DOI: 10.1534/genetics.106.057364
Unusual DNA Structures Associated With Germline Genetic Activity in
Caenorhabditis elegans
Andrew Fire,
1Rosa Alcazar
2and Frederick Tan
3Departments of Pathology and Genetics, Stanford University School of Medicine, Stanford, California 94305-5324, Department of Biology, Johns Hopkins University, Baltimore, Maryland 21218 and Department of Embryology,
Carnegie Institution of Washington, Baltimore, Maryland 21218 Manuscript received February 17, 2006
Accepted for publication April 21, 2006
ABSTRACT
We describe a surprising long-range periodicity that underlies a substantial fraction ofC. elegans ge-nomic sequence. Extended segments (up to several hundred nucleotides) of theC. elegansgenome show a strong bias toward occurrence of AA/TT dinucleotides along one face of the helix while little or no such constraint is evident on the opposite helical face. Segments with this characteristic periodicity are highly overrepresented in intron sequences and are associated with a large fraction of genes with known germline expression inC. elegans. In addition to altering the path and flexibility of DNAin vitro, sequences of this character have been shown by others to constrain DNATnucleosome interactions, potentially
producing a structure that could resist the assembly of highly ordered (phased) nucleosome arrays that have been proposed as a precursor to heterochromatin. We propose a number of ways that the periodic occurrence of An/Tnclusters could reflect evolution and function of genes that express in the germ cell
lineage ofC. elegans.
D
NA carries biological information on a variety of levels. Some of the information is evident from well-defined sequence features such as the genetic code and certain reproducible transcription factor binding sites. Other (perhaps equally important) information in the genome may involve less precise sequence rules and longer-range interactions that are likely to be more difficult to detect and understand. One approach to dissecting the information encoded in the genome is to search for nonrandom character in the DNA sequence. Protein-coding constraints, for example, produce a non-random distribution in the utilization of base triplets. Likewise, certain transcription factors have a tendency to recognize multiple target sequences in a short in-terval, producing a localized increase in the incidence of one or more motifs. An intriguing means to inves-tigate nonrandom features of DNA sequence involves searches for periodic appearance of specific sequence elements (see Trifonov 1989 and Minsky 2004 forreviews). Previous analyses of this type have identified, among other nonrandom features, a strong tendency for 3n repeats in coding sequences (due to nonrandom usage of the genetic code and of amino acid sequences)
and a periodicity of 10–11 bp in occurrence of AA/TT dinucleotides. The latter periodicity has been observed rather strikingly in sequences that display intrinsic curvature in vitro (e.g., Koo et al. 1986; Ulanovsky
and Trifonov 1987; Goodsell and Dickerson 1994
and references therein). AA/TT dinucleotides are also unusual in having less flexibility under certain circum-stances than other dinucleotide pairs (e.g., Nelsonet al.
1987) and in their apparent ability to contribute to the lateral positioning of nucleosomes along DNA (e.g., Satchwellet al. 1986).
Although the bulk genome periodicity analysis has substantial power to detect patterns in the sequence, the biological significance of these patterns has remained somewhat of a mystery, in particular due to challenges in identifying and finding functional consequences corre-sponding to individual sequence characteristics. From this perspective, well-characterized model systems such asCaenorhabditis elegans provide a tool of considerable value.C. elegans has been reported to exhibit a strong
10-base periodicity signal (e.g., VanWye et al. 1991;
Widom 1996; Fukushima et al. 2002) and is among
the most extensively characterized both in structure (a complete sequence) and function (both individual ge-netic studies and whole-genome expression/phenotype analysis).
In this article, we analyze the surprisingly prevalent and extensive periodic character in the C. elegans ge-nome. Rather than underlying the entire genome sequence, the periodic regions appear enriched in a number of ‘‘islands’’ throughout the genome. These
1Corresponding author:Departments of Pathology and Genetics, Stan-ford University School of Medicine, 300 Pasteur Dr., Room L235, Mail Stop M5324, Stanford, CA 94305-5324. E-mail: afire@stanford.edu
2Present address:Pathology Department, Mail Stop M5324, Stanford University School of Medicine, 300 Pasteur Dr., Stanford, CA 94305-5324. 3Present address:Department of Biology, Johns Hopkins University, 3400 N. Charles St., Baltimore, MD 21218.
islands are for the most part unique in sequence; strikingly, they appear to delineate transcribed regions for a large group of genes that are expressed in the germ cell lineage ofC. elegans.
MATERIALS AND METHODS
Sequence versions and software: Sequences used for this analysis were downloaded from GenBank (Wheeler et al. 2005),Wormbase(Chenet al. 2005a), or theIntronerator(Kent and Zahler2000) databases at the times noted in describing each experiment. Use of distinct versions of certain sequences was in some cases necessitated by the availability of annotation files of a given type only for those versions. Basic observations [such as the strong enrichment of periodicity in theC. elegans genome and the propensity for periodic An/Tncluster (PATC)
islands to appear in autosomal arms] have been confirmed for numerous releases of the genome sequence and using several independent software tools with a wide variety of parameters.
SAGE data:Serial analysis of gene expression (SAGE) data were obtained from the Genome British Columbia (BC)C. ele-gansGene Expression Consortium (http://elegans.bcgsc.bc.ca/) (McKayet al. 2003; Blacqueet al. 2005; Chenet al. 2005b; B. Goszczynski and J. McGhee, personal communication; K. Wong, M. Marra, S. Jones, D. Baillieand D. Moerman, personal communication).
Computational analysis: Stringent repeat masking of ge-nomes was carried out as described by Kurtzand S chleier-macher(1999), using a PC-compatible machine running the Red Hat version of Linux. All other analyses were carried out on Macintosh systems using Pascal-based programming for processor-intensive tasks (Metrowerks Code Warrior v. 7) and Hypercard-based programming for text-intensive tasks. Sev-eral randomized sequences were derived independently from scripts in Perl, Hypercard, and Pascal with no evident differ-ences in periodicity properties.
RESULTS
Unusual character of some C. elegans DNA
frag-ments: This work started from an observation that
certain DNA sequences from C. elegans exhibited un-usual electrophoretic mobility on agarose gels at low temperature (S. White-Harrison, J. Fleenor and A.
Fire, unpublished observations). Retarded
electropho-resis, seen for DNAs from diverse biological sources, can be induced by specific sequence elements that can produce either a static bend or increased bendability in the otherwise straight helix (e.g., Mariniet al. 1982;
Olson and Zhurkin 2000). Several rather precise
models for predicting the three-dimensional path of the DNA helix as a function of base sequence (roll, tilt, and helical periodicity) have been published; these models can predict certain anomalies in electrophoretic behavior with considerable sensitivity and specificity, although the details of predicted trajectories differ considerably between different models (e.g., Goodsell
and Dickerson1994) and are thus at best a very rough
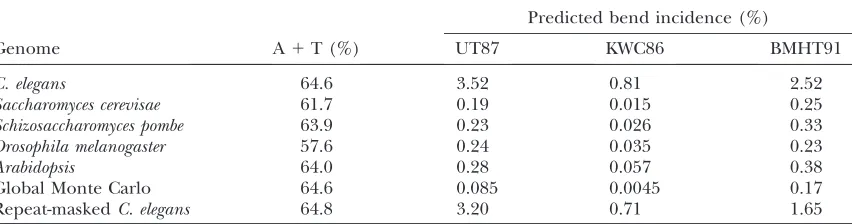
approximation. All of these algorithms predict strongly bent character in the segments ofC. elegans DNA for which we had observed abnormal mobility. The algo-rithms are readily scaled for high-throughput analysis of large numbers of sequences. This analysis revealed a higher overall ‘‘bend’’ density forC. elegansDNA than for other nonnematode genomes with a comparable base composition (Table 1).
Periodicity analysis of C. elegans DNA: While the ‘‘bending’’ analysis above demonstrates a nonrandom character in the DNA sequence, it is by no means clear
TABLE 1
High incidence of predicted static bends inC. elegansDNA
Predicted bend incidence (%)
Genome A1T (%) UT87 KWC86 BMHT91
C. elegans 64.6 3.52 0.81 2.52
Saccharomyces cerevisae 61.7 0.19 0.015 0.25
Schizosaccharomyces pombe 63.9 0.23 0.026 0.33
Drosophila melanogaster 57.6 0.24 0.035 0.23
Arabidopsis 64.0 0.28 0.057 0.38
Global Monte Carlo 64.6 0.085 0.0045 0.17
Repeat-maskedC. elegans 64.8 3.20 0.71 1.65
Predicted prevalence of DNA bending in several eukaryotic genomes is shown. Genomes of the indicated species were analyzed using the algorithms of Ulanovskyand Trifonov(1987) (‘‘UT87’’), Kooet al. (1986) (‘‘KWC86’’), and Bolshoyet al. (1991) (‘‘BMHT91’’). The UT87 and BMHT91 algorithms predict a trajectory of the DNA through three-dimensional space on the basis of perturbations to the helix from specific dinucle-otide sequences. The KWC86 algorithm predicts static bends at the borders of longer Anor Tnstrings. Values
that bendingper seis the biologically selected basis for this nonrandomness. Indeed, the diverse algorithms used above are all based onin vitrobehavior of DNA at relatively low temperatures; little or no gel mobility anomaly is generally observed at physiological temper-atures (Diekmann1987). Thus the unusual shapes of
DNAs predicted by the algorithms are unlikely to ac-curately represent the biological properties of the DNA in vivo.
When a set of predicted ‘‘bent’’ sequences from theC. elegansgenome was inspected in detail, we found that the most prominent common feature was a propensity for occurrence of AA or TT dinucleotides on one face of the helix over tens of base pairs (e.g., Figure 1). This is consistent with reports in the literature that periodic distributions of AA/TT dinucleotides can produce some of the largest anomalies in DNA structure (e.g., Bolshoy et al. 1991). To extend this analysis to the
whole genome, we carried out a general analysis of periodicity for the C. elegans genome. Figure 2 and supplemental Figure S1A (http://www.genetics.org/ supplemental/) show such an analysis for each of the 256 tetranucleotide ‘‘words.’’ These figures represent the number of times that two occurrences of a given word are separated by each integral number of base pairs.
As expected from previous analyses of periodicity in sequence databases (e.g., Trifonov1989), a variety of
different characteristics are evident from the periodicity analysis ofC. elegans.
1. For some tetranucleotides (e.g., TGTG; supplemental Figure S1A at http://www.genetics.org/supplemental/), a 2-base periodicity is present, apparently represent-ing segments of the genome with extended runs of alternating purine–pyrimidine.
2. A 3-base periodicity is evident for a number of the tetranucleotides (e.g., GTAG; supplemental Figure S1A). This type of periodicity is likely to represent the triplet nature of the genetic code combined with nonrandom codon choices and distributions of amino acids.
3. Additional 3n periodicities of 6, 9, etc. (e.g., GATT, TCAG; supplemental Figure S1A) presumably reflect (at least in part) coding for protein motifs such as beta sheet that have periodicity.
4. A number of the distributions show a strong and very discrete signal at a unique periodicity (e.g., GCCG; Figure 2C). These longer periodicities represent highly repeated tandem sequences in the genome. 5. An 10-base periodicity is present in words
con-taining multiple AA/TT dinucleotides (e.g., AAAA, Figure 2B).
Filtering the genome to remove repetitive and protein-coding sequences: To avoid the biases of re-petitive DNA and coding sequences, we sought to produce a version of the genome sequence in which repetitive sequences were stringently removed and coding sequences (as much as possible) ignored.
Many repeat masking algorithms are very limited in that only repeats from a limited and defined database are masked in the target sequence (e.g., the repeat-masker algorithm of Smit and Green) (Thomaset al.
2003). This type of operation was at best incomplete for the task at hand. Instead, we chose to apply an algorithm (reputer) and tools developed by Kurtzand S chleier-macher(1999). Starting with two parameters, window
(n) and stringency (m),reputer efficiently removes any segment ofn bp that can be aligned elsewhere in the genome with at leastmbases matching. We usedn¼25, m ¼ 25 (i.e., removal of all sequences of 25 bp that precisely match another sequence in the genome) and n¼28,m¼27 (i.e., removal of all sequences of 28 bp that match 27/28 to another sequence in the genome) with highly similar results. For the discussion below, we make use of the 25/25 genome masks.
utilizing a version of the genome in which coding sequences annotated by the genome project consor-tium had been removed (Kent and Zahler 2000).
Figure 2, C and E, and supplemental Figure S1B (http:// www.genetics.org/supplemental/) show the tetranucle-otide periodicity analysis for the repeat-subtracted, coding-region-depleted version of the C. elegans ge-nome. Two aspects of the genome emerge from this analysis. First, tetranucleotides rich in AA or TT dinu-cleotides retain their10-base periodicity, and second, the complex set of patterns for other tetranucleotides is substantially or completely removed.
The periodic distribution of AA/TT-containing tetra-nucleotides is consistent with previous observations (e.g., Widom 1996; Fukushima et al. 2002) as well as
the above analysis of bending predictions. Not as evident from some of the earlier analyses was the extent to which the periodicities can be detected over relatively long molecular distances. In Figure 2B, showing the frequencies of AAAA to AAAA distance as a function of base pair separation, a clear periodicity can easily be seen to extend beyond 200 bp.
Islands of periodicity in theC. elegansgenome:It was conceivable that the periodic signal might derive either uniformly from the whole genome or from some number of highly periodic areas or islands. Under the latter scenario, periodic signals might be enhanced by (i) selecting a tetranucleotide that was overrepresented in periodic islands and then (ii) calculating distance distributions from this tetranucleotide to other (arbi-trary) tetranucleotides. Such an analysis is shown in supplemental Figures S2A (distributions of distances between AAAA and an antecedent arbitrary tetranucleo-tide sequence) and S2B (distributions of distances be-tween AAAA and a subsequent arbitrary tetranucleotide sequence) (http://www.genetics.org/supplemental/). Particularly striking is the propensity for AA/TT-con-taining tetranucleotides to concentrate in one face of the helix(e.g., start-to-start separations 10, 20, 30 from adjacent AAAA/TTTT sequences), while non-AA/TT-containing tetranucleotides concentrate in intervening regions (e.g., 5, 15, 25, etc). These analyses indicate a periodic character to virtually all tetranucleotides when located in the vicinity of AAAA/TTTT sequences. The enhanced periodicity for arbitrary tetranucleotides in the vicinity of AAAA/TTTT tetranucleotides is indica-tive of a mosaic character to the genome,i.e., ofislands of periodicity. We give theseislandsthe formal designa-tion PATCs.
Detailed analysis of the histograms in supplemental Figure S2 (http://www.genetics.org/supplemental/) re-veals an interesting asymmetry in that complementary tetranucleotides can show different location profiles [e.g., (AAAA to GTCG) compared to (AAAA to CGAC), supplemental Figure S2B]. Similar asymmetry can be seen with AA/TT-rich tetranucleotides [e.g., (AAAA to TTTT) compared to (TTTT to AAAA) as expanded
in supplemental Figure S3 (http://www.genetics.org/ supplemental/)]. The latter comparison in particular shows significant differences in the$100-nt range that would not be expected with a simple localized deviation in DNA sequence. Rather, this comparison suggests a relatively intricate and asymmetric interaction that can cover many helical turns in the DNA.
Identification of individual periodic islands in theC. elegans genome: Although the statistical analysis above demonstrates nonrandom aspects of genome structure, our ability to understand the significance of the unusual structures depends on being able to identify and draw functional correlations between individual structures and genome function. To accomplish this, we needed algorithms that were able to detect a significant fraction (preferably as high as possible) of sequences with strong periodicity while showing maximal selectivity in avoid-ing detection of ‘‘false’’ positives (e.g., predicted highly periodic regions in random DNA sequence). Several different algorithms were evaluated for this purpose, including examination of localized sequence trajecto-ries using available models for DNA bending, variations of a hidden Markov model, a localized form of Fourier analysis, and a heuristic algorithm (PATC) described in Figure 3. All of these approaches gave similar results in the incidence and distribution of unusual DNA struc-tures in theC. elegansgenome, with general agreement on both localized and overall patterns of these struc-tures. The heuristic PATC algorithm was chosen for detailed analysis as it has given somewhat better signal-to-noise responses. For subsequent analysis we chose a cutoff score that excludes 99.999% of random se-quence. On the basis of this cutoff score (which has an arbitrary numerical value of 95), 6.14% of theC. elegans genome is within identified PATCs.
PATC motifs show a striking global pattern within the genome (Figure 4) with enrichment on the terminal approximately one-third of each autosome and on the extreme left tip of the X chromosome.C. elegans auto-somes are known to have distinct central and peripheral characteristics, with genes more densely packed in the center and more frequent recombination and occurrence of certain transposons in peripheral regions (Brenner
1974; Barnes et al. 1995; C. elegans Sequencing
Consortium1998; Duretet al. 2000).
that all PATCs are within transcribed regions. Periodic-ity of 59- and 39-UTR regions is also of interest and is described in Figure 5B as the best that could be extracted from current databases. Both 59- and 39-UTR regions clearly show above-background periodicity, although we stress that current annotation of UTR sequences is much less complete than that for introns and exons, potentially biasing the genomewide values for periodicity in UTRs.
To understand the functional significance of peri-odic An/Tnclusters in theC. elegans genome, we next
sought a rough size distribution for individual islands. Although the AAAA/TTTT separation plot in Figure 2C demonstrates strong periodicity beyond 200 bp, a
longer-range analysis of this plot becomes difficult as noise becomes sufficient to hide any signal. As an alternative approach, we examine the distribution of lengths for which the PATC algorithm described in Figure 3A can define a periodicity-enriched face of the helix. As shown in supplemental Figure S4A (http:// www.genetics.org/supplemental/), this distribution con-tinues beyond 1000 bp. Because the latter method could fortuitously ‘‘fuse’’ two adjacent periodic regions that happened to be in the same register, we applied a simpler and less ‘‘tuned’’ algorithm to look for periodic correlations over long distances. This algorithm simply assigns a weight to each 5-base word on the basis of the number of AA/TT dinucleotides and then sums the coincidence value (product) as a function of distance. As a starting data set for the latter procedure, we use the component of the genome most susceptible to high periodicity: introns present on autosomal arms. The resulting plot (supplemental Figure S4B at http:// www.genetics.org/supplemental/) shows striking albeit imperfect periodicity well beyond 500 bp. Interestingly, if we require that two intron-contained words be sep-arated not only bynbases, but by at least one exon se-quence, we maintain the strong periodicity (supplemental Figure S4C at http://www.genetics.org/supplemental/). This indicates that the specific periodicity can be main-tained even through a region of minimally periodic exon sequence.
Associations between periodic character and gene expression in C. elegans: The strong preference for periodicity in a subset of transcribed intron sequences raises the question of whether there might be correla-tion(s) between gene function and periodic character. To address this question, we first constructed a gene-by-gene list of periodicity values for introns, exons, and upstream and downstream segments (supplemental Fig-ures S5 and S6 at http://www.genetics.org/supplemental/). Given the genomewide preference for periodicity in intron sequences, we then used the intron PATC score for evaluating expression–periodicity associations.
Despite the remarkable functional genomic analysis available for C. elegans, the majority of the coding re-gions have been subject to only limited experimental characterization. We expected that the most reliable evaluation would come from the relatively small num-ber of genes for which functional data from classical genetics have been available (generally these are genes with classical genetic nomenclature and references to alleles isolated in forward mutagenic screens). Of the 10 most highly periodic genes on this list (sorted by intron periodicity), 9 (par-2,smu-2,mrt-2,ced-10,mel-46,sqv-2, hmp-2,ced-2, andapx-1) are known to express and/or function in the adult hermaphrodite germline, while only 1 gene in this set (ced-1) has not been reported to function in the germline [we note, however, that the observed lack of maternal rescue by ced-1(1) (Ellis
et al.1991) does not rule out expression during germline Figure3.—An algorithm for detecting local An/Tn
period-icity. (A) A schematic of the PATC algorithm for detecting and assigning a score to periodic An/Tnclusters in theC. elegans
development] (Kemphueset al. 1988; Elliset al. 1991;
Mello et al. 1994; Costa et al. 1998; Ahmed and
Hodgkin2000; Hwang et al. 2003; Spartzet al. 2004;
R. Minasakiand A. Streit, personal communication).
Cross-referencing of a somewhat larger subset of this list with data in both the C. elegansdatabase ‘‘Wormbase’’ and openly available articles inPubMed(supplemental Figure S7 at http://www.genetics.org/supplemental/) again revealed an unexpectedly high fraction (45/62) from this class with known roles or expression in the hermaphrodite germline. A comparable set of genes culled at random from the least periodic portion of the list included a much lower fraction with known germ-line roles (14/69). It should be stressed that although the difference in incidence of germline activity is highly statistically significant (P, 108), this analysis is quite imperfect, since annotation of gene expression and function in Wormbase and indeed in the literature as a whole is by nature incomplete.
A more objective (although still imperfect) assess-ment of association between periodic character and gene expression should be derivable from expression data obtained on a genomewide scale. Although micro-array assays have been the most frequently employed in such analyses (e.g., Kimet al. 2001), the resulting data are
complicated by a marginal signal-to-noise ratio among low-level-expressed genes. Another technique, SAGE
offers a somewhat better opportunity to definitively detect rare RNAs in a mixture (Velculescuet al. 1995).
Through the heroic efforts of the Genome BCC. elegans Gene Expression Consortium and their various collab-orators, the C. eleganscommunity is fortunate to have access to an extensive set of published and unpublished SAGE data (McKay et al. 2003; Blacque et al. 2005;
Chen et al. 2005b; B. Goszczynski and J. McGhee,
personal communication; K. Wong, M. Marra, S.
Jones, D. Baillieand D. Moerman, personal
reflective of expression levels, SAGE frequencies are certainly related to the underlying mRNA level. These data thus suggest a ‘‘reverse bell curve’’ association be-tween oocyte expression and periodic character.
Although the analyses in Figure 6B suggest a relation-ship between ooctye expression level and periodicity, there is no reason to assume that this relationship is ex-clusive. Indeed many of the genes described as periodic (e.g., supplemental Figures S5–S7 at http://www.genetics. org/supplemental/) are putative housekeeping genes involved in processes such as basal transcription that are essential for all cells. To determine which, if any, tissues were most tightly associated with periodic character, we needed a SAGE-compatible numerical method to eval-uate the quality of associations such as the bell-shaped curve in Figure 6A. To this end, we adapted a tech-nique from communications signal analysis (detailed in supplemental Methods at http://www.genetics.org/ supplemental/) that yields a result in the form of a likeli-hood ratio. Using this method to compare the expression/ periodicity association for the oocyte SAGE data to a
null hypothesis (no significant association) gives a total likelihood ratio of.1019. As shown in Figure 6B, a much less significant likelihood ratio is obtained from the other major tissues (gut, hypodermis, muscle, neurons, and pharynx). Better likelihood ratios were obtained from two additional SAGE data sets (pharyngeal mar-ginal cells and AFD neurons). It should be noted that the marginal cell and AFD data sets derive from a much rarer tissue (a small number of cells in each case). Contamination by germline or oocyte RNAs becomes a greater concern under such circumstances, and in-deed we observe that several genes that are thought to be germline specific in their activity are represented in these data sets. We stress, however, that no contam-ination has been demonstrated and thus the possibil-ity remains that these two tissues share considerable portions of their gene expression profiles with the germline. On the basis of this analysis of the SAGE data, we conclude that oocyte SAGE data representation provides the best associative model for predicting periodicity.
DISCUSSION
We have described an unusual DNA structure that underlies a significant fraction of theC. elegansgenome. This structure shares certain features with DNA that has been shown to curve or bend in several biochemical and ultrastructural assays (Crotherset al. 1990).
Nonethe-less, we note that the unusual sequence characteristics are not accounted for in detail by assumptions of a specific role in producing a bent naked DNAin vivo.
Nature of periodic An/Tnsequences in theC. elegans
genome: The long-range nature of the periodicities
observed inC. eleganssuggests that we are detecting the ‘‘shadow’’ of a large nuclear structure as it is reflected in DNA sequence. Our working hypothesis is that ex-tended regions with highly periodic character (‘‘PATC islands’’) correspond with the ability of the DNA to interact with some surface within the nucleus. Extended surfaces in the nucleus include the nuclear envelope,
the outer surfaces of nucleosome cores, neighboring DNA duplexes, and a number of other organelles (nu-clear granules and speckles, higher-order chromatin struc-tures, nuclear scaffolds, etc.) (Kornberg and Lorch
1999; Gallet al. 2004; Taddeiet al. 2004; Gruenbaum
et al. 2005).
Among the various nuclear structures, the outer surface of the nucleosome core provides perhaps the most fertile speculation. In particular, it is this surface that is thought to associate most prominently with the bulk of nuclear DNA (Kornberg and Lorch 1999).
AA/TT dinucleotides have been suggested to be in-volved in specific positioning of nucleosomes through a preference for their minor grooves positioned facing inward in the structure (e.g., Calladine and Drew
dinucleotides face minor-groove-inward would provide a substantial free energy benefit. Such structural inter-actions have thus been proposed to be capable of constraining nucleosomes by impeding their ability to slide along the DNA (e.g., Flausand Richmond1998;
Kiyama and Trifonov 2002; Kulic and Schiessel
2003). Although nucleosome mobilityin vivo is likely to involve a number of biophysical modalities (Flaus
and Owen-Hughes 2003), some sequence-dependent
kinetic impedance of nucleosome translocation could be expected from all of these modalities. A useful (if incomplete) analogy would be to a bicycle chain that prefers to limit its interaction with the corresponding gears to a quantized set of positions, thereby generating traction.
Several observations are consistent with the sugges-tion that PATC clusters reflect nucleosomal posisugges-tioning in C. elegans. First, nucleosomes association seems to be the default state for the bulk of eukaryotic nuclear DNA (including that ofC. elegans) (Dixonet al. 1990;
Kornberg and Lorch 1999). Second, nucleosome
modification complexes are critical in setting up appro-priate patterns of gene expression in the germline (Ahringer 2000; Shin and Mello 2003). Third, the
minor peaks in the Fourier plot in supplemental Figure S4E (http://www.genetics.org/supplemental/) and the observation in supplemental Figure S3 (http://www. genetics.org/supplemental/) that AAAA/TTTT sep-arations have a distinct profile from TTTT /AAAA separations is consistent with previous reports of non-symmetric dinucleotide preferences and nonuniform helical repeats at different positions within a nucleo-some (e.g., see Hayeset al. 1991; Ioshikheset al. 1996;
Lugeret al. 1997).
Although these data are consistent with a nucleoso-mal connection to the structural anonucleoso-maly in PATC sequences, we point out several caveats. First, regions of the DNA may adopt different protein associations under distinct cellular conditions; thus association with nucleosomes could be limited to a subset of tissues. Second, helical periodicities of DNA have not been determined for potential alternative models (e.g., for nuclear envelope-associated DNA). Third, differences and heterogeneity in TTvs. AA dinucleotide distribu-tion might also be expected for other physical inter-actions, either as a result of intrinsic structure and flexibility or as a result of specific protein binding.
One feature of the sequence anomalies that initially seems unexpected on the basis of nucleosomal models is the persistence of periodic structure. Individual nu-cleosomes would be expected to cover only 146–147 bp of DNA, with some extension potentially coming from defined linker proteins. Two situations could poten-tially extend this footprint: first, an array of tandemly arranged nucleosomes could potentially produce a longer periodicity, assuming that the spacing between nucleosomal cores was such that the 10.# base
peri-odicity was maintained (Widom1992). Second, as has
been proposed for a number of systems, a region of DNA might be specialized on the basis of constraining one or more nucleosomes not to a single position but rather to a quantized set of positions,e.g., to impede its movement on DNA. Such a situation might involve a larger region of DNA than would be covered by individual nucleosomes, just as a bicycle chain is much longer than the circumference of the gears that it connects with.
For purposes of discussion of functionality and evo-lution, we refer in the remainder of this discussion to the unknown surfaces that align to periodic regions as ‘‘nucleosomes.’’ We note, however, that all of the sub-sequent arguments could apply equally to another type of subnuclear surface.
Functional correlates of strong periodic character in the C. elegans genome: The strong bias for abundant periodicity in a discrete subset of genes suggests a functional commonality among these genes. Manual annotation of gene lists and objective comparisons of genomic data both indicate a statistically significant association between gene activity in the hermaphrodite germline and high sequence periodicity. Nonetheless, it should be stressed that this correlation could be secondary and that the functional link could be a complex one, for example, involving another cell type with similar expression profiles to the hermaphrodite germline or some metabolic process that is simply more effective in these cells. The C. elegansgermline passes through a rather dramatic series of transitions during development. Examining the lists of genes that show highly periodic character, we note that the majority of strongly periodic genes appear to be active in meiotic cells of the distal adult germline that act simultaneously (1) as nurse cells for mature oocytes and (2) as ‘‘oocytes-in-training.’’ Interestingly, we observed no strong period-icity in a trio of genes [him-17(Reddyand Villeneuve
2004), spo-11 (Dernburg et al. 1998), and rad-51
(Colaiacovoet al. 2003)] that are located on autosomal
arms and are thought to be active at premeiotic stages in germline function. Likewise we fail to observe strong periodicity in a large group of genes (many on the X chromosome) that are activated in a late burst of transcription during the last stages of oocyte maturation (Kellyet al. 2002). Spermatogenesis also has a
charac-teristic pattern of gene expression (L’Hernault and
Roberts 1995), and we see no evidence for unusual
periodicity of the majority of genes active in this aspect of germline development. These observations are con-sistent with the hypothesis that highly periodic charac-ter associates with expression at a stage of germline development in which oocyte precursors slowly traverse the early stages of meiosis.
Do An/Tnperiodicities in the C. elegansgenome
con-tribute to macroscopic fitness? Given the strongly
pressure must have been present for their origin and maintenance. Consistent with this expectation, we ob-serve regions with comparably high periodicity signals in a number of additional nematode genomes for which draft or fragmentary sequences are available [e.g.,C. briggsae (draft genome from Steinet al. 2003) andC. remanei,
Caenorhabditis species CB5161, and Caenorhabditis species PS110 (fragments from theama-1gene; Kiontke
et al. 2004)]. We envision two types of selective pressures that could contribute to forming and maintaining these structures: macroscopic selection (accumulation of peri-odic structures in DNA due to increased fitness for the animal) andmicroscopic bias (accumulation of periodic structures in DNA due to nonrandom mutagenesis and or repair).
For a macroscopic selection model to be applicable, periodic segments that are sufficiently unusual to be absent in random DNA each would need to contribute in some way to the phenotypic fitness of the animal. Such contributions might be relatively subtle, such that no single base mutation would produce a visible phe-notype in the laboratory. Alternatively, the effects could be dramatic, in which case one might expect to uncover a set of mutations whose phenotypic effects might be difficult to explain on the basis of standard aspects of DNA and RNA structure. Such ‘‘unexplained’’ muta-tional consequences have been rarely if ever reported in theC. elegans genetic literature, although biases for examining mutations with stronger phenotypes and biases in choices of mutagen may have masked the relevant interpretations. Estimates of total constraint due to highly periodic islands in theC. elegansgenome are rather substantial (with on the order of 106 non-coding bases showing nonrandom identity related to these structures). Spontaneous mutation rates in C. elegans(108/base/generation) are such that selective pressure on 106noncoding bases would pose a slight but evolutionarily significant burden on the species. At the same time, the likely nonlethal character of individual mutations that affect only one or a few periodic regions raises some interesting population genetic questions in terms of whether simple selection could allow highly periodic structures to be fixed and maintained within a population.
PATC islands in C. elegans would presumably make their contribution to organismal fitness through mod-ulation of nearby genetic elements (coding regions or other functional chromosomal features). The most attractive hypothesis to us is that these sequences im-pede nucleosome rearrangement in genes for which germline expression is important. Formation of inactive heterochromatin has been shown at least in a subset of cases to be associated with the assembly of nucleosomes into arrays with a specific and uniform spacing (e.g., Sun
et al. 2001). Constraints on nucleosome mobility (or alternatively, strong positioning signals that failed to match the required heterochromatic spacing) would be
expected to increase the energy and time needed to assemble these uniformly spaced arrays. This in turn could inhibit or delay the assembly of specific chromo-somal regions into higher-order (and potentially si-lenced) chromatin structures. Such a situation might serve to protect a subset of genes from active and somewhat indiscriminant silencing mechanisms that normally protect the germline from unwanted expres-sion of genes whose expresexpres-sion might be harmful. Harmful genes might include both somatic specifica-tion and differentiaspecifica-tion components and selfish DNAs such as transposons or viruses.
Although the forces that shape patterns of hetero-chromatin in the genome are not fully understood, they are almost certain to involve nucleation events (de novo initiation of heterochromatin structures) combined with a tendency of heterochromatin structures to spread lat-erally in a stepwise and somewhat deliberate manner until sequences that prevent their spread are encoun-tered (Richards and Elgin 2002). Initiation events
involve a mixture of triggers: specific DNA sequence elements that are targeted by interaction with proteins that initiate heterochromatic silencing, repeated struc-tures in DNA that may be recognized by meiotic or recombination machinery, parasitic genetic elements, and certain types of modulatory RNA interactions (Turker
and Bestor1997; Hsiehand Fire2000; Selker2002).
Interestingly, the outer autosomal regions where many of the highly periodic genes in C. elegans reside are enriched in repeated DNA and putative selfish elements (C. elegans Sequencing Consortium 1998). One
model would be that these elements contribute to a relatively inhospitable genomic environment for those genes whose activity in the germline is important. This might necessitate the ability of such genes to evolve mechanisms to resist the encroachment of heterochro-matic structure (which could potentially render them useless for the germline). A situation in which periodic regions of the germline-expressed genes were refractory to ordered nucleosome phasing and thus to higher-order heterochromatin assembly might provide these genes substantial protection from silencing.
Could biased mutagenesis and/or repair mecha-nisms contribute on a microscopic scale to periodicity in the C. elegansgenome?As a (nonexclusive) alterna-tive to models proposing an organismal selection for periodic regions in DNA, it is conceivable that such structures arise due to biases in mutagenesis and repair at the level of individual genomic regions. Biased muta-genesis and repair processes have certainly been docu-mented in numerous systems and could reflect both nucleosome positioning and transcriptional properties (Smerdon1991; Miller2005). The observed
periodic-ity inC. elegansDNA could be explained if a number of conditions were met.
would either (1) have distinct sensitivity to mutagenic hits or (2) be repaired with altered efficiency or spec-ificity after such hits. The most likely scenario here would be for transcription in germline tissue to be asso-ciated with altered mutability characteristics. The rela-tively stable early meiotic states slowly traversed by oocyte precursors in the distal germline make a good candidate for the tissue of interest here since these chromosomes can persist in a stage-specific conforma-tion for a considerable time period. Another possibility would be targeting of specific regions of chromatin that are accessible in diapause(e.g., dauer) animals, again at a stage where cells persist for a long period.
Second, the mutagenic processes in the animal would need to be influenced by local structural characteristics of the chromosome and its environment. Most attractive as a model here would be to propose that nucleosome interactions might influence the distribution of muta-genic hits and/or the spectrum of repair functions. Such influences have been hypothesized (Holmquist
1994) and demonstrated in model systems (e.g., Smerdon 1991; Schieferstein and Thoma 1996). In
theC. eleganscase, nucleosome positions would need to alter mutagenic consequences on a base-by-base level, skewing the eventual spectrum of mutations such that AA/TT dinucleotides would form more frequently on one face of a surface contacting the DNA.
Numerous mutational events would be required to produce a periodic structure. Assuming that these hits occurred over considerable evolutionary time, the mi-croscopic bias model requires that some force must constrain the nucleosome. One possibility would be a constraint of nucleosomes independent of the AA/TT periodicity: the AA/TT periodicity would then appear as a shadow of this underlying positional constraint. Alternatively, the AA/TT periodicity itself may work to constrain nucleosome positions, so that positioning of nucleosomes may become more and more constrained as further point mutations are accumulated. The latter model works under the interesting constraint that biases in mutagenesis and repair serve to reinforce current nucleosome positioning(e.g., Holmquist1994). This
would certainly be the case if mutagenic lesions that produce AA/TT dinucleotides with their minor grooves facing into the nucleosome were more frequently formed or fixed in the DNA sequence. Curiously, this set of conditions would set up a situation where strong nucleosome positioning signals could coalescede novo in DNA sequence through an increasingly energetically favorable and self-reinforcing process.
We have relatively little information from which to speculate on potential mutagenic processes that might produce the types of nonrandomness observed as periodicity in theC. elegansgenome. Virtually any point mutagen that was sensitive to DNA structure or context could produce such a bias. Likewise almost any repair process could be biased for or against events that occur
on surface-associated microregions in the DNA. For exemplary purposes, we note the following aspects of ultraviolet-induced mutagenesis: (a) UV has been shown to preferentially induce lesions at TT dimers, (b) repair can proceed by either an error-free mechanism (pho-toreactivation) or an error-prone mechanism (excision and gap repair), and (c) it is conceivable that gap repair would predominate on the outer (exposed) surface of nucleosomes while photoreactivation would predomi-nate on inner (sterically protected) faces. Combining this with other mutagenic processes that might result in formation of A- or T-rich segments in transcriptionally accessible regions of the genome, (Daiet al.2005) one
might expect the types of structures that we observe in nematodes.
On the possibility of macroscopic/microscopic co-evolution: Proposals that the strong periodicity seen in the C. elegansgenome might reflect a combination of structural constraints and mutagenesis/repair biases led to a plausible hypothesis that the resulting anoma-lies may have no particular fitness value for the or-ganism. While this would certainly be a valid hypothesis, an attractive alternative would be to propose that the aggregate changes occurring in certain germline-active genomic regions would confer structural characteristics that would yield a fitness advantage for the organism. One possible modality for this advantage would be the formation, over evolutionary time, of certain germ-line-active chromatin regions that would be protected from epigenetic silencing. Other potential roles for these sequences could be envisioned in chromosome structure, replication, recombination, segregation, or maintenance. Whatever the role in the function of the or-ganism, the unexpected suggestion here is that mu-tagenic and repair functions in the nematode phylum have been tuned over evolutionary time so that beneficial changes to certain regions of the genome accumulate without any immediate selection for the individual events. The proposal that organisms might evolve to manage their own evolutionary change is by no means original to this system (e.g., McClintock1984). As additional
nematode genomes are sequenced and functionally characterized, we expect that further information about the large-scale processes directing their change will become clearer.
Could PATC islands contribute to germline genome
defense? Organisms use numerous mechanisms to
protect themselves from selfish or unwanted informa-tion (transposons, viruses, etc.). These mechanisms include silencing of repeated DNA, meiotic silencing of unpaired DNA, RNAi, recognition and clearance of extended regions of ssDNA and untranslated RNA, and gatekeeping processes that prevent nuclear export of unspliced RNAs (Fire2006). Even with these defense
Unusual structural characteristics of germline ex-pressed genes could potentially provide a species with an additional level of protection. Foreign DNAs (whether viral, transposon, or from another species or phylum) would generally lack such characteristics and might thus be recognized as foreign. In the case of the periodic character of many germline-expressed genes in C. elegans, an ability of PATC islands to resist encroach-ment of silencing would allow the organism to employ a highly aggressive process for spreading of germline heterochromatin into unprotected regions of the ge-nome. The vast majority of sequences that entered the genome from outside sources would then be subject to effective silencing for germline expression during their first few generations in theC. elegansgenome, a pro-cess that is readily seen with many different transgene constructs that are introduced into C. elegans (Kelly
et al. 1997; Hsieh and Fire2000). The process of
in-jected transgene silencing inC. elegansis not completely understood but is thought to involve the progressive recruitment of heterochromatin-related histone mod-ifications to the extrachromosomal array structures into which the foreign DNA is incorporated (Hsiehand Fire
2000; Bean et al. 2004). Several different vectors are
available that seem to partially limit germline silencing of injectedC. eleganstransgenes; intriguingly the major-ity of these vectors are derived from genes with sub-stantial periodicity scores (let-858, mex-3, pie-1, smu-2, smu-1, andama-1) (Kellyet al. 1997; Reeseet al. 2000;
Sparzet al. 2004; M. Montgomery, S. Xu, W. Kellyand
A. Fire, unpublished data; M. Dunnand G. Seydoux,
personal communication). By contrast, germline-expressed genes with no detectable periodicity have (with one exception; W. Johnson and J. Dennis,
per-sonal communication; Lee and Schedl 2004) been
ineffective tools for vector development. Periodicity diagrams for several of the genes that have been tested in vector development are shown in supplemental Figure S8 (http://www.genetics.org/supplemental/).
Given that a number of germline-expressedC. elegans genes lack any detectable periodicity signal, one expects that the periodicity may not be the only force that can prevent heterochromatinization. The difference in peri-odic character between central and peripheral genes may indicate distinct optima in terms of protection from silencing, with periodicity-related processes the most effective means in the harsh environment of autosomal arms (which may have a greater proportion of silenced DNA) while alternatives may be most effective in tran-scription-rich chromosomal centers.
Although we see PATC periodicity most strikingly in nematode sequences, the existence of genome de-fense mechanisms based on species-specific or phylum-specific properties of DNA could in principle be quite general. One expects that other species could use features such as base composition (GC%), presence of specific protein-binding sites, or conserved DNA/
miRNA interactions to provide a layer of self–nonself discrimination and thus to protect their cells (and most importantly germline cells) from invasion or activity by unwanted information.
Tool building based on genome structure:Tools that manipulate gene expression in a specific cell type are critical for many experimental and therapeutic ma-nipulations. The highly periodic structure in a large number ofC. elegansgermline-expressed genes certainly argues that we should be aware of periodicity in de-signing germline expression vectors. This might guide the choice of promoters, coding regions, and reporters (synthetic or natural) for intended germline expres-sion. Ultimately, it may be useful to design our own reporter coding sequencesde novoto maximize periodic character (particularly in introns) and thus potentially prevent epigenetic silencing of resulting transgenes.
For experimental manipulations of nonnematode species (e.g., human cell lines and transgenic mice) some of the same challenges of gene silencing have been described and discussed (e.g., Bacheler et al.
1979; see Bestor2000 for review). Although the
struc-tures described in this article are somewhat specific to C. elegans, gene silencing applications outside of nem-atodes may still be worthy of investigation. First, the type of AA/TT periodicity described in this article, if it fun-damentally alters nucleosome positioning or mobility, might be directly useful to avoid silencing even in systems that lack these signals normally. Second, a full knowl-edge of nucleosome positioning signals in vertebrates might allow the construction of vertebrate-specific vec-tors that would likewise constrain nucleosomes from forming highly ordered heterochromatin.
We thank K. Wong, M. Marra, S. Jones, D. Baillie, D. Moerman, J. McGhee, B. Goszczynski, the M. Smith Genome Sciences Centre, Genome British Columbia, and Genome Canada for providing C. elegansSAGE data prior to publication. Additional help, suggestions, and support from Anne Villeneuve, Tim Schedl, Robert Herman, Bill Kelly, Susan Strome, Geraldine Seydoux, Barbara Meyer, David Schwartz, Phil Beachy, Robert Schleif, Don Brown, Ed Trifonov, Alan Wolffe, Adrian Streit, Roger Kornberg, Jim Priess, Dennis Dixon, SiQun Xu, Jamie Fleenor, Susan White-Harrison, Javier Lopez-Molina, Jeb Gaudet, Dave Hansen, Andrew Travers, Wendy Johnston, Jim Dennis, Weng-Onn Lui, Lia Gracey, Jonathan Gent, Rayka Yokoo, Cecilia Mello, Steve Johnson, Julia Pak, Hann Yew, Blake Hill, and the National Institutes of Health (grants R01GM37706 and T32GM07231) are gratefully acknowledged.
LITERATURE CITED
Ahmed, S., and J. Hodgkin, 2000 MRT-2 checkpoint protein is required for germline immortality and telomere replication in
C. elegans. Nature403:159–164.
Ahringer, J., 2000 NuRD and SIN3 histone deacetylase complexes in development. Trends Genet.16:351–356.
Bacheler, L., R. Jaenischand H. Fan, 1979 Highly inducible cell lines derived from mice genetically transmitting the Moloney murine leukemia virus genome. J. Virol.29:899–906.
Barnes, T., Y. Kohara, A. Coulsonand S. Hekimi, 1995 Meiotic recombination, noncoding DNA and genomic organization in
Bean, C., C. Schaner and W. Kelly, 2004 Meiotic pairing and imprinted X chromatin assembly inCaenorhabditis elegans. Nat. Genet.36:100–105.
Bestor, T., 2000 Gene silencing as a threat to the success of gene therapy. J. Clin. Invest.105:409–411.
Blacque, O., E. Perens, K. Boroevich, P. Inglis, C. Li et al., 2005 Functional genomics of the cilium, a sensory organelle. Curr. Biol.15:935–941.
Bolshoy, A., P. McNamara, R. Harrington and E. Trifonov, 1991 Curved DNA without A-A: experimental estimation of all 16 DNA wedge angles. Proc. Natl. Acad. Sci. USA88:2312–2316. Brenner, S., 1974 The genetics ofCaenorhabditis elegans. Genetics
77:71–94.
C.elegansSequencingConsortium, 1998 Genome sequence of the nematode C. elegans: a platform for investigating biology. Science282:2012–2018.
Calladine, C., and H. Drew, 1986 Principles of sequence-dependent flexure of DNA. J. Mol. Biol.192:907–918. Chen, N., T. Harris, I. Antoshechkin, C. Bastiani, T. Bieriet al.,
2005a WormBase: a comprehensive data resource for Caenor-habditis biology and genomics. Nucleic Acids Res. 33: D383– D389.
Chen, N., S. Pai, Z. Zhao, A. Mah, R. Newburyet al., 2005b Iden-tification of a nematode chemosensory gene family. Proc. Natl. Acad. Sci. USA102:146–151.
Colaiacovo, M., A. MacQueen, E. Martinez-Perez, K. McDonald, A. Adamoet al., 2003 Synaptonemal complex assembly inC.
el-egans is dispensable for loading strand-exchange proteins but critical for proper completion of recombination. Dev. Cell 5: 463–474.
Costa, M., W. Raich, C. Agbunag, B. Leung, J. Hardin et al., 1998 A putative catenin-cadherin system mediates morphogen-esis of theCaenorhabditis elegansembryo. J. Cell Biol.141:297– 308.
Crothers, D., T. Haran and J. Nadeau, 1990 Intrinsically bent DNA. J. Biol. Chem.265:7093–7096.
Dai, J., R. Chuangand T. Kelly, 2005 DNA replication origins in theSchizosaccharomyces pombegenome. Proc. Natl. Acad. Sci. USA 102:337–342.
Dernburg, A., K. McDonald, G. Moulder, R. Barstead, M. Dresseret al., 1998 Meiotic recombination inC. elegans ini-tiates by a conserved mechanism and is dispensable for homolo-gous chromosome synapsis. Cell94:387–398.
Diekmann, S., 1987 Temperature and salt dependence of the gel migration anomaly of curved DNA fragments. Nucleic Acids Res.15:247–265.
Dixon, D., D. Jonesand E. Candido, 1990 The differentially ex-pressed 16-kD heat shock genes ofCaenorhabditis elegansexhibit differential changes in chromatin structure during heat shock. DNA Cell Biol.9:177–191.
Duret, L., G. Maraisand C. Biemont, 2000 Transposons but not ret-rotransposons are located preferentially in regions of high recom-bination rate inCaenorhabditis elegans. Genetics156:1661–1669. Ellis, R., D. Jacobsonand H. Horvitz, 1991 Genes required for
the engulfment of cell corpses during programmed cell death inCaenorhabditis elegans. Genetics129:79–94.
Fire, A., 2006 Nucleic acid structure and intracellular immunity: some recent ideas from the world of RNAi. Q. Rev. Biophys. (in press). Flaus, A., and T. Owen-Hughes, 2003 Mechanisms for nucleosome
mobilization. Biopolymers68:563–578.
Flaus, A., and T. Richmond, 1998 Positioning and stability of nucle-osomes on MMTV 39LTR sequences. J. Mol. Biol.275:427–441. Fukushima, A., T. Ikemura, M. Kinouchi, T. Oshima, Y. Kudoet al., 2002 Periodicity in prokaryotic and eukaryotic genomes identi-fied by power spectrum analysis. Gene300:203–211.
Gall, J., Z. Wu, C. Murphyand H. Gao, 2004 Structure in the amphibian germinal vesicle. Exp. Cell Res.296:28–34. Goodsell, D., and R. Dickerson, 1994 Bending and curvature
cal-culations in B-DNA. Nucleic Acids Res.22:5497–5503. Gruenbaum, Y., A. Margalit, R. Goldman, D. Shumaker and
K. Wilson, 2005 The nuclear lamina comes of age. Nat. Rev. Mol. Cell Biol.6:21–31.
Hayes, J., D. Clarkand A. Wolffe, 1991 Histone contributions to the structure of DNA in the nucleosome. Proc. Natl. Acad. Sci. USA88:6829–6833.
Holmquist, G., 1994 Chromatin self-organization by mutation bias. J. Mol. Evol.39:436–438.
Hsieh, J., and A. Fire, 2000 Recognition and silencing of repeated DNA. Annu. Rev. Genet.34:187–204.
Hwang, H., S. Olson, J. Brown, J. Eskoand H. Horvitz, 2003 The
Caenorhabditis elegansgenessqv-2andsqv-6, which are required for vulval morphogenesis, encode glycosaminoglycan galactosyltrans-ferase II and xylosyltransgalactosyltrans-ferase. J. Biol. Chem.278:11735–11738. Ioshikhes, I., A. Bolshoy, K. Derenshteyn, M. Borodovskyand E. Trifonov, 1996 Nucleosome DNA sequence pattern revealed by multiple alignment of experimentally mapped sequences. J. Mol. Biol.262:129–139.
Kelly, W. G., S. Xu, M. K. Montgomeryand A. Fire, 1997 Distinct requirements for somatic and germline expression of a generally expressedCaernorhabditis elegansgene. Genetics146:227–238. Kelly, W., C. Schaner, A. Dernburg, M. Lee, S. Kimet al., 2002
X-chromosome silencing in the germline ofC. elegans. Develop-ment129:479–492.
Kemphues, K., J. Priess, D. Mortonand N. Cheng, 1988 Identif-ication of genes required for cytoplasmic localization in early
C. elegansembryos. Cell52:311–320.
Kent, W., and A. Zahler, 2000 The intronerator: exploring introns and alternative splicing inCaenorhabditis elegans. Nucleic Acids Res.28:91–93.
Kim, S., J. Lund, M. Kiraly, K. Duke, M. Jianget al., 2001 A gene expression map for Caenorhabditis elegans. Science 293: 2087– 2092.
Kiontke, K., N. Gavin, Y. Raynes, C. Roehrig, F. Piano et al., 2004 Caenorhabditis phylogeny predicts convergence of her-maphroditism and extensive intron loss. Proc. Natl. Acad. Sci. USA101:9003–9008.
Kiyama, R., and E. Trifonov, 2002 What positions nucleosomes?— A model. FEBS Lett.523:7–11.
Koo, H., H. Wuand D. Crothers, 1986 DNA bending at adenine. thymine tracts. Nature320:501–506.
Kornberg, R., and Y. Lorch, 1999 Twenty-five years of the nucleo-some, fundamental particle of the eukaryote chromosome. Cell 98:285–294.
Kulic, I., and H. Schiessel, 2003 DNA spools under tension. Phys. Rev. Lett.9114:8103.
Kurtz, S., and C. Schleiermacher, 1999 REPuter: fast computa-tion of maximal repeats in complete genomes. Bioinformatics 15:426–427.
Lee, M., and T. Schedl, 2004 Translation repression by GLD-1 pro-tects its mRNA targets from nonsense-mediated mRNA decay in
C. elegans. Genes Dev.18:1047–1059.
L’Hernault, S., and T. Roberts, 1995 Cell biology of nematode sperm. Methods Cell Biol.48:273–301.
Lowe, T., and S. Eddy, 1997 tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res.25:955–964.
Luger, K., A. Mader, R. Richmond, D. Sargentand T. Richmond, 1997 Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature389:251–260.
Marini, J., S. Levene, D. Crothersand P. Englund, 1982 Bent he-lical structure in kinetoplast DNA. Proc. Natl. Acad. Sci. USA79: 7664–7668.
McClintock, B., 1984 The significance of responses of the genome to challenge. Science226:792–801.
McKay, S., R. Johnsen, J. Khattra, J. Asano, D. Baillie et al., 2003 Gene expression profiling of cells, tissues, and develop-mental stages of the nematodeC. elegans. Cold Spring Harbor Symp. Quant. Biol.68:159–169.
Mello, C., B. Draperand J. Priess, 1994 The maternal genesapx-1 andglp-1and establishment of dorsal-ventral polarity in the early
C. elegansembryo. Cell77:95–106.
Miller, J., 2005 Perspective on mutagenesis and repair: the stan-dard model and alternate modes of mutagenesis. Crit. Rev. Bio-chem. Mol. Biol.40:155–179.
Minsky, A., 2004 Information content and complexity in the high-order organization of DNA. Annu. Rev. Biophys. Biomol. Struct. 33:317–342.
Olson, W., and V. Zhurkin, 2000 Modeling DNA deformations. Curr. Opin. Struct. Biol.10:286–297.
Reddy, K., and A. Villeneuve, 2004 C. elegansHIM-17 links chroma-tin modification and competence for initiation of meiotic recom-bination. Cell118:439–452.
Richards, E., and S. Elgin, 2002 Epigenetic codes for heterochro-matin formation and silencing: rounding up the usual suspects. Cell108:489–500.
Satchwell, S., H. Drewand A. Travers, 1986 Sequence periodici-ties in chicken nucleosome core DNA. J. Mol. Biol.191:659– 675.
Schieferstein, U., and F. Thoma, 1996 Modulation of cyclobutane pyrimidine dimer formation in a positioned nucleosome con-taining poly, dA.dT tracts. Biochemistry35:7705–7714. Selker, E., 2002 Repeat-induced gene silencing in fungi. Adv.
Genet.46:439–450.
Shin, T., and C. Mello, 2003 Chromatin regulation duringC. elegans germline development. Curr. Opin. Genet. Dev.13:455–462. Smerdon, M., 1991 DNA repair and the role of chromatin structure.
Curr. Biol.3:422–428.
Spartz, A., R. Herman and J. Shaw, 2004 SMU-2 and SMU-1,
Caenorhabditis elegans homologs of mammalian spliceosome-associated proteins RED and fSAP57, work together to affect splice site choice. Mol. Cell. Biol.24:6811–6823.
Stein, L., Z. Bao, D. Blasiar, T. Blumenthal, M. Brent et al., 2003 The genome sequence of Caenorhabditis briggsae: a plat-form for comparative genomics. PLoS Biol.1:E45.
Sun, F., M. Cuaycongand S. Elgin, 2001 Long-range nucleosome ordering is associated with gene silencing in Drosophila
mela-nogaster pericentric heterochromatin. Mol. Cell. Biol.21:2867– 2879.
Taddei, A., F. Hediger, F. Neumannand S. Gasser, 2004 The func-tion of nuclear architecture: a genetic approach. Annu. Rev. Genet.38:305–345.
Thomas, J., J. Touchman, R. Blakesley, G. Bouffard, S. Beckstrom -Sternberget al., 2003 Comparative analyses of multi-species sequences from targeted genomic regions. Nature424:788–793. Trifonov, E., 1989 The multiple codes of nucleotide sequences.
Bull. Math. Biol.51:417–432.
Turker, M., and T. Bestor, 1997 Formation of methylation patterns in the mammalian genome. Mutat. Res.386:119–130. Ulanovsky, L., and E. Trifonov, 1987 Estimation of wedge
compo-nents in curved DNA. Nature326:720–722.
VanWye, J., E. Bronson and J. Anderson, 1991 Species-specific patterns of DNA bending and sequence. Nucleic Acids Res.19: 5253–5261.
Velculescu, V., L. Zhang, B. Vogelsteinand K. Kinzler, 1995 Serial analysis of gene expression. Science270:484–487.
Wheeler, D., T. Barrett, D. Benson, S. Bryant, K. Caneseet al., 2005 Database resources of the National Center for Biotechnol-ogy Information. Nucleic Acids Res.33:D39–D45.
Widom, J., 1992 A relationship between the helical twist of DNA and the ordered positioning of nucleosomes in all eukaryotic cells. Proc. Natl. Acad. Sci. USA89:1095–1099.
Widom, J., 1996 Short-range order in two eukaryotic genomes: rela-tion to chromosome structure. J. Mol. Biol.259:579–588.