OncoTargets and Therapy
Dove
press
O r i g i n a l r e s e a r c h
open access to scientific and medical research
Open access Full Text article
Four-mirna signature as a prognostic tool
for lung adenocarcinoma
Yan lin1
Yufeng lv1
rong liang1
chunling Yuan1
Jinyan Zhang1
Dan he2
Xiaowen Zheng2
Jianfeng Zhang2
1Department of Medical Oncology,
affiliated Tumor hospital of guangxi Medical University, 2Department of
emergency, The second affiliated hospital of guangxi Medical
University, nanning, People’s republic of china
Purpose: The aim of this study was to generate a novel miRNA expression signature to accurately predict prognosis for patients with lung adenocarcinoma (LUAD).
Patients and methods: Using expression profiles downloaded from The Cancer Genome Atlas database, we identified multiple miRNAs with differential expression between LUAD and paired healthy tissues. We then evaluated the prognostic values of the differentially expressed miRNAs using univariate/multivariate Cox regression analysis. This analysis was ultimately used to construct a four-miRNA signature that effectively predicted patient survival. Finally, we analyzed potential functional roles of the target genes for these four miRNAs using Gene Ontology and Kyoto Encyclopedia of Genes and Genomes pathway enrichment analyses.
Results: Based on our cutoff criteria (P,0.05 and |log2FC| .1.0), we identified a total of 187 differentially expressed miRNAs, including 148 that were upregulated in LUAD tissues and 39 that were downregulated. Four miRNAs (miR-148a-5p, miR-31-5p, miR-548v, and miR-550a-5p) were independently associated with survival based on Kaplan–Meier analysis. We generated a signature index based on the expression of these four miRNAs and stratified patients into low- and high-risk groups. Patients in the high-risk group had significantly shorter survival times than those in the low-risk group (P=0.002). A functional enrichment analysis suggested that the target genes of these four miRNAs were involved in protein phosphorylation and the Hippo and sphingolipid signaling pathways.
Conclusion: Taken together, our results suggest that our four-miRNA signature can be used as a prognostic tool for patients with LUAD.
Keywords: lung adenocarcinoma, miRNA, prognosis
Introduction
Lung cancer is a highly prevalent disease and the most frequently diagnosed cancer. It is also highly fatal, being the most common cause of cancer-related deaths in males
and the second most common cause in females.1 Lung adenocarcinoma (LUAD) is the
most often diagnosed histological subtype of non-small-cell lung cancer (NSCLC).2
Despite the advent of targeted therapies that have improved outcomes for a subset
of patients,3–6 the overall 5-year survival rate for lung cancer patients remains low at
17.4%.7 For patients with NSCLC, the best prognosis is achieved using a complete
surgical resection of stage IA disease, yielding a 5-year survival rate up to 70%. How-ever, this requires early and accurate disease prognosis. Lung cancer can be detected early by a computed tomography scan in high-risk individuals, but this method has
a high false-positive rate that can lead to unnecessary treatment.8 Therefore, more
prognostic indicators are urgently needed.
miRNAs are highly conserved, endogenous noncoding RNAs (20–22 bp in length) that function in a variety of biological processes at the transcriptional or
correspondence: Jianfeng Zhang Department of emergency, The second Affiliated Hospital of Guangxi Medical University, no 166 Daxuedong road, nanning, guangxi 530007, People’s republic of china
Tel +86 771 327 7199 Fax +86 771 327 7285 email drzhangjf@163.com
Journal name: OncoTargets and Therapy Article Designation: Original Research Year: 2018
Volume: 11
Running head verso: Lin et al
Running head recto: Four-miRNA signature for lung adenocarcinoma DOI: http://dx.doi.org/10.2147/OTT.S155016
OncoTargets and Therapy downloaded from https://www.dovepress.com/ by 118.70.13.36 on 25-Aug-2020
For personal use only.
Dovepress
lin et al
posttranscriptional level. Abnormal expression of miRNAs is a common problem in cancer development. Specifically, there is growing evidence for a role of various miRNAs in
the progression and biology of NSCLC.9 Several miRNAs
have been identified as markers in cancers, including a
four-miRNA signature in colon cancer10 and a three-miRNA
signature in cervical cancer.11 Therefore, miRNAs are
promising indicators for cancer diagnosis, prognosis, and targeted therapies.
The Cancer Genome Atlas Project (TCGA) is a National Cancer Institute effort to profile 29 different tumor types using genomic platforms and to make raw and processed data
readily available to all researchers.12 The TCGA has released
massive datasets of miRNA sequencing data from patients with LUAD. For this study, we downloaded high-throughput miRNA data from the TCGA database to identify several miRNAs with differential expression between LUAD and paired healthy lung tissues. We then evaluated the prognostic value of these differentially expressed miRNAs and found a four-miRNA signature that could effectively predict patient survival. Furthermore, we analyzed the pathways and func-tions of the target genes modulated by these four miRNAs. Ultimately, we hope that this signature can be used as a novel prognostic tool for LUAD, and that our functional analysis may provide more insights into the molecular mechanisms of this prevalent and devastating disease.
Patients and methods
Data processing
The preprocessed mature miRNA expression profiles, dis-played as reads per million, and clinical information from the TCGA (TCGA data version 2016_01_28 for LUAD) were
downloaded from Broad Institute GDAC Firebrowse (http://
firebrowse.org/). The inclusion criteria were as follows: 1) the sample included both miRNA expression profiles and clinical information, and 2) the sample included prognosis information.
generation of a prognostic signature
for lUaD using differentially expressed
mirnas
To create a LUAD prognostic signature, we first removed
miRNAs with expression levels of zero in .50% of the
patients. We then normalized the miRNA expression profiles by log2 transformation using the R language project. We identified the miRNAs that were differentially expressed between LUAD and paired healthy tissues using the “limma” package in R. Finally, we calculated fold changes in the
expression of individual miRNAs between LUAD and
healthy tissues and considered miRNAs with a |log2FC| .1
and a P-value ,0.05 to be significant.
The four-mirna signature index and
survival
In our univariate Cox proportional hazards regression analysis, we selected the median expression level for each miRNA as the cutoff point to dichotomize patients into high- and low-expression groups. To determine if the expression level was associated with a prognostic outcome, we applied a Kaplan–Meier survival analysis with the log-rank method. We then confirmed the prognostic value of the four statistically significant miRNAs using a multivariate Cox proportional hazards regression analysis. For each of these four miRNAs, we assigned each patient one point for the high-risk expression level. Thus, each patient received a score ranging from 0 to 4, which we called the signature index. We plotted a survival curve based on the patients’ survival status, survival days, and signature index using the Kaplan–Meier survival method (log-rank). We considered patients with a signature index of 3 or 4 to be high risk and
those ,3 to be low risk.
Prediction of the target genes of the
prognostic mirna signature
We predicted the target genes of the four prognostic miRNAs
using the TargetScan (http://www.targetscan.org/), miRDB
(http://www.mirdb.org/miRDB/), and miRTarBase (http://
mirtarbase.mbc.nctu.edu.tw/index.php) online analysis tools.
To further enhance the reliability of the bioinformatic analy-sis, we created a Venn diagram for each miRNA to identify the overlapping results among the three prediction tools. We performed Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses of the overlapping target genes using the “clusterProfiler”
package in R.13 The clusterProfiler package implements
methods to analyze and visualize the functional profiles of genes and gene clusters. We set the cutoff criteria as a
P-value ,0.05 and a gene count $2.
statistical analysis
We analyzed the expression levels of the miRNAs in LUAD and matched healthy tissues using unpaired t-tests. We performed Kaplan–Meier survival analysis and univariate/ multivariate Cox proportional hazards regression analysis to compare the expression levels (low vs high) and prog-nostic significance (low risk vs high risk) of each miRNA.
OncoTargets and Therapy downloaded from https://www.dovepress.com/ by 118.70.13.36 on 25-Aug-2020
Dovepress Four-mirna signature for lung adenocarcinoma
We considered P-values ,0.05 to be statistically significant.
Statistical analysis was performed using IBM SPSS Statis-tics software program version 22.0 (IBM Corp., Armonk, NY, USA).
Results
Differentially expressed mirnas in
lUaD and healthy tissues
Our analysis included a total of 438 LUAD tissues and 45 matched healthy tissues from the same patients. The detailed clinical characteristics including age at diagnosis, metastasis, lymph node status, pathological stage, and T stage are
listed in Table 1. According to the cutoff criteria (P,0.05
and |log2FC| .1), 187 miRNAs were differentially expressed
between LUAD and matched healthy tissues. This included 148 miRNAs that were upregulated in the LUAD tissues and 39 miRNAs that were downregulated. The results of the expression analysis are presented as a volcano plot (Figure 1) to demonstrate that the distribution of P-values and |log2FC| was reasonable with respect to each other.
Identification of four miRNAs associated
with survival in lUaD
For each of the 187 differentially expressed miRNAs, we used the median expression level as a cutoff to stratify the 438
patients into two groups: a high-level group and a low-level group. The univariate Cox proportional hazards regression analysis revealed that a total of 22 miRNAs had prognostic value. We subsequently applied a multivariate Cox pro-portional hazards regression analysis to verify these results and identified four miRNAs, miR-148a-5p (Figure 2A), 31-5p (Figure 2B), 548v (Figure 2C), and miR-550a-5p (Figure 2D), as independent prognostic indicators. The results of the univariate and multivariate analyses for the 22 miRNAs are displayed in Table 2. All four miRNAs were upregulated in the LUAD tissues. Patients with high expres-sion levels of miR-148a-5p, miR-31-5p, and miR-1548v had longer survival times compared with patients with low expression levels of those miRNAs. In contrast, patients with high expression levels of miR-550a-5p had shorter survival times than those with low expression levels.
a four-mirna signature index for lUaD
prognosis
We scored the four-miRNA signature by value assignment for each miRNA (Table 3). Each patient was assigned one score for a low expression level of miR-148a-5p, miR-31-5p, and miR-548v, respectively, and one score for a high expression
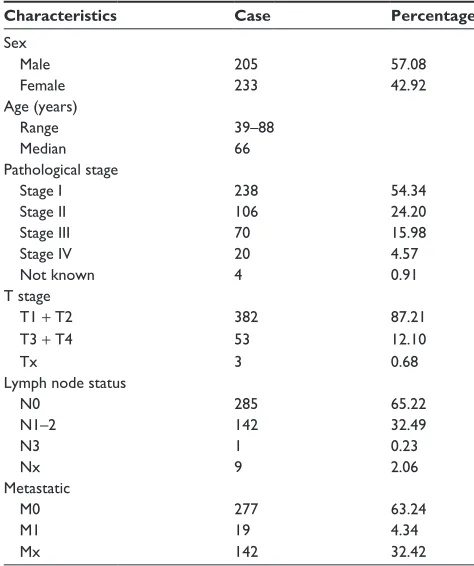
Table 1 summary of patient cohort information
Characteristics Case Percentage
sex
Male 205 57.08
Female 233 42.92
age (years)
range 39–88
Median 66
Pathological stage
stage i 238 54.34
stage ii 106 24.20
stage iii 70 15.98
stage iV 20 4.57
not known 4 0.91
T stage
T1 + T2 382 87.21
T3 + T4 53 12.10
Tx 3 0.68
lymph node status
n0 285 65.22
n1–2 142 32.49
n3 1 0.23
nx 9 2.06
Metastatic
M0 277 63.24
M1 19 4.34
Mx 142 32.42
± ±
ORJ)&
íORJDGMXVWHG
3
YDOXH
Figure 1 Volcano plot of the differentially expressed mirnas between lUaD and
paired healthy tissues.
Notes: red indicates high expression in lUaD tissues, while blue indicates low
expression. gray shows the mirnas with a |log2Fc| ,1 and/or an adjusted
P-value $0.05. The volcano plot was constructed using the ggplot2 package of the r language.
Abbreviation: lUaD, lung adenocarcinoma.
OncoTargets and Therapy downloaded from https://www.dovepress.com/ by 118.70.13.36 on 25-Aug-2020
Dovepress
lin et al
level of miR-550a-5p. Then, we summed the scores for each patient, resulting in a signature index ranging from 0 to 4. All patients were divided into high- or low-risk groups accord-ing to this index. Specifically, we considered patients with a signature index of 3 or 4 to be high risk and those with an
index ,3 to be low risk. We compared the two groups in a
Kaplan–Meier survival analysis using the log-rank method.
This revealed that patients in the high-risk group (n=123)
had significantly shorter survival times than those in the
low-risk group (n=315) (P=0.002; Figure 3). This indicates
that our four-miRNA signature created was associated with survival in LUAD.
Target prediction and functional analysis
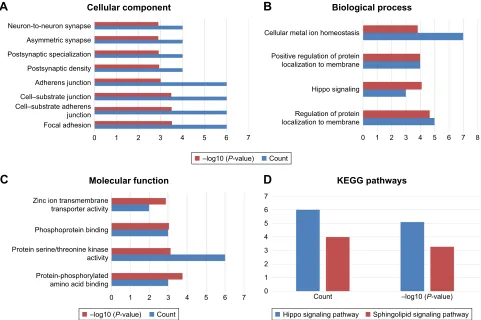
To explore the potential biological functions of the four miRNAs (miR-148a-5p, miR-31-5p, miR-548v, andmiR-550a-5p), we predicted their respective target genes using the TargetScan, miRDB, and miRTarBase online analysis tools. Each analysis tool identified unique target genes for the four miRNAs, but all three tools identified 12 overlapping target genes for miR-148a-5p, 17 for miR-31-5p, nine for miR-548v, and 6 for miR-550a-5p (Figure 4A–D). We analyzed the 44 potential target genes using the KEGG signal pathway and GO databases. Those 44 potential target genes are shown in Table 4. The GO annotation results have three parts: a cellular component, a biological process, and a molecular function. The results of the significantly enriched
GO term analysis are shown in Figure 5A–C (P,0.05, gene
count $2). This analysis revealed that the molecular
func-tions primarily associated with these genes were protein serine/threonine kinase activity and protein-phosphorylated amino acid binding. The KEGG pathways identified in the
$
PL5DS
3
7LPHGD\V
)UDFWLRQVXUYLYDO
3
PL5S
)UDFWLRQVXUYLYDO
7LPHGD\V
%
3 PL5Y
7LPHGD\V
)UDFWLRQVXUYLYDO
&
3
PL5DS
7LPHGD\V
)UDFWLRQVXUYLYDO
'
+LJKOHYHO /RZOHYHO
Figure 2 Four mirnas were associated with overall survival in lUaD patients using Kaplan–Meier curves and log-rank tests.
Notes: The patients were stratified into high- and low-risk groups according to the median expression of each miRNA. Each plot represents the Kaplan–Meir curves obtained
using a different mirna: (A) mir-148a-5p, (B) mir-31-5p, (C) mir-548v, and (D) mir-550a-5p. Abbreviation: lUaD, lung adenocarcinoma.
OncoTargets and Therapy downloaded from https://www.dovepress.com/ by 118.70.13.36 on 25-Aug-2020
Dovepress Four-mirna signature for lung adenocarcinoma
analysis were significantly enriched with the Hippo and
sphingolipid signaling pathways (P,0.05, gene count $2)
(Figure 5D). Together, these results suggest that the differen-tially expressed miRNAs may modulate protein phosphoryla-tion, particularly in the identified signaling pathways.
Discussion
LUAD is a devastating cancer subtype with a low survival rate. Despite its prevalence, prognostic tools for LUAD are limited. The prognosis of patients with LUAD would be
improved considerably if tumor behavior could be reliably predicted at the time of the initial diagnosis. This requires the identification of novel biomarkers and an understanding of the molecular mechanisms underlying LUAD development. In this paper, we identified a total of 187 miRNAs that were differentially expressed between LUAD and matched healthy tissues, 4 of which were associated with overall survival in patients with LUAD. Because miRNAs are known to modu-late gene expression, we screened the target genes of those four miRNAs and used bioinformatics to predict the pathways and biological functions associated with their targets.
Several miRNA markers have been identified for the prediction and prognosis of cancers such as head and neck
squamous cell carcinomas,14 bile duct cancer,15 and glioma.16
In this paper, we sought to identify miRNA markers associ-ated with prognosis for LUAD. We creassoci-ated a four-miRNA signature index that successfully separated patients into low- and high-risk groups. Specifically, patients deemed high risk by our four-miRNA signature index had significantly shorter
survival times than those in the low-risk group (P=0.002).
We additionally performed a receiver operating characteristic (ROC) curve study after multivariate analysis, but were unable
to achieve the suggested area under the curve (AUC) .0.7
(data not shown). However, this discrepancy is similar to the results of previous studies. Several studies have investigated the relationship between miRNA expression patterns and the survival and prognosis of patients with LUAD. Li et al reported that miR-101-1, miR-220a, miR-4661, and miR-450a-2 were
associated with overall survival in patients with LUAD.17
Table 2 Univariate and multivariate analyses in lUaD patients
miRNA Univariate analysis Multivariate analysis
HR (95% CI) P-value HR (95% CI) P-value
let-7c-5p 0.512–0.966 0.029 0.639–1.395 0.772
mir-148a-3p 1.053–1.993 0.022 0.889–2.135 0.151 mir-148a-5p 1.052–1.987 0.023 0.399–0.931 0.022* mir-161-3p 1.204–2.277 0.002 0.606–1.363 0.643 mir-199b-5p 1.036–1.970 0.028 0.593–1.414 0.690 mir-200a-5p 1.119–2.116 0.008 0.582–1.261 0.432
mir-21-5p 0.467–0.889 0.007 0.966–2.136 0.074
mir-29b-1-5p 1.070–2.023 0.017 0.535–1.277 0.390 mir-29b-3p 1.111–2.106 0.009 0.590–1.454 0.738
mir-31-5p 0.495–0.945 0.020 1.132–2.417 0.009*
mir-3200-3p 1.018–1.925 0.037 0.630–1.333 0.648 mir-3607-3p 1.167–2.218 0.003 0.496–1.197 0.245
mir-375 1.045–1.979 0.025 0.651–1.367 0.757
mir-423-5p 0.486–0.934 0.016 0.936–1.972 0.107 mir-516a-5p 0.520–0.981 0.037 0.973–1.931 0.071
mir-548v 1.207–2.294 0.002 0.446–0.946 0.024*
mir-550a-5p 0.517–0.997 0.035 1.114–2.394 0.012* mir-551b-3p 1.025–1.939 0.034 0.618–1.434 0.779 mir-628-5p 1.020–1.923 0.037 0.616–1.258 0.483 mir-660-5p 1.058–2.011 0.020 0.533–1.184 0.259
mir-9-5p 0.527–0.995 0.046 0.793–1.664 0.463
mir-940 1.117–2.125 0.008 0.500–1.074 0.111
Note: *P value of ,0.05 was considered statistically significant difference.
Abbreviations: LUAD, lung adenocarcinoma; HR, hazard ratio; CI, confidence
interval.
Table 3 Patient scoring method using the four-mirna signature
index
miRNA Median
(RPM)
log2 (median)
Expression level
Scorea
mir-148a-5p 56.96 5.832 .Median 0
#Median 1
mir-31-5p 2.093 1.066 .Median 0
#Median 1
mir-548v 1.298 0.376 .Median 0
#Median 1
mir-550a-5p 3.350 1.744 .Median 1
#Median 0
Note: aa score of 3 or 4 indicates the patients to be high risk, while a score ,3 indicates low risk.
Abbreviation: rPM, reads per million.
3
7LPHGD\V
)UDFWLRQVXUYLYDO
+LJKULVNJURXS /RZULVNJURXS
Figure 3 The Kaplan–Meier curves obtained using the four-mirna signature to
separate patients into high- and low-risk groups.
OncoTargets and Therapy downloaded from https://www.dovepress.com/ by 118.70.13.36 on 25-Aug-2020
Dovepress
lin et al
This prognosis model had significant efficacy for Caucasian patients, but as in the current study, their model did not reach
an AUC of 0.7 (AUC =0.629). This study also did not show
their sensitivity and specificity, so it is difficult to fully evaluate the efficacy of their model. Alternatively, Peng et al performed a comprehensive analysis based on LUAD miRNome profiling studies using the robust rank aggregation method. They identified a panel of six miRNAs (miR-21-5p, miR-210-3p, miR-182-5p, miR-183-5p, miR-126-3p, and miR-218-5p) associated with LUAD and evaluated the expression and
prognostic values of those miRNAs in an independent cohort.18
They did not perform a ROC curve study. Sathipati and Ho
proposed an optimized support vector regression method to identify an miRNA signature for estimating LUAD patients survival using their miRNA expression profiles. Their miRNA signature consists of 18 miRNAs associated with LUAD: hsa-let-7f-1, hsa-miR-16-1, hsa-miR-152, hsa-miR-217, miR-18a, has-miR-193b, miR-3136, let-7g, hsa-miR-155, hsa-miR-3199-1, hsa-miR-219-2, hsa-miR-1254, 1291, 192, 3653,
hsa-miR-3934, hsa-miR-342, and hsa-miR-141.19 In our study, we
determined that increased expression levels of four mature miRNAs (148a-5p, 31-5p, 548v, and miR-550a-5p) were associated with clinical outcomes in patients with LUAD. Interestingly, the miRNA signatures generated by these various studies do not overlap, perhaps because of variability in the methods used to identify miRNAs. And, no study can build a predictive tool to achieve the desired AUC. Therefore, predicting the prognosis of LUAD with miRNA signature still needs more in-depth research. How-ever, our study and the previous studies all emphasize new and constructive strategies to identify prognostic markers for LUAD. One strength of our study is that we removed miRNAs
with expression levels of zero in .50% of the patients,
which would enhance the feasibility of implementation.
7DUJHWJHQHVRIPL5DS
7DUJHW6FDQ
PL57DU%DVH PL5'%
7DUJHWJHQHVRIPL5Y
7DUJHW6FDQ
PL57DU%DVH PL5'%
7DUJHWJHQHVRIPL5S
7DUJHW6FDQ
PL57DU%DVH PL5'%
7DUJHW6FDQ
7DUJHWJHQHVRIPL5DS
PL57DU%DVH PL5'%
$
%
&
'
Figure 4 The overlapping target genes of the respective mirnas were predicted using the mirDB, Targetscan, and mirTarBase online analysis tools: (A) mir-148a-5p,
(B) mir-31-5p, (C) mir-548v, and (D) mir-550a-5p.
Table 4 The overlapping potential target genes of the four
mirnas
miRNA Overlapping target genes
mir-148a-5p PDPK1, RBMXL1, AFF2, UBR3, CDCA4, RBM24, TPST2, STK17A, MOB3B, SLC30A7, MYC, ID4
mir-31-5p PPP2R2A, LATS2, NUMB, KLF13, JAZF1, FZD3, HIF1AN, PRKCE, RASA1, STK40, ZC3H12C, TBXA2R, ARID1A, YWHAE, RHOBTB1, SYDE2, CCNT1
mir-548v SGPL1, CDK5R1, SCD, TCF12, NCK2, PPIA, CSTF2T, SLC39A6, SLC10A7
mir-550a-5p CDCA7, SEMA4C, MKI67, STX16, LIMD1, TMEM151B
OncoTargets and Therapy downloaded from https://www.dovepress.com/ by 118.70.13.36 on 25-Aug-2020
Dovepress Four-mirna signature for lung adenocarcinoma
To gain insight into the molecular functions of the four miRNAs, we predicted the target genes for the four miRNAs and identified their associated KEGG pathways and GO annotations. The molecular functions of the target genes were mainly associated with protein serine/threonine kinase activity and protein-phosphorylated amino acid binding. Thus, the four miRNAs might be closely related to protein phosphorylation. The target genes were also signifi-cantly enriched for the Hippo signaling pathway (six genes,
P=7.87×10−6). Abnormal regulation of the Hippo signaling
pathway is involved in the development of a variety of human cancers such as breast cancer, hepatocellular carcinoma,
hematological cancer, and lung cancer.20 Thus, abnormal
regulation of signaling pathways, including Hippo, may play a crucial role in the pathogenesis and progression of LUAD. The interaction between the four miRNAs and the Hippo pathway merits further exploration, as more research is necessary to determine the exact molecular mechanisms of the role of miRNAs in LUAD.
Our study has a few limitations. Although we included a relatively large sample size of 438 patients, we used
the median expression level as the cutoff value to stratify patients. Despite the ability of our model to successfully stratify patients into high- and low-risk groups, it did not yield the suggested AUC. However, this was similar to the results of previous studies examining the relationship between miRNA expression and LUAD prognosis. Thus, the prognostic value of the four-miRNA signature needs further validation with other methods and a larger sample size. Further functional investigation is also required to explore the molecular functions of the four miRNAs in LUAD progression.
Conclusion
Taken together, our results identified a four-miRNA signa-ture that could be a prognostic tool for patients with LUAD.
The four miRNAs were expressed in .50% of patients with
LUAD and were more highly expressed in LUAD tissues than in matched healthy ones. These miRNAs modulated genes associated with protein phosphorylation and the Hippo sig-naling pathway, which has previously been associated with cancer progression. Ultimately, we hope that this miRNA
0ROHFXODUIXQFWLRQ
=LQFLRQWUDQVPHPEUDQH WUDQVSRUWHUDFWLYLW\
3KRVSKRSURWHLQELQGLQJ
3URWHLQVHULQHWKUHRQLQHNLQDVH DFWLYLW\
3URWHLQSKRVSKRU\ODWHG DPLQRDFLGELQGLQJ
.(**SDWKZD\V
&RXQW ±ORJ3YDOXH
±ORJ3YDOXH &RXQW
±ORJ3YDOXH &RXQW
+LSSRVLJQDOLQJSDWKZD\ 6SKLQJROLSLGVLJQDOLQJSDWKZD\
&HOOXODUFRPSRQHQW %LRORJLFDOSURFHVV
1HXURQWRQHXURQV\QDSVH $V\PPHWULFV\QDSVH 3RVWV\QDSWLFVSHFLDOL]DWLRQ 3RVWV\QDSWLFGHQVLW\ $GKHUHQVMXQFWLRQ &HOO±VXEVWUDWHMXQFWLRQ &HOO±VXEVWUDWHDGKHUHQV MXQFWLRQ )RFDODGKHVLRQ
&HOOXODUPHWDOLRQKRPHRVWDVLV
3RVLWLYHUHJXODWLRQRISURWHLQ ORFDOL]DWLRQWRPHPEUDQH
+LSSRVLJQDOLQJ
5HJXODWLRQRISURWHLQ ORFDOL]DWLRQWRPHPEUDQH
$
%
&
'
Figure 5 The significantly enriched Gene Ontology annotation of the target genes of the four miRNAs including their (A) cellular component, (B) biological process, and
(C) molecular function. (D) Kyoto encyclopedia of genes and genomes pathway analysis of the predicted targets of the four mirnas.
OncoTargets and Therapy downloaded from https://www.dovepress.com/ by 118.70.13.36 on 25-Aug-2020
OncoTargets and Therapy
Publish your work in this journal
Submit your manuscript here: http://www.dovepress.com/oncotargets-and-therapy-journal
OncoTargets and Therapy is an international, peer-reviewed, open access journal focusing on the pathological basis of all cancers, potential targets for therapy and treatment protocols employed to improve the management of cancer patients. The journal also focuses on the impact of management programs and new therapeutic agents and protocols on
patient perspectives such as quality of life, adherence and satisfaction. The manuscript management system is completely online and includes a very quick and fair peer-review system, which is all easy to use. Visit http://www.dovepress.com/testimonials.php to read real quotes from published authors.
Dovepress
Dove
press
lin et al
signature can help to predict LUAD prognosis and uncover the mechanisms underlying LUAD development.
Acknowledgments
This study was supported by the National Natural Science Foundation of China (Grant No 81360290), the Guangxi Natural Science Foundation (Grant Nos 2017GXNS-FAA198249, 2016GXNS FBA380090 and 2015GXNS-FAA139128), the Key Research and Development project of Guangxi (Grant No Guike AB17195002), the Basic Ability Enhancement Program for Young and Middle-age Teach-ers of Guangxi (Grant No 2017KY0120), the Self-raised Scientific Research Funds of Ministry of Health of Guangxi Province (Grant No Z2016480) and the China Scholarship Council (201608455001).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65(2):87–108. 2. Travis WD. Pathology of lung cancer. Clin Chest Med. 2011;32(4):
669–692.
3. Zhou C, Wu Y-L, Chen G, et al. Erlotinib versus chemotherapy as first-line treatment for patients with advanced EGFR mutation-positive non-small-cell lung cancer (OPTIMAL, CTONG-0802): a multicentre, open-label, randomised, phase 3 study. Lancet Oncol. 2011;12(8):735–742. 4. Maemondo M, Inoue A, Kobayashi K, et al. Gefitinib or chemotherapy
for non-small-cell lung cancer with mutated EGFR. N Engl J Med. 2010;362(25):2380–2388.
5. Paez JG, Jänne PA, Lee JC, et al. EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science. 2004; 304(5676):1497–1500.
6. Kwak EL, Bang Y-J, Camidge DR, et al. Anaplastic lymphoma kinase inhibition in non-small-cell lung cancer. N Engl J Med. 2010;363(18): 1693–1703.
7. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer
J Clin. 2016;66(1):7–30.
8. Aberle DR, Abtin F, Brown K. Computed tomography screening for lung cancer: has it finally arrived? Implications of the National Lung Screening Trial. J Clin Oncol. 2013;31(8):1002–1008.
9. Boeri M, Pastorino U, Sozzi G. Role of microRNAs in lung cancer.
Cancer J. 2012;18(3):268–274.
10. Xu J, Zhao J, Zhang R. Four microRNAs signature for survival prog-nosis in colon cancer using TCGA data. Sci Rep. 2016;6:38306. 11. Liang B, Li Y, Wang T. A three miRNAs signature predicts survival
in cervical cancer using bioinformatics analysis. Sci Rep. 2017; 7(1):1–8.
12. Chandran UR, Medvedeva OP, Barmada MM, et al. TCGA expedition: a data acquisition and management system for TCGA data. PLoS One. 2016;11(10):e0165395.
13. Yu G, Wang L-G, Han Y, He Q-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012; 16(5):284–287.
14. Wong N, Khwaja SS, Baker CM, et al. Prognostic microRNA signatures derived from The Cancer Genome Atlas for head and neck squamous cell carcinomas. Cancer Med. 2016;5(7):1619–1628.
15. Wang M, Wen T-F, He L-H, Li C, Zhu W-J, Trishul NM. A six-microRNA set as prognostic indicators for bile duct cancer. Int J Clin
Exp Med. 2015;8(10):17261–17270.
16. Yan W, Li R, Liu Y, et al. MicroRNA expression patterns in the malig-nant progression of gliomas and a 5-microRNA signature for prognosis.
Oncotarget. 2014;5(24):12908–12915.
17. Li X, An Z, Li P, Liu H. A prognostic model for lung adenocarci-noma patient survival with a focus on four miRNAs. Oncol Lett. 2017;14(3):2991–2995.
18. Peng Z, Pan L, Niu Z, et al. Identification of microRNAs as potential biomarkers for lung adenocarcinoma using integrating genomics analy-sis. Oncotarget. 2017;8(38):64143–64156.
19. Sathipati SY, Ho S-Y. Identifying the miRNA signature associated with survival time in patients with lung adenocarcinoma using miRNA expression profiles. Sci Rep. 2017;7(1):7507.
20. Yu F-X, Zhao B, Guan K-L. Hippo pathway in organ size control, tissue homeostasis, and cancer. Cell. 2015;163(4):811–828.
OncoTargets and Therapy downloaded from https://www.dovepress.com/ by 118.70.13.36 on 25-Aug-2020