organic papers
o3176
Meng and Wu C12H18O4 doi:10.1107/S1600536805027790 Acta Cryst.(2005). E61, o3176–o3178 Acta Crystallographica Section E
Structure Reports
Online
ISSN 1600-5368
(
RS
)-2,6-Dimethoxy-4-(2-methoxypropyl)phenol
Xiang-Gao Meng* and An-Xin Wu*
Department of Chemistry, Central China Normal University, Wuhan 430079, People’s Republic of China
Correspondence e-mail: mengxianggao@mail.ccnu.edu.cn, chwuax@mail.ccnu.edu.cn
Key indicators
Single-crystal X-ray study
T= 292 K
Mean(C–C) = 0.007 A˚
Rfactor = 0.057
wRfactor = 0.164 Data-to-parameter ratio = 8.3
For details of how these key indicators were automatically derived from the article, see http://journals.iucr.org/e.
#2005 International Union of Crystallography Printed in Great Britain – all rights reserved
The title compound, C12H18O4, crystallizes with two molecules
in the asymmetric unit. The structure features O—H O hydrogen-bonded double chains.
Comment
The mechamism of phenol oxidation by tyrosinase has been well studied (Krol & Bolton, 1997). However, little work has been done to determine the influence of substituents on the reaction. Recently, we have synthesized the title compound, (I), and its crystal structure is reported here.
Compound (I) crystallizes in the non-centrosymmetric space group Pc, with two independent molecules in the asymmetric unit. In the molecular structure (Fig. 1 and Table 1), the two molecules have similar geometric parameters about the two chiral centres, C10 and C22; the bond lengths and angles are unremarkable. The crystal structure is stabil-ized by intermolecular O—H O hydrogen bonds, with the phenol groups acting as donors and the ether O atoms as acceptors (Table 2). These hydrogen bonds link the molecules into double chains (Fig. 2).
Experimental
2,6-Dimethoxyphenol, (1), was synthesized according to the literature procedure of Wu et al.(1997). To a basic acetone solution of 2,6-dimethoxyphenol (0.35 g, 2 mmol) was added K2CO3 (0.14 g); a solution of 3-bromoprop-1-ene (15 ml) in acetone was then added dropwise. The mixture was stirred at 308 K for 5 h, filtered and
[image:1.610.209.460.299.516.2]purified by column chromatography on silical gel, using petroleum– ethyl acetate (3:2 v/v), to give compound (2) (0.30 g, 1.5 mmol). Product (2) (0.19 g, 1 mmol) in tetrahydrofuran (THF, 20 ml) was refluxed for 1 h at 393 K and then cooled to room temperature. To the solution was added hydrobromic acid (5 ml, 6M) and the mixture stirred for 1 h. Removal of the solvent under reduced pressure and subsequent purification by flash chromatography (using acetone as eluent) afforded compound (4) (0.15 g, 0.5 mmol). The hydroxyl group of compound (4) was protected with acetic andydride, yielding compound (5). To a solution of (5) in THF (20 ml) was added sodium (0.07 g) and the mixture was refluxed for 12 h at 323 K to afford the title compound, (I). Crystals suitable for X-ray analysis were obtained by recrystallization from ethanol.
Crystal data
C12H18O4 Mr= 226.26
Monoclinic,Pc a= 13.740 (3) A˚
b= 9.405 (2) A˚
c= 9.997 (2) A˚
= 102.190 (4)
V= 1262.8 (5) A˚3 Z= 4
Dx= 1.190 Mg m 3
MoKradiation Cell parameters from 2536
reflections
= 2.6–25.4
= 0.09 mm1 T= 292 (2) K Block, colorless 0.300.200.10 mm
Data collection
Bruker SMART APEX CCD area-detector diffractometer
!scans
Absorption correction: none 6654 measured reflections 2481 independent reflections
1809 reflections withI> 2(I)
Rint= 0.081 max= 26.0 h=16!16
k=11!6
l=12!11
Refinement
Refinement onF2 R[F2> 2(F2)] = 0.057 wR(F2) = 0.164 S= 0.97 2481 reflections 299 parameters
H-atom parameters constrained
w= 1/[2(F
o2) + (0.1154P)2]
whereP= (Fo2+ 2Fc2)/3
(/)max< 0.001 max= 0.37 e A˚
3 min=0.19 e A˚
3
Table 1
Selected geometric parameters (A˚ ,).
C9—C10 1.502 (6) C10—O4 1.437 (5) C10—C11 1.491 (7)
C21—C22 1.501 (8) C22—O8 1.431 (6) C22—C23 1.555 (10) O4—C10—C11 111.5 (4)
O4—C10—C9 105.4 (3) C11—C10—C9 113.1 (4)
O8—C22—C21 105.3 (4) O8—C22—C23 112.4 (5) C21—C22—C23 111.8 (5)
Table 2
Hydrogen-bond geometry (A˚ ,).
D—H A D—H H A D A D—H A
O2—H2 O1 0.82 2.26 2.676 (4) 112 O2—H2 O8i
0.82 2.05 2.774 (5) 148 O6—H6A O5 0.82 2.25 2.661 (4) 111 O6—H6A O4ii
0.82 2.05 2.823 (4) 156
Symmetry codes: (i)xþ1;y;z; (ii)x;y1;zþ1.
H atoms were positioned geometrically (methyl C—H = 0.96 A˚ , methylene C—H = 0.97 A˚ , methine C—H = 0.98 A˚, aromatic C—H = 0.93 A˚ and hydroxy O—H = 0.82 A˚) and included in the refinement in the riding-model approximation, with Uiso(H) = xUeq(carrier
atom), wherex= 1.5 for methyl and hydroxy, andx = 1.2 for the others. In the absence of significant anomalous dispersion effects, Friedel pairs were averaged.
Data collection:SMART(Bruker, 2001); cell refinement: SAINT-Plus(Bruker, 2001); data reduction:SAINT-Plus; program(s) used to solve structure: SHELXS97(Sheldrick, 1997); program(s) used to refine structure:SHELXL97(Sheldrick, 1997); molecular graphics:
PLATON (Spek, 2003); software used to prepare material for publication:PLATON.
The authors are grateful to the Central China Normal University, the National Natural Science Foundation of China (grant No. 20472022), and the Hubei Province Natural Science Fund (grant Nos. 2004ABA085 and 2004ABC002) for finan-cial support.
References
Bruker (2001). SAINT-Plus (Version 6.45) and SMART (Version 5.628). Bruker AXS Inc., Madison, Wisconsin, USA.
organic papers
Acta Cryst.(2005). E61, o3176–o3178 Meng and Wu C
[image:2.610.318.567.70.237.2]12H18O4
o3177
Figure 1
[image:2.610.311.564.278.463.2]The molecular structure of the asymmetric unit of the title compound, showing 30% probablity displacement ellipsoids.
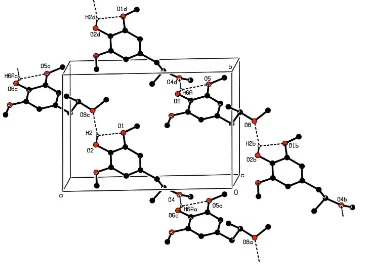
Figure 2
Krol, E. S. & Bolton, J. L. (1997).Chem. Biol. Interact.104, 11–27. Sheldrick, G. M. (1997). SHELXS97 and SHELXL97. University of
Go¨ttingen, Germany.
Spek, A. L. (2003).J. Appl. Cryst.36, 7–13.
Wu, A. X., Wang, M. Y. & Pan, X. F. (1997).Hua Xue Tong Bao,8, 42–43. (In Chinese.)
organic papers
o3178
Meng and Wu Csupporting information
sup-1
Acta Cryst. (2005). E61, o3176–o3178supporting information
Acta Cryst. (2005). E61, o3176–o3178 [doi:10.1107/S1600536805027790]
(
RS
)-2,6-Dimethoxy-4-(2-methoxypropyl)phenol
Xiang-Gao Meng and An-Xin Wu
S1. Comment
The mechamism of phenol oxidation by tyrosinase has been well studied (Krol & Bolton, 1997). However, little work has
been done to determine the influence of substituents on the reaction. Recently, we have synthesized the title compound,
(I), and its crystal structure is reported here.
Compound (I) crystallizes in the non-centrosymmetric space group Pc, with two independent molecules in the
asymmetric unit. In the molecular structure (Fig. 1 and Table 1), the two molecules have similar geometric parameters
about the two chiral centres, C10 and C22; the bond lengths and angles are unremarkable. The crystal structure is
stabilized by intermolecular O—H···O hydrogen bonds, with the phenol groups acting as donors and the ether O atoms as
acceptors (Table 2). These hydrogen bonds link the molecules into double chains (Fig. 2).
S2. Experimental
2,6-Dimethoxyphenol, (1), was synthesized according to the literature procedure of Wu et al. (1997). To a basic acetone
solution of 2,6-dimethoxyphenol (0.35 g, 2 mmol) was added K2CO3 (0.14 g); a solution of 3-bromoprop-1-ene (15 ml)
in acetone was then added dropwise. The mixture was stirred at 308 K for 5 h, filtered and purified by column
chromatography on silical gel, using petroleum–ethyl acetate (3:2 (v/v), to give compound (2) (0.30 g, 1.5 mmol).
Product (2) (0.19 g, 1 mmol) in tetrahydrofuran (THF, 20 ml) was refluxed for 1 h at 393 K and then cooled to room
temperature. To the solution was added hydrobromic acid (5 ml, 6 M) and the mixture stirred for 1 h. Removal of the
solvent under reduced pressure and subsequent purification by flash chromatography (using acetone as eluent) afforded
compound (4) (0.15 g, 0.5 mmol). The hydroxyl group of compound (4) was protected with acetic andydride, yielding
compound (5). To a solution of (5) in THF (20 ml) was added sodium (0.07 g) and the mixture was refluxed for 12 h at
323 K to afford the title compound, (I). Crystals suitable for X-ray analysis were obtained by recrystallization from
ethanol.
S3. Refinement
H atoms were positioned geometrically (methyl C—H = 0.96 Å, methylene C—H = 0.97 Å, methine C—H = 0.98 Å,
aromatic C—H = 0.93 Å and hydroxy O—H = 0.82 Å) and included in the refinement in the riding-model approximation,
with Uiso(H) = xUeq(carrier atom), where x = 1.5 for methyl and hydroxy, and x = 1.2 for the others. In the absence of
supporting information
[image:5.610.125.487.69.310.2]sup-2
Acta Cryst. (2005). E61, o3176–o3178Figure 1
The molecular structure of the asymmetric unit of the title compound, showing 30% probablity displacement ellipsoids.
Figure 2
Plot of the crystal packing, showing the linkage of molecules by O—H···O hydrogen bonding (dashed lines). Labels a, b,
[image:5.610.126.495.352.618.2]supporting information
sup-3
Acta Cryst. (2005). E61, o3176–o3178(RS)-2,6-Dimethoxy-4-(2-methoxypropyl)phenol
Crystal data
C12H18O4
Mr = 226.26
Monoclinic, Pc
Hall symbol: p -2yc
a = 13.740 (3) Å
b = 9.405 (2) Å
c = 9.997 (2) Å
β = 102.190 (4)°
V = 1262.8 (5) Å3
Z = 4
F(000) = 488
Dx = 1.190 Mg m−3
Mo Kα radiation, λ = 0.71073 Å Cell parameters from 2536 reflections
θ = 2.6–25.4°
µ = 0.09 mm−1
T = 292 K Block, colorless 0.30 × 0.20 × 0.10 mm
Data collection
Bruker SMART APEX CCD area-detector diffractometer
Radiation source: fine focus sealed Siemens Mo tube
Graphite monochromator
ω scans
6654 measured reflections
2481 independent reflections 1809 reflections with I > 2σ(I)
Rint = 0.081
θmax = 26.0°, θmin = 2.6°
h = −16→16
k = −11→6
l = −12→11
Refinement
Refinement on F2
Least-squares matrix: full
R[F2 > 2σ(F2)] = 0.057
wR(F2) = 0.164
S = 0.97 2481 reflections 299 parameters 2 restraints
Primary atom site location: structure-invariant direct methods
Secondary atom site location: difference Fourier map
Hydrogen site location: inferred from neighbouring sites
H-atom parameters constrained
w = 1/[σ2(F
o2) + (0.1154P)2]
where P = (Fo2 + 2Fc2)/3
(Δ/σ)max < 0.001
Δρmax = 0.37 e Å−3
Δρmin = −0.19 e Å−3
Special details
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes.
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2,
conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used
only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2
are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger.
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
x y z Uiso*/Ueq
C1 0.6045 (4) 0.5670 (6) 0.4852 (6) 0.0891 (15)
H1A 0.5831 0.6425 0.5370 0.134*
H1B 0.6266 0.4879 0.5445 0.134*
H1C 0.5499 0.5379 0.4137 0.134*
supporting information
sup-4
Acta Cryst. (2005). E61, o3176–o3178H2A 0.7639 1.0064 −0.0065 0.161*
H2B 0.8803 1.0136 0.0179 0.161*
H2C 0.8230 1.1163 0.0978 0.161*
C3 0.6677 (3) 0.7268 (4) 0.3394 (4) 0.0567 (9)
C4 0.7519 (3) 0.7675 (4) 0.2909 (4) 0.0563 (9)
C5 0.7455 (3) 0.8834 (4) 0.2053 (4) 0.0575 (9)
C6 0.6549 (3) 0.9541 (5) 0.1646 (4) 0.0615 (10)
H6 0.6511 1.0320 0.1064 0.074*
C7 0.5725 (3) 0.9122 (5) 0.2075 (4) 0.0604 (10)
C8 0.5778 (3) 0.7984 (5) 0.2970 (4) 0.0618 (10)
H8 0.5217 0.7701 0.3284 0.074*
C9 0.4728 (3) 0.9858 (6) 0.1579 (4) 0.0741 (12)
H9A 0.4778 1.0476 0.0820 0.089*
H9B 0.4229 0.9142 0.1236 0.089*
C10 0.4377 (3) 1.0722 (5) 0.2650 (4) 0.0640 (10)
H10 0.4291 1.0092 0.3397 0.077*
C11 0.5079 (4) 1.1886 (5) 0.3224 (6) 0.0885 (14)
H11A 0.5254 1.2417 0.2489 0.133*
H11B 0.5669 1.1485 0.3783 0.133*
H11C 0.4766 1.2507 0.3769 0.133*
C12 0.2791 (4) 1.1581 (10) 0.2848 (6) 0.106 (2)
H12A 0.2581 1.0713 0.3206 0.159*
H12B 0.2220 1.2083 0.2350 0.159*
H12C 0.3133 1.2163 0.3589 0.159*
C13 0.0655 (4) 0.1060 (8) 0.9756 (8) 0.1043 (19)
H13A 0.0276 0.1819 1.0037 0.156*
H13B 0.0755 0.0324 1.0437 0.156*
H13C 0.0300 0.0679 0.8900 0.156*
C14 0.3877 (4) 0.4636 (7) 0.6184 (6) 0.0979 (18)
H14A 0.3470 0.4237 0.5371 0.147*
H14B 0.4567 0.4514 0.6159 0.147*
H14C 0.3733 0.5631 0.6231 0.147*
C15 0.1611 (3) 0.2431 (5) 0.8537 (4) 0.0631 (10)
C16 0.2586 (3) 0.2793 (4) 0.8426 (4) 0.0550 (9)
C17 0.2706 (3) 0.3661 (5) 0.7367 (4) 0.0635 (10)
C18 0.1891 (4) 0.4235 (5) 0.6469 (5) 0.0837 (14)
H18 0.1995 0.4826 0.5766 0.100*
C19 0.0936 (3) 0.3948 (5) 0.6597 (5) 0.0730 (12)
C20 0.0797 (3) 0.3014 (6) 0.7632 (5) 0.0798 (13)
H20 0.0155 0.2780 0.7717 0.096*
C21 0.0041 (4) 0.4538 (5) 0.5613 (6) 0.0873 (15)
H21A −0.0415 0.4939 0.6132 0.105*
H21B 0.0255 0.5300 0.5090 0.105*
C22 −0.0501 (4) 0.3442 (6) 0.4642 (6) 0.0849 (14)
H22 −0.0698 0.2654 0.5170 0.102*
C23 0.0156 (6) 0.2853 (9) 0.3679 (7) 0.125 (3)
H23A −0.0244 0.2278 0.2979 0.188*
supporting information
sup-5
Acta Cryst. (2005). E61, o3176–o3178H23C 0.0434 0.3631 0.3263 0.188*
C24 −0.2195 (6) 0.3201 (9) 0.3515 (14) 0.177 (5)
H24A −0.1962 0.2309 0.3235 0.266*
H24B −0.2668 0.3606 0.2768 0.266*
H24C −0.2506 0.3049 0.4277 0.266*
O1 0.6832 (2) 0.6152 (4) 0.4274 (4) 0.0855 (10)
O2 0.8411 (2) 0.7024 (4) 0.3321 (4) 0.0844 (10)
H2 0.8335 0.6277 0.3709 0.127*
O3 0.8320 (2) 0.9199 (4) 0.1675 (4) 0.0853 (10)
O4 0.3421 (2) 1.1273 (4) 0.1988 (3) 0.0772 (9)
O5 0.1574 (2) 0.1577 (4) 0.9601 (3) 0.0859 (10)
O6 0.33905 (18) 0.2330 (3) 0.9336 (3) 0.0658 (8)
H6A 0.3219 0.2050 1.0028 0.099*
O7 0.3678 (2) 0.3950 (4) 0.7325 (4) 0.0867 (11)
O8 −0.1376 (3) 0.4144 (4) 0.3907 (4) 0.1023 (13)
Atomic displacement parameters (Å2)
U11 U22 U33 U12 U13 U23
supporting information
sup-6
Acta Cryst. (2005). E61, o3176–o3178O6 0.0360 (13) 0.0909 (19) 0.0688 (16) 0.0089 (12) 0.0074 (11) 0.0195 (15) O7 0.0435 (16) 0.117 (3) 0.100 (2) −0.0062 (15) 0.0182 (15) 0.040 (2) O8 0.070 (2) 0.093 (2) 0.121 (3) 0.0152 (18) −0.031 (2) 0.015 (2)
Geometric parameters (Å, º)
C1—O1 1.405 (6) C13—H13A 0.9600
C1—H1A 0.9600 C13—H13B 0.9600
C1—H1B 0.9600 C13—H13C 0.9600
C1—H1C 0.9600 C14—O7 1.388 (6)
C2—O3 1.428 (7) C14—H14A 0.9600
C2—H2A 0.9600 C14—H14B 0.9600
C2—H2B 0.9600 C14—H14C 0.9600
C2—H2C 0.9600 C15—O5 1.343 (5)
C3—O1 1.357 (5) C15—C20 1.394 (6)
C3—C8 1.392 (5) C15—C16 1.408 (5)
C3—C4 1.399 (5) C16—O6 1.347 (4)
C4—O2 1.355 (5) C16—C17 1.374 (6)
C4—C5 1.377 (5) C17—O7 1.372 (5)
C5—O3 1.364 (5) C17—C18 1.388 (6)
C5—C6 1.394 (6) C18—C19 1.372 (7)
C6—C7 1.352 (6) C18—H18 0.9300
C6—H6 0.9300 C19—C20 1.402 (7)
C7—C8 1.387 (6) C19—C21 1.509 (7)
C7—C9 1.521 (6) C20—H20 0.9300
C8—H8 0.9300 C21—C22 1.501 (8)
C9—C10 1.502 (6) C21—H21A 0.9700
C9—H9A 0.9700 C21—H21B 0.9700
C9—H9B 0.9700 C22—O8 1.431 (6)
C10—O4 1.437 (5) C22—C23 1.555 (10)
C10—C11 1.491 (7) C22—H22 0.9800
C10—H10 0.9800 C23—H23A 0.9600
C11—H11A 0.9600 C23—H23B 0.9600
C11—H11B 0.9600 C23—H23C 0.9600
C11—H11C 0.9600 C24—O8 1.421 (8)
C12—O4 1.373 (6) C24—H24A 0.9600
C12—H12A 0.9600 C24—H24B 0.9600
C12—H12B 0.9600 C24—H24C 0.9600
C12—H12C 0.9600 O2—H2 0.8200
C13—O5 1.391 (6) O6—H6A 0.8200
O1—C1—H1A 109.5 H13A—C13—H13C 109.5
O1—C1—H1B 109.5 H13B—C13—H13C 109.5
H1A—C1—H1B 109.5 O7—C14—H14A 109.5
O1—C1—H1C 109.5 O7—C14—H14B 109.5
H1A—C1—H1C 109.5 H14A—C14—H14B 109.5
H1B—C1—H1C 109.5 O7—C14—H14C 109.5
supporting information
sup-7
Acta Cryst. (2005). E61, o3176–o3178O3—C2—H2B 109.5 H14B—C14—H14C 109.5
H2A—C2—H2B 109.5 O5—C15—C20 126.1 (3)
O3—C2—H2C 109.5 O5—C15—C16 113.8 (3)
H2A—C2—H2C 109.5 C20—C15—C16 120.0 (4)
H2B—C2—H2C 109.5 O6—C16—C17 119.9 (3)
O1—C3—C8 125.4 (3) O6—C16—C15 121.8 (3)
O1—C3—C4 114.1 (3) C17—C16—C15 118.4 (3)
C8—C3—C4 120.5 (3) O7—C17—C16 114.6 (3)
O2—C4—C5 118.7 (3) O7—C17—C18 124.0 (4)
O2—C4—C3 122.2 (3) C16—C17—C18 121.2 (4)
C5—C4—C3 118.9 (3) C19—C18—C17 121.3 (4)
O3—C5—C4 115.3 (3) C19—C18—H18 119.4
O3—C5—C6 125.1 (3) C17—C18—H18 119.4
C4—C5—C6 119.6 (3) C18—C19—C20 118.4 (4)
C7—C6—C5 121.7 (4) C18—C19—C21 122.1 (5)
C7—C6—H6 119.2 C20—C19—C21 119.5 (5)
C5—C6—H6 119.2 C15—C20—C19 120.6 (4)
C6—C7—C8 119.7 (4) C15—C20—H20 119.7
C6—C7—C9 121.4 (4) C19—C20—H20 119.7
C8—C7—C9 118.9 (4) C22—C21—C19 113.2 (4)
C7—C8—C3 119.5 (3) C22—C21—H21A 108.9
C7—C8—H8 120.2 C19—C21—H21A 108.9
C3—C8—H8 120.2 C22—C21—H21B 108.9
C10—C9—C7 114.6 (3) C19—C21—H21B 108.9
C10—C9—H9A 108.6 H21A—C21—H21B 107.8
C7—C9—H9A 108.6 O8—C22—C21 105.3 (4)
C10—C9—H9B 108.6 O8—C22—C23 112.4 (5)
C7—C9—H9B 108.6 C21—C22—C23 111.8 (5)
H9A—C9—H9B 107.6 O8—C22—H22 109.1
O4—C10—C11 111.5 (4) C21—C22—H22 109.1
O4—C10—C9 105.4 (3) C23—C22—H22 109.1
C11—C10—C9 113.1 (4) C22—C23—H23A 109.5
O4—C10—H10 108.9 C22—C23—H23B 109.5
C11—C10—H10 108.9 H23A—C23—H23B 109.5
C9—C10—H10 108.9 C22—C23—H23C 109.5
C10—C11—H11A 109.5 H23A—C23—H23C 109.5
C10—C11—H11B 109.5 H23B—C23—H23C 109.5
H11A—C11—H11B 109.5 O8—C24—H24A 109.5
C10—C11—H11C 109.5 O8—C24—H24B 109.5
H11A—C11—H11C 109.5 H24A—C24—H24B 109.5
H11B—C11—H11C 109.5 O8—C24—H24C 109.5
O4—C12—H12A 109.5 H24A—C24—H24C 109.5
O4—C12—H12B 109.5 H24B—C24—H24C 109.5
H12A—C12—H12B 109.5 C3—O1—C1 119.1 (4)
O4—C12—H12C 109.5 C4—O2—H2 109.5
H12A—C12—H12C 109.5 C5—O3—C2 116.5 (4)
H12B—C12—H12C 109.5 C12—O4—C10 115.0 (4)
supporting information
sup-8
Acta Cryst. (2005). E61, o3176–o3178O5—C13—H13B 109.5 C16—O6—H6A 109.5
H13A—C13—H13B 109.5 C17—O7—C14 118.7 (4)
O5—C13—H13C 109.5 C24—O8—C22 112.6 (4)
O1—C3—C4—O2 −1.7 (6) O6—C16—C17—C18 −175.5 (4)
C8—C3—C4—O2 178.5 (4) C15—C16—C17—C18 3.5 (7)
O1—C3—C4—C5 −177.4 (4) O7—C17—C18—C19 −177.4 (5)
C8—C3—C4—C5 2.8 (6) C16—C17—C18—C19 −0.5 (8)
O2—C4—C5—O3 1.3 (6) C17—C18—C19—C20 −2.3 (8)
C3—C4—C5—O3 177.2 (4) C17—C18—C19—C21 −178.7 (5)
O2—C4—C5—C6 −178.3 (4) O5—C15—C20—C19 176.9 (5)
C3—C4—C5—C6 −2.5 (6) C16—C15—C20—C19 0.8 (8)
O3—C5—C6—C7 −179.3 (4) C18—C19—C20—C15 2.1 (8)
C4—C5—C6—C7 0.3 (6) C21—C19—C20—C15 178.7 (5)
C5—C6—C7—C8 1.6 (6) C18—C19—C21—C22 106.4 (6)
C5—C6—C7—C9 −177.3 (4) C20—C19—C21—C22 −70.1 (7)
C6—C7—C8—C3 −1.2 (6) C19—C21—C22—O8 173.7 (5)
C9—C7—C8—C3 177.7 (4) C19—C21—C22—C23 −64.0 (6)
O1—C3—C8—C7 179.3 (4) C8—C3—O1—C1 −1.7 (7)
C4—C3—C8—C7 −1.0 (6) C4—C3—O1—C1 178.5 (4)
C6—C7—C9—C10 −110.2 (5) C4—C5—O3—C2 169.2 (5)
C8—C7—C9—C10 70.9 (6) C6—C5—O3—C2 −11.2 (7)
C7—C9—C10—O4 −178.0 (4) C11—C10—O4—C12 −82.5 (6)
C7—C9—C10—C11 59.9 (6) C9—C10—O4—C12 154.3 (6)
O5—C15—C16—O6 −1.2 (6) C20—C15—O5—C13 8.5 (8)
C20—C15—C16—O6 175.4 (4) C16—C15—O5—C13 −175.2 (5)
O5—C15—C16—C17 179.9 (4) C16—C17—O7—C14 170.4 (5)
C20—C15—C16—C17 −3.5 (7) C18—C17—O7—C14 −12.5 (8)
O6—C16—C17—O7 1.6 (6) C21—C22—O8—C24 −148.7 (8)
C15—C16—C17—O7 −179.4 (4) C23—C22—O8—C24 89.3 (8)
Hydrogen-bond geometry (Å, º)
D—H···A D—H H···A D···A D—H···A
O2—H2···O1 0.82 2.26 2.676 (4) 112
O2—H2···O8i 0.82 2.05 2.774 (5) 148
O6—H6A···O5 0.82 2.25 2.661 (4) 111
O6—H6A···O4ii 0.82 2.05 2.823 (4) 156