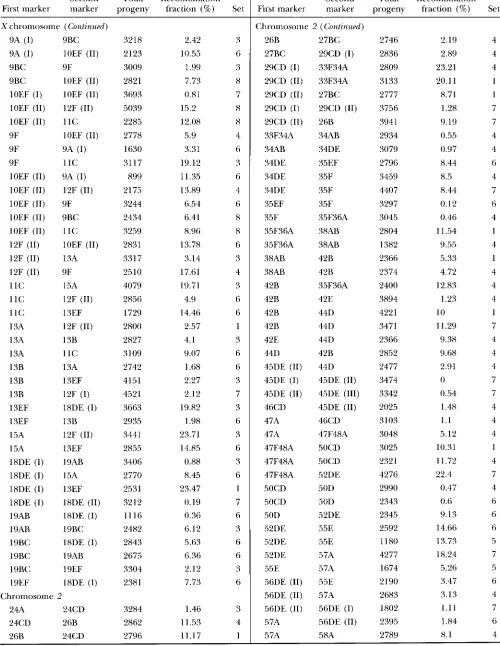
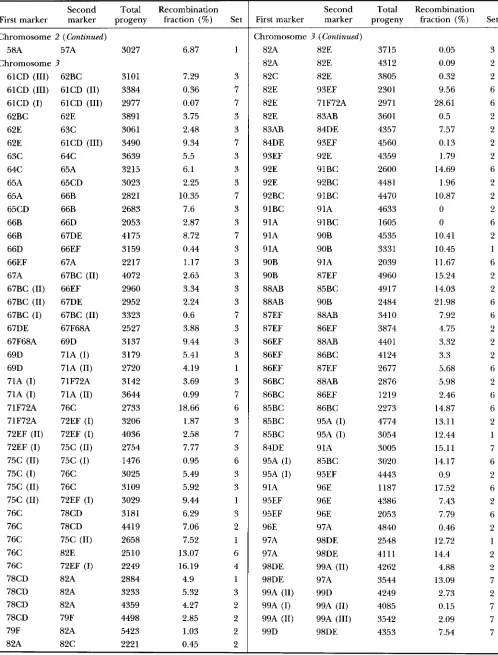
genetic maps of D. mauritiana: A total of 114 P[lac-711+] inserts were localized by in situ hybridization to the salivary gland polytene chromo- somes (Figure 1 ) . These locations represent a total of 94 clearly distinct positions, which are well distributed across the genome (although somewhat sparser on the second chromosome than the Xor third). This sample represents only the inserts isolated as single-insert, ho- mozygous viable transformant lines, but the distribution is very similar to that found in a mdclnogastPr sample that also included lethal lines (LAURIE-AIII.RERG and STAM 1987).The genetic map was constructed by estimating the recombination fraction between each pair of adjacent insert positions. The raw data are provided in APPENDIX.
male gametes. Females tmnsheterozygous for X-linked inserts can produce X0 males that lack an insert and therefore have white eyes, just like male progeny with a recombinant X lacking both inserts. To determine the frequency of exceptional males, females homozy- gous for an X-linked insert from each of six different lines, as well as females from each of two wildtype lines, were crossed with 70- males. All eight of these lines
produced patroclinous 7u- males at about the same rate,
which ranges from two of 4340 progeny (0.05%) to four of 2519 progeny (0.16%), with a pooled value of 29 of 27,696 progeny (0.10%). All but three of the 29 70-
males obtained in this experiment were sterile, as ex- pected for X 0 males. Among the three fertile males, which were crossed to ?(I- females, two produced all
white progeny. The third male produced male progeny that were all white and female progeny that all had the orange eyes typical of a single insert genotype. Evi- dently, this male had an X-linked insert with v e y low o r n o rnini-7uhil~ expression, while the other two may have had XXY mothers.
512 J. R. True, J. M. Mercer, and C. C . Laurie
o
D.
muuritiana
*
D.
simulans
D.
mehogaster
a.
X
major cytological division
0 Y 8 7 6 5 4 3 2 I 0 Y 8 7 6 5 4 3 2 1 2 1 1 1 1 I I 1 I I I
L I I I I I I I 1 1 1 1 1 1 I I l l l l l
125
-
O P
b. X
major cytological division
O Y 8 7 6 5 4 3 2 1 0 Y 8 1 6 5 4 3 2 1 2 1 I l l I I I I 1 I
J I I I I I I 1 1 1 1 1 I f l l ’ f l J
125
-
1O P
,/
75
f
50b
i?
3
f
250
0 250 5 0 0 750 IO00
bands from centromere
c. 2L
4 3 3 3 3 3 3 3 3 3 3 2 2 2 2 2 2 2 2 2 0 9 8 7 6 5 4 3 2 1 0 9 8 7 6 5 4 3 2 1
major cytoiogical division
8’
I t I
250 500 750
bands from centromere
e. 3L
major cytological division
O Y 8 1 6 5 4 3 2 l O Y 8 1 6 5 4 3 2 I
8 1 7 1 7 7 1 1 1 7 1 6 6 6 6 6 6 6 6 6
I I I
0 750 500 250
I
bands from centromere
0 750 250 500 IO00
bands from centromere
d.2R
75
e!
B
0
2
508
E
‘
253
0
major cytological division 4 4 4 4 4 4 4 4 4 5 5 5 5 5 5 5 5 5 5 6
1 2 3 4 S 6 7 8 Y 0 I 2 3 4 5 6 7 8 9 0
L l 1 l I I I / 1 1 1 1 ‘ 1 ’ 1 ’ “ ‘ 1
I
I
0 1000 250 750 500bands from centromere
0 250 1000 750 500
Recombination in Drosophila Species 513
a very small amount
(2
x
0.10% Equation 0.20 cM). We did not attempt to correct this bias in analyses of the relationship between genetic and physical maps dis- cussed below, but we did correct the estimate of total X chromosome map length by subtracting the product of the number of intervals(22)
and the bias in the rate estimate per interval (0.20 cM), which is 4.4 cM.In addition to the X chromosome inserts, crosses were made between females homozygous for a second or third chromosome insert and w- males. No white progeny of either sex were obtained from a total of 5028 progeny. These X , second and third chromosome insert crosses provide evidence of the expected P insert stability, because no white females were observed in a total of 32,997 progeny examined.
Two inserts were localized to the fourth chromo- some, providing the possibility of estimating a recombi- nation fraction for this chromosome, which is normally achiasmate in
D.
melanogaster (HOCHMAN 1976). How- ever, we have been unable to obtain a reliable estimate of recombination between these two fourth chromo- some inserts because of complications that appear to be due to fourth chromosome nondisjunction and pos- sibly a suppression of mini-white gene expression in one of the fourth chromosome inserts in X 0 males.Interspecific comparisons of total genetic map length: Figure 2 presents a comparison of total genetic map length between the Pbased mauritiana map and the standard melanogaster map (LINDSLEY and ZIMM 1992; FLYBASE 1994). These comparisons were made by find- ing genetic markers in the melanogaster map that are located at essentially the same cytological positions as the terminal markers in mauritiana. For the extent of the genome spanned by the P[ lac-w+] markers, the mau- ritiana genetic map is l .8 times longer than the standard melanogaster map. Ratios of mauritiana to melanogaster genetic map lengths are 1.8, 1.6, and 2.1 for the X, second, and third chromosomes, respectively.
The map lengths of melanogasterand simulunswere com- pared between pairs of homologous loci near the chro- mosomal termini. The map distances for melanogaster and simulans, respectively, are 66.0 and 66.3 between
y
and bb; 84.0 and 108.1 between @and P U , and 81.5 and 135.9 between j v and ca. Ratios of the simulans to mlanogaster map lengths are 1 .O, 1.3 and 1.7 for the X, second and third chromosomes, respectively. The total simulans map is-
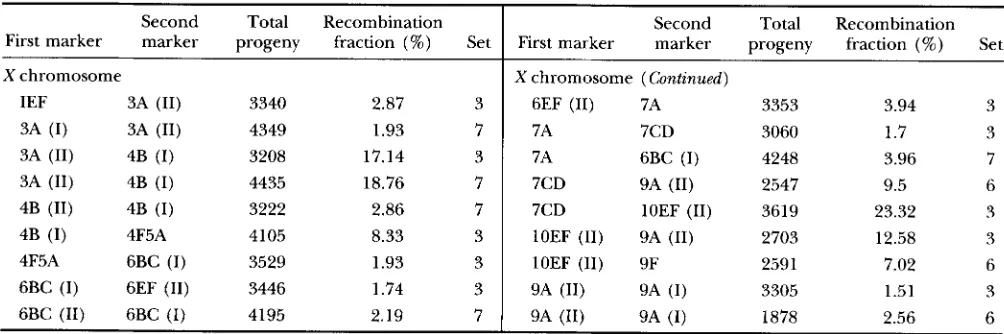
1.3 times longer than the melanogaster map.Relationship between genetic and physical maps: The relationships between the genetic and physical maps for each chromosome arm in melanogaster, mauri- tiana, and simulans are shown in Figure 3. The abscissa represents the physical distance between a genetic marker and its centromere in terms of the number of polytene bands, which is proportional to DNA content. The ordinate represents the genetic map distance, which is proportional to the expected number of cross- overs per meiosis between the genetic marker and its centromere (50 map units = crossover). The fitted curves in Figure 3 show that this relationship is de- scribed very well by an incomplete beta function, which has two shape parameters.
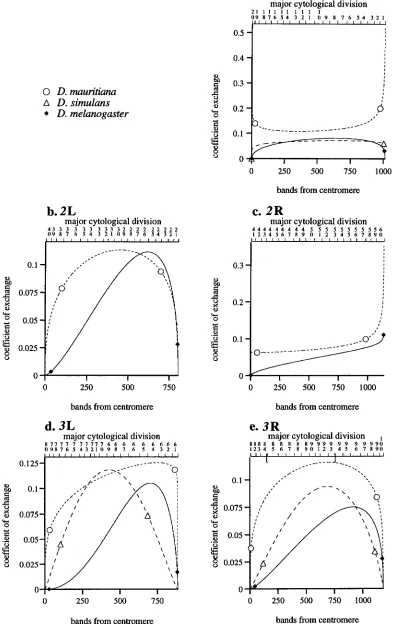
The slope of the incomplete beta function is an esti- mate of the coefficient of exchange (CE) , a measure of the expected number of crossovers per unit of physical distance (centimorgans per band). The differentiated functions (which are proportional to the beta density function) are shown in Figure 4. These plots provide a rough estimate of how the coefficient of exchange var- ies along each chromosome arm at least within the range of observed data, the endpoints of which are marked in Figure 4. Outside that range (near the ter- mini of each chromosome arm), the CE curves are ex- trapolations that probably do not accurately represent the recombination pattern. Even within the observed range, distortions of the real situation may occur be- cause measurement errors of terminal data points often have an undue influence on parameters estimated from curve fitting. Therefore, we can only interpret broad features of the CE curves, since the reliability of some of the detailed shape variation is unknown.
In melanogaster, the crossover distributions differ markedly among the chromosome arms, particularly between the Xand autosomes. On each autosomal arm, the CE (slope of the curves in Figure 3) is low near the centromere, rises gradually in the proximal half and then more steeply in the distal half. Near the telomere, there may be a decrease in the CE for 2L, ?L and
X,
but probably not for 2R. These data indicate a strong “centromere effect” (recombination suppression) for each of the autosomal arms, although it is somewhat less strong for 2R than for the others. A suppressing effect of the autosomal telomeres may also be present, but is not pronounced. In contrast, the Xchromosome shows only a slight depression near the centromere, butFIGURE 3.-Relationship between genetic and physical distance for D. melanogaster, D. simulans and D. mauritiana. The X is represented twice because the melanogaster and simulans data points for this chromosome largely coincide. The brackets within the major cytological division scale for ?R indicate breakpoints of an inversion that distinguishes melanogaster from the other
two species. The fitted curves are proportional to the incomplete beta function such that genetic distance from the centromere equals cB(z, a, 6), where z is standardized physical distance (band index) from the centromere and c, a and 6 are estimated parameters (see MATERIALS AND METHODS). The parameter estimates are: melanogastm r.x = 84.3, ax = 0.86, bS = 0.73, czl. =
143.5, aZ1. = 1.37, 621. = 1.29, c213 = 67.2 a2,< = 0.99, 6 2 ~ = 0.74, c3l. = 127.1, a31. = 1.27, 631. = 1.05, rJR = 194.6, a3R = 1.31, bjll =
1.22; mauritiana: ex = 129.9, ax = 1.44, bX = 1.26, cZl. = 244.6, aZ1. = 2.43, bZ1. = 1.43, czIl = 82.4, aZ1( = 1.64, bZR = 0.93, cgl, =
400.7, a3/. = 3.27, 631~ = 1.59, c3R = 217.6, a 3 ~ = 2.31, 63,< = 1.38 ; simulans: ex = 83.2, ax = 1.14, bx = 1.07, cjI. = 542.7, a3/, =
2.14, 671, = 2.23, r7R = 411.1, U ~ R = 2.13, 6 7 H = 1.81. The total band numbers for each arm are: X = 1011, 2L = 803.5, 2R =
514 J. R. True, J. M. Mercer, and C . C. Laurie
0
D.
muritiana
A
D.
simulans
4
D. melanogaster
rcl 0b. 2L
major cytological division
4 3 3 3 3 3 3 3 3 3 3 2 2 2 2 2 2 2 2 2 119 X 7 6 5 4 3 2 1 O Y X 1 6 5 4 3 2 1
a.
X
2 1 I l l 1 1 I I 1 I
O Y X 7 6 5 4 3 2 I 11 Y X 1 6 5 4 3 2 1
major cytological division
I I 1 l l l l
0.3
-
I0.2
-
0.
""-
0.1
-
"--0 250 500 750 1000
bands from centromere
c. 2 R
major cytological division
1 2 3 4 5 6 7 8 ~ n 1 2 3 4 5 6 7 8 9 0 4 4 4 4 4 4 4 4 4 5 5 5 5 5 5 5 5 5 5 6
I I I I I I I 1 1 1 I I 1 1 1 I 1 1 1 1 I
0.2
0 . 3 1 I
O . l k
__
-
0'"""*
0 ---
0
0 750 500 250 750 0 500 250 1000
bands from centromere bands from centromere
d. 3L
e. 3 R
major cytological division major cytological division I
o Y x 7 6 5 4 3 2 1 0 U 7 1 1 7 1 7 1 7 7 7 6 6 6 6 Y n I 6 6 5 4 3 2 6 6 6 6 I u u n x u n x x X Y Y Y Y Y Y
Y Y Y Y U 1 2 3 4 5 6 I X Y O I 2 3 4 5 6 I U Y I I
1 1 1 1 I I 1 I l l I I I I I I I I I I I I I I I I I 1 1 1 I $ I I I I I I I
0.125-
-
"""
""" "-" "
""
/
.-
\9
""
-.
/."
,
/ '0 500 250
750 1000 750 0 500 250
bands from centromere bands from centromere
FIGURE 4.-Coefficient of exchange curves for U . melanogaster, D. simulans and U . mauritiana. These curves are the derivatives
Recombination in Drosophila Species
TABLE 1
Interspecific comparison of recombination in centromeric regions
515
D. mauritiana D. melanogaster"
Chromosome Cytology cM/Band Cytology cM/Band
X
2
3
NA
38AB, 42B
79F, 82A
NA
0.031 0.025
19E7-F1, 20EF
38B3-6, 43A1-3 77EF, 83D5-El
0.0o2oh
0.0045' 0.0033"
FLY BASE (1994). " Between unc and bb.
Between tyrl and p k . 'Between ri and Tpl.
has a marked telomere effect. There is little variation in CE along the X in comparison with the autosomes. These general features have been noted previously (e.g.,
LINDSLEY and SANDLER 1977).
The total map length differences noted above dem- onstrate a marked difference in the frequency of cross- overs between melanogaster and mauritiana for all of the chromosome arms. In addition, there are clear differ- ences in the general pattern of crossover distribution on the autosomal arms. The autosomes of mauritiana show much less regional variation in the CE than those of melanogaster. In particular, none show the pro- nounced centromere effect that is so characteristic of melanogaster autosomes. Although the differences in crossover frequency on the autosomes are most pro- nounced in the centromere-proximal regions, the curves in Figure 4 suggest that the CE of mauritiana is generally higher even in distal regions, with the possible exception of 2L. On the X , both species have a more nearly constant CE, but mauritiana values are higher. Although these patterns are evident from the fitted curves in Figures 3 and 4, there could be local hot or cold spots for recombination in which the generaliza- tions do not hold.
Table 1 provides additional information about the CE in the most centromere-proximal regions. For chro- mosomes 2 and 3, the CE is an order of magnitude less in melanogaster than in mauntiana. Because there is no marker at the mauritiana X chromosome centromere, a direct estimate of recombination in this region is not possible. It should be noted that even though the mela- nogaster X shows relatively little long-range centromeric suppression of recombination in comparison with the autosomes, the CE from 19E to 20F on the standard map is comparable with the CEs for the centromeric regions of the autosomes, which do show long-range suppression. The reduced crossing over in the very proximal Xeuchromatin is apparently due to proximity to the centromere, because crossing over increases when that region is moved distally by inversions (see SCHALET and LEFEVRE 1976).
Figures 3 and 4 also provide information about the frequency and distribution of crossovers in simulans compared with the other two species. The X chromo-
some of simulans appears to have essentially the same frequency and distribution as melanogaster. Data for the third chromosome indicates that simulans has an inter- mediate level of centromeric suppression. As noted ear- lier, the third chromosome genetic map of simulans is -70% longer than melanogaster. This length difference is not due to a uniform increase in recombination throughout the chromosome, but appears to be due mainly to a higher CE in the proximal half of each arm. In fact, the melanogaster CEs appear to be higher near the distal regions (Figure 4).
Direct comparisons of recombination fraction: The "standard" genetic maps of melanogaster, simulans and
mauntiana compared above were constructed in differ-
ent laboratories and at different times. To provide more direct comparisons among these species, we estimated the recombination fractions for each of four intervals defined by homologous loci
(yw,
wg and gf on the X and bcn on the second chromosome). Estimates for each species (as well as mauritiana/simuZans hybrids) were made simultaneously and under the same environ- mental conditions. In addition, for two of these inter- vals, double mutant tester stocks were available, so intra- specific variation in recombination rate was assessed by crossing the double mutant to each of two wild-type strains. The results are summarized in Figure 5. This figure also provides recombination values from the stan- dard genetic maps of melanogaster, simulans and mauri- tiana (see legend for details).There is significant intraspecific variation in recombi- nation frequency, as found previously in other studies (BROOKS 1988). For the
yw
interval, each of the three pairwise intraspecific comparisons is significant in a chi- square contingency test. For the bcn interval, the mela- nogaster intraspecific comparison is highly significant, but the other two intraspecific comparisons are not. Some of these differences are quite large. For example, the two melanogaster strains differ by approximately sev- enfold in recombination fraction for the y w and bcn intervals, but in opposite directions. Such large differ- ences suggest the involvement of inversion polymor- phism. However, examination of the polytene chromo- somes from a few individuals of each genotype ( b cn/516
J .
R. True,J. M. Mercer, and<:.
C . 1,aurieD. ntelunognster
[7
D.
sinlrtlansw
D.
mmtritiunaw
I). si/77ttlurc.s/D.
ncnrtritiarm hybridY - l V
v-g
0. I25 0.2
c 0.1
3
'J
n
C .- d
.3 0.15
5
rt:
t 0.075
E
-
'5 0.1.f 0.05
5 5
2 0.025
8
2 0.05n
S 0
.-
-
c
.-
c c
-
>
0
4 p \ I z I W z + P J
0IC' " 2 - 3
= ;
5" . / . " + = = " " ? > . % > S < >
2 2 ; . 5 . % z
s s
2 2= " = "
" "- -
-
.5 a . 2 : : " % *z
2
c
2 3
"
"
-
.-
-
.--
2.
c
"
-
.-"
.- .- + 2.
0.3
1
b-al 0.251
I:
-f
C c 0.2
2 0.2
"
.-
-
0L)
d
.-
-
t
c S
0 3
n n
c c
f.
k 0.15
.-
-
.--
.- 5 0.1
E 0.1 E
5
2 2 0.05
0
'J
0
.
r - ] +P I
4 r.1 :
-
(u 0-
- 2 " - : 2b - * = X '
-
"? s ; . z . z
=, =,- -
-
5 %
" " - * , 5 2 = - -" 2 + S " S % "
-
b U
"
" 2 " "
" " "
" "
-
-
-
.- "- -
-
"
c
.- " .-
.e. 2,
.e.
arm, indicating no fixed differences between the wild- type and double mutant lines (although inversions could be segregating within the wild-type lines, because they are not isochromosomal).
In spite of the considerable intraspecific variability, data from the four homologous intervals support the map length differences indicated by the standard ge- netic maps. The sum of the average recombination frac- tions for each of the four intervals is 0.304 for mdnnogns- to-, 0.459 for simulnns and 0.556 for mnun'tinn,n. T h e ratio of the mnuritinnn sum to the m~lnnognstm sum is
1.8, which is the same as the ratio of total map distance of the standard maps. In addition, for each of the four intervals, the individual recombination fraction esti- mates for mnun'tinnn are each greater than the corre- sponding mPlnnognsfPr estimates. Therefore, it is likely that mnurilinnn characteristically has a substantially longer genetic map than melanogasto-. As in the standard map comparisons, simulnns appears to be intermediate.
FKXXK 5.--Comparison of re- combination fractions for four ho-
mologous i n tends in I). nwlnnogns-
I n ; I). s i m d n n s and I). n m o i t i n n n
and .simltlnn.s/mcrltn'tin~cc/ hybrids.
The "I" and "2" labels refer to dif- ferent wild-type stocks to which the double mutant ?' 7 0 and 1) f n stocks were crossed. Error bars are 9.5%
confidence intends. The "stcl" val-
ues (without error bars) are values
from the standard maps o f n w l m o -
gnstcr or sinculms. A Ipcn value w a s
not available for simctlnns. For the
7+g antl g-Jintenals in mnloitinnn, a
"standard" value was estimated
from the P-hased map by substitut- ing the band index for each marker into the incomplete beta function that describes the relationship be- tween the genetic and physical maps for the X chromosome. A
standard valr~e was not estimated for the y 7 I J antl I x n i n t e n d s in n m t -
n'tinnn because they span terminal
regions of chromosome arms, which are not well described b y the incomplete beta frmction.
DISCUSSION
Recombination in Drosophila Species 517
both in terms of total map length and intensity of the autosomal centromeric effect.
Methodological issues: In discussing the possible causes of the species differences in recombination, we must first consider differences in the methods used in constructing the standard genetic maps that are being compared. The mauritiana map was constructed by esti- mating recombination fractions between a series of P-
element insertions within a fairly uniform genetic back- ground over a short time period. The melanogaster and
simulans maps were constructed using a number of dif-
ferent mutants arising in various stocks over a longer time period. The methods used for estimating map dis- tances from recombination fractions also differ to some extent. However, the following arguments suggest that the standard maps are fairly comparable in spite of these methodological differences.
For both mauritiana and melanogaster, distances in the standard map take multiple crossovers into account be- cause, in essence, they are calculated as the sum of a series of recombination fractions observed for short intervals (within which the probability of double cross- over is negligible). For simulans, the observed recombi- nation fractions are for somewhat longer intervals, but those have been converted to map distance using a mapping function (Foss et al. 1993). Further details are provided in MATERIALS AND METHODS.
The use of P-element insertions as genetic markers in mnuritiana probably has little influence on genetic map characteristics relative to melanogaster because a large fraction of the spontaneous visible mutants used in constructing the melanogaster map also are caused by transposable element insertions (GREEN 1988). In addition, the size of an interval between insertions is generally enormous relative to the size of the transpos- able element marker, so any local effects of these inser- tions on pairing are likely to have a negligible influence on the total crossover frequency within the interval.
The relative genetic homogeneity of the stock used for map construction in mauritiana could bias the map length estimates, if there is a relationship between rate of meiotic recombination and level of heterozygosity (see RADMAN and WAGNER 1993). This relationship in
D.
melanogaster was investigated by RUTHERFORD and CARPENTER (1988) by measuring the frequency of non- exchange X chromosomes (that segregate distributively from aY)
in XXYfemales where the two X i are either isogenic or of different origins. They found no signifi- cant effect of isogenicity, which suggests that using a single mauritiana stock for mapping is probably not an important factor. Furthermore, our analysis of four ho- mologous intervals (Figure 5) indicates that a stock ho- mogeneity effect, if present at all, is likely to be very small. In this experiment, the females from which re- combination fractions are estimated were produced by crosses between mauritiana strains of different origins, yet the map length comparisons with melanogaster are very similar to those involving the standard map dis-tances. Also, the v g and gf distances predicted from the standard mauritiana map are similar to those ob- served in this experiment. Finally, it should also be noted that the simulans/mauritiana hybrids give recom- bination fractions very similar to those of the pure spe- cies, even though they presumably have somewhat higher heterozygosity.
Intra- us. interspecific variation: As noted earlier, many previous studies have shown that natural popula- tions of melanogaster are polymorphic for modifiers of recombination. Here we have provided additional evi- dence of intraspecific variation in all three species, par- ticularly for the y w interval.
The high degree of intraspecific variation suggests that some of the differences among the standard ge- netic maps may be strain-specific rather than species- specific. A proper evaluation of this suggestion will require much more extensive data on intraspecific variation than is currently available for mauritiann or simulans. Genetic map distances estimated for the four homologous intervals (Figure 5) are generally consis- tent with the standard maps in indicating that mauri- tiana has a substantially longer genetic map than mela- nogaster and that simulans is intermediate. However, we have no basis for determining whether the differ- ences in distribution of recombination that are evident from the standard maps (Figures 3 and 4) are species- specific characteristics.
Genetic and mechanistic basis of recombination dif- ferences: There is very little evidence concerning the genetic basis of the differences in recombination in- ferred from the standard genetic maps of the three species, but some possibilities will be mentioned. 1) Differences in recombination frequency may reflect dif- ferences in the physical length of the interval. 2) Trans acting factors, such as those identified as meiotic mu- tants (BAKER et al. 1976) may alter both the level and distribution of recombination throughout the genome.
3) Differences in the sequence and structure of chro- mosomal features, such as the centromere and te- lomere, can cause long-range, polar effects on recombi- nation. 4) Cis- or transacting factors may have restricted regional effects [e.g., mei-1 in D. melanogaster (VALENTIN 1973) or cog sites in Neurospora (CATCHESIDE 1977)].
Three types of evidence argue against the formal pos- sibility that changes in physical distance have caused differences in recombination among the three species. 1) The salivary chromosomes have essentially identical banding patterns except for a few simple rearrange- ments (LEMEUNIER and A~HBURNER 1976). 2) J. SPEN-
CER JOHNSTON (personal communication) has used
518 J. R. True, J. M. Mercer. and C. C. Laurie
two species, even though it has a smaller genetic map. 3) Our own preliminary observations of mitotic chro- mosomes from larval brain cells of mauritiana/melano- gaster hybrid females indicate no obvious differences in chromosome length.
An important question is whether the observed differ- ences in crossover distribution between melanogasterand mauritiana are due to differences in the mechanisms that control crossover positions, or whether they could be explained simply by changes in crossover frequency. An association between changes in crossover frequency and the cumulative distribution of crossover position is expected because of chiasma interference, which affects how multiple crossovers within a bivalent are positioned along the chromosome. Multipoint recombination data for the Xchromosome of melanogasterhave been used to estimate the distribution of exchange points for tetrads with one, two or three crossovers (WEINSTEIN 1936; CHARL.ES 1938; CARPENTER 1988; LANDE and STAHI, 1993). Crossovers in single exchange tetrads (E,s) tend to occur in the medial region of the chromosome arm, while in double exchange tetrads (E2s), one crossover tends to occur proximally and the other distally. In triple exchange tetrads (E3s), the crossovers are evenly spaced along the chromosome: one proximal, one me- dial and one distal. Cumulative distributions of cross- over position, such as those shown in Figures 3 and 4, represent a weighted sum of the distributions for single, double and all higher order tetrad types. Therefore, the cumulative distribution can change simply by alter- ing the relative frequencies of these tetrad types, which is a necessary consequence of changing the total fre- quency of crossing over.
Thus, it appears that an increase in the frequency of crossovers from about one per arm (as in melanogaster) to nearly two per arm (as in mauritiana) could change the shape of the cumulative distribution without neces- sarily changing the mechanisms that determine the po- sitions of crossovers. In particular, increasing the fre- quency of the double crossover class at the expense of the single crossover class is expected to flatten the distribution and thereby diminish regional differences such as the apparent centromere effect. However, a consideration of the differences in crossover distribu- tion between the X and autosomal arms within melano- gaster (discussed below) suggests that additional factors are probably involved i n causing the species differences. Nevertheless, further work using multipoint recombina- tion data is needed to determine whether the species differ in their distribution of exchange points in single crossover tetrads and/or in the pattern of interference shown in multiple crossover tetrads.
The cumulative crossover distributions for melanogas- terin Figures 3 and 4 show that the Xchromosome has much less regional variation in CE than the autosomal arms. This difference is not simply due to a difference in total frequency of exchange, because the X and 3R
show markedly different distributions even though they
have nearly equal genetic lengths. Thus, it seems likely that modal positions of crossovers within each tetrad class differ between Xand 3R. We are not aware of any quantitative comparisons of crossover position among chromosome arms, but there is evidence that the modal position of crossovers in single exchange tetrads is more distal for 3R (and other autosomal arms) than for the X (A. T. C. CARPENTER, personal communication). This difference may be due to the fact that the X chromo- some has much more centric heterochromatin than the autosomes (ASHBURNER 1989, p. 26). An extensive dele- tion analysis provides strong evidence that the X chro- mosome centric heterochromatin buffers the sup- pressing effects of the centromere on recombination in nearby euchromatin (YAMAMOTO and MIKLOS 1978). However, the In(1)sc' rearrangement, which moves a large segment of heterochromatin away from the cen- tromere, has no obvious effect on recombination in the proximal euchromatin, as would be expected from the buffering hypothesis (SLAUTER 1984; CARPENTER 1988). More work is needed to settle this issue.
If centric heterochromatin does play an important role in modulating centromere effects on recombina- tion, then evolutionary changes in crossover distribu- tion may be due to heterochromatic sequence changes. In fact, heterochromatin appears to be a good candi- date for explaining species differences, because it evolves rapidly in many organisms (JOHN and MIKIOS 1979; JOHN 1988). Some differences in heterochro- matic sequences among the three sibling Drosophila species have been observed, particularly between melu- nogaster and simulans. Differences in the staining pat- tern of heterochromatic regions of mitotic chromo- somes (LEMEUNIER et al. 1978) and in satellite sequence composition (LOHE and ROBERTS 1988; CARMENA ~t ab 1993) have been reported, but the functional signifi- cance of these changes (if any) is unknown.
X chromosome nondisjunction: In crosses invoving
X-linked inserts, we observed a frequency of 0.105% patroclinous males (29 out of 27,696) in mauritiana while a compilation of melanogaster results in A H -
BURNER (1989, p. 829) shows a frequency of 0.028%
patroclinous males (140 out of 490,182)