ABSTRACT
GUSTILO, ESTELLA MAILUM. Post-transcriptional Modifications of the tRNA Anticodon Stem and Loop (ASL) Affect the Ability of tRNA to Bind Synonymous Codons. (Under the direction of Dr. Paul F. Agris).
The Genetic Code is arranged into sixteen codon boxes, where the four codons in each box are similar in their first two letters but differ at the third position (the wobble position). In the universal Genetic Code, each amino acid, except for Tryptophan and Methionine that have one codon each, is encoded by two to six codons (two to six-fold degenerate). There are fewer tRNA species than codons; therefore, a tRNA species can recognize more than one codon. This flexibility in recognition resides in the third position of the codon:anticodon pairing, the wobble position.
anticodon restrict codon recognition to the two codons specific for that 2-fold degenerate amino acid. For example, Lysine has two codons (AAA and AAG) that share a codon box with Asparagine codons (AAU and AAC). The modified nucleoside 5-methoxycarbonylmethyl-2-thiouridine at the wobble position of human tRNA (tRNALys mcm5s2U34) confers this tRNA’s ability to restrict codon recognition to the two Lysine codons only.
Post-transcriptional Modifications of the tRNA Anticodon Stem and Loop (ASL) Affect the Ability of tRNA to Bind Synonymous Codons
by
Estella M. Gustilo
A dissertation submitted to the Graduate Faculty of North Carolina State University
In partial fulfillment of the Requirements for the degree of
Doctor of Philosophy
Molecular and Structural Biochemistry
Raleigh, North Carolina June 5, 2009
APPROVED BY:
_______________________________ ______________________________
Dr. Paul F. Agris Dr. E. Stuart Maxwell
Committee Chair
DEDICATION
To my father who showed me the magic in books and the wonders of learning. To my
mother who expressed unconditional love. To my sister Marietta who offered unconditional
BIOGRAPHY
Estella M. Gustilo was born in the Philippines and immigrated to the United States with her
family at a young age. She grew up in Charleston, SC, where she attended college. Upon
graduation from college she worked in molecular biology research, where she found her
ACKNOWLEDGMENTS
The process of earning my Ph.D. has blessed me with wonderful opportunities of
learning. I will be forever grateful for those who contributed to my growth during these
years. First, I extend my deepest thanks to my mentor, Dr. Paul F. Agris, whose brilliance
and dedication to his craft has been inspiring. Dr. Agris has shown me amazing possibilities.
I thank each of the other members of my graduate committee; Dr. E. Stuart Maxwell, Dr.
Linda Spremulli, and Dr. Paul Wollenzien have given me valuable advice, collaborated in
these studies, and expanded my horizons.
I acknowledge my fellow lab members, especially William Darnell Graham, Dr.
Franck A. P. Vendeix, and William A. Cantara for providing me with dependable support.
My fellow biochemistry graduate students made this experience so much fun. Dr. Tatjana
Shapkina and Dr. Keith Gagnon have shared technical knowledge that has helped make this
study become more successful. Dr. Simpson Joseph has been extremely kind and opened his
lab and resources to me for several weeks to learn precious methods used in this work. This
study wouldn’t have been possible without technical training provided by Dr. Steven S.
Phelps. Steve has been such a wonderful, supportive friend who has taught me so much.
As always, my close family and friends have been instrumental in all the good things
that have come from my life. Even as science fills my mind, they fill my heart. My parents
and my sister Marietta have never failed me. Finally, but not least, I must thank the Creator,
whose mind I’ve seen only glimpses of in my study of science but whose heart I know fully.
TABLE OF CONTENTS
LIST OF TABLES………. vii
LIST OF FIGURES ……….. viii
CHAPTER 1. Post-transcriptional modifications are a trademark of tRNA…….….. 1
1.1 The roles of transfer ribonucleic Acid (tRNA) and the significance of post-transcriptional modifications to tRNA functions………..………. 1
1.2 The Genetic Code………..………. 7
1.3 The effects of post-transcriptional modifications of tRNA on synonymous-codon reading………..………….. 9
REFERENCES………..……… 16
CHAPTER 2. Modifications of the anticodon stem and loop of human tRNALys (ASLLysUUU) restrict codon recognition to Lysine codons AAA and AAG……...……... 24
2.1 Introduction………...……….. 24
2.2 Materials and Methods………...………. 26
2.3 Results………...……….. 28
2.4 Conclusions………...……….. 28
REFERENCES……….……..………... 32
CHAPTER 3. Anticodon domain modifications contribute order to tRNA for ribosome-mediated codon binding………...……….. 35
ABSTRACT………...………… 36
INTRODUCTION………. 39
EXPERIMENTAL PROCEDURE………..……..……… 42
RESULTS………..……… 47
DISCUSSION ……….….………. 62
CONCLUSION………..…..…….. 70
ACKNOWLEDGEMENTS……….……..…… 71
SUPPORTING INFORMATION AVAILABLE………..……..…….. 71
REFERENCES……….………. 72
ABSTRACT……….………….. 96
INTRODUCTION……….……… 96
MATERIALS AND METHODS……….………. 98
RESULTS AND DISCUSSIONS……….………….……... 107
FUNDING……….………… 114
ACKNOWLEDGEMENTS……….………. 114
REFERENCES……….………. 116
CHAPTER 5. The wobble position modified nucleoside of human mitochondrial tRNAMet decodes unconventional codons in the mitochondrial Genetic Code……….. 134
ABSTRACT……….. 135
INTRODUCTION……….……… 136
RESULTS……….………. 140
DISCUSSION ………... 146
MATERIALS AND METHODS……….………. 151
ACKNOWLEDGEMENTS……….…….………. 155
REFERENCES……….………. 157
CHAPTER 6. Summary and Insights……….……….. 172
6.1 The Evolution of tRNA and the Genetic Code……… 172
6.2 Implications of tRNA Modifications on Gene-Expression Regulation………... 175
6.3 tRNA codon-reading flexibility is Modulated by the Modifications at the Anticodon Loop……… 181
LIST OF TABLES
CHAPTER 1
Table 1. Distribution of modified nucleosides………. 11
Table 2. The recognition of tRNA wobble position-34 for the wobble position (N3) of the
codon………. 15
CHAPTER 3
Table 1. Thermal Parameters of Unmodified and Modified ASLVal3UAC ... 77
Table 2. Affinity of Unmodified and Modified ASLVal3UAC for the Valine Codons ……... 77
SUPPLEMENTARY MATERIALS:
Table 1. Local base step parameters of the unmodified ASL Val3UAC……….……….. 93
CHAPTER 4
Table 1. Thermodynamic contributions of f5C34………..……….. 125
CHAPTER 5
Table 1. Affinity of Unmodified and f5C34-Modified hmtASLMetCAU-Ψ27 for the
Mitochondrial Methionine Codons at E. coli Ribosomal P or A-site ………... 161
CHAPTER 6
Table 1. Contribution of modifications to codon recognition, frameshifting, and as
LIST OF FIGURES
CHAPTER 1
Figure 1. Bacterial tRNAs on the ribosome during translation……… 2
Figure 2. The tRNA……….………. 3
Figure 3. The Universal Genetic Code……….……… 8
CHAPTER 2
Figure 1. The anticodon stem and loop of human tRNALys3 (ASLLys3UUU-mcm5s2U34,
ms2t6A37)………..………….. 30
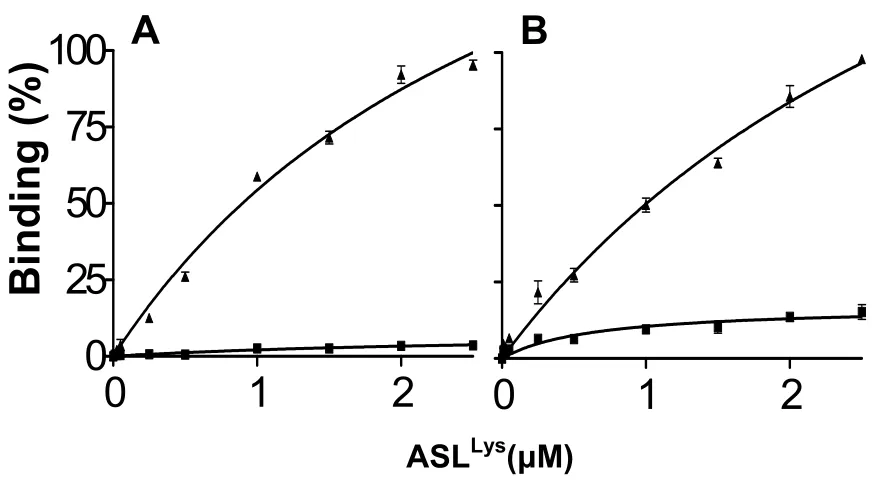
Figure 2. Ribosomal, equilibrium binding curves of the fully-modified human ASLLys3UUU-
mcm5s2U34, ms2t6A37 or the unmodified ASLLys3UUU………...…………. 31
CHAPTER 3
Figure 1. Nucleotide sequence of the Escherichia coli tRNAVal3UAC anticodon stem and loop
(ASLVal3UAC) and its modified nucleosides……… 82
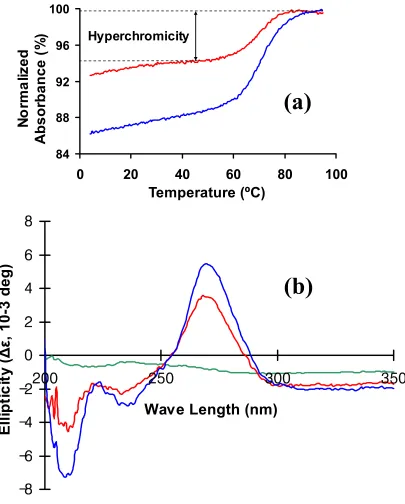
Figure 2. UV-monitored, thermal transitions and circular dichroism specta of unmodified
and modified ASLVal3UAC……….……….. 83
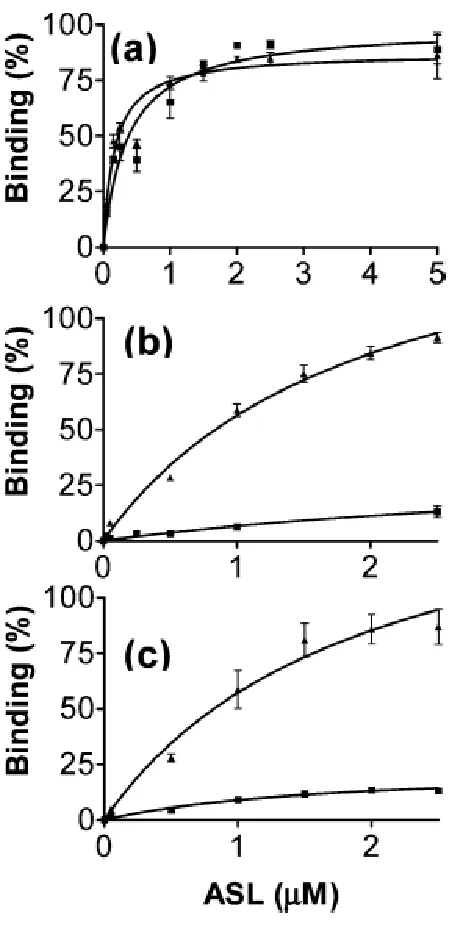
Figure 3. Binding of unmodified and modified ASLVal3UAC to valine codons at the A-site of
E. coli 30S ribosomal subunits……….. 84
Figure 4. Detection of the base-paired imino protons of the ASLs by NMR………... 85
Figure 5. Superimposed 1H-1H COSY spectra of unmodified and modified ASLVal3UAC… 86
Figure 7. Solution structures of ASLVal3UAC -cmo5U34;m6A37 and ASLVal3UAC and the rmsd for each of their nucleotides……….……….. 88
Figure 8. Superimposition of the average structures of the loop residues of ASLVal3UAC -cmo5U34; m6A37 with that of ASLVal3UAC……….……….. 89
Figure 9. Loop structure of ASLVal3UAC-cmo5U34;m6A37 in solution compared to that of the crystallographic structure on the ribosome……… 90
SUPPLEMENTARY MATERIALS:
Figure 1. Superposition of the 1H-31P HETCOR spectra of ASLVal3UAC and ASLVal3
UAC-cmo5U34;m6A37………..…………. 91
Figure 2. Anomeric to aromatic connectivity for ASLVal3UAC and ASLVal3UAC-cmo5U34;m6A37 ……….……….. 92
CHAPTER 4
Figure 1. Human mitochondrial tRNAMetCAU ………... 126
Figure 2. Synthesis of the 5-formylcytidine phosphoramidite………... 127
Figure 3. NMR spectra of the hmASLMetCAU ………... 128
Figure 4. HPLC nucleoside composition and NMR analyses of hmASLMetCAU and f5C…... 129
Figure 5. Thermal denaturations and circular dichroism spectra of the hmASLMetCAU -Ψ27 and
the hmASLMetCAU -Ψ27;f5C34……….……. 130
Figure 6. Codon binding by hmASLMetCAU -Ψ27 and the hmASLMetCAU -Ψ27;f5C34……….. 131 Figure 7. Analysis of the pKa of cytidine in comparison to that of 5-formylcytidine……... 132
Supplemental Figure. Heavy atoms root mean square deviation (r.m.s.d.) variations as a function of the molecular dynamics simulation time for the hmASLMetCAU-Ψ27;f5C34 and for
the hmASLMetCAU-Ψ27 ………... 133
CHAPTER 5
Figure 2. Human mitochondrial tRNAMetCAU……….………….. 166
Figure 3. Ribosomal binding curves of the wobble-modified hmtASLMetCAU-Ψ27;f5C34 or the
hmtASLMetCAU-Ψ27……….……… 167
Figure 4. Binding kinetics of the hmtASLMetCAU-Ψ27;f5C34 and the hmtASLMetCAU-Ψ27 to AUG or AUA at the P- and A-sites……….……….. 168
Figure 5. Codon binding of the hmtASLMetCAU-Ψ27;f5C34 and the hmtASLMetCAU-Ψ27 on the bovine 55S mitochondrial ribosome……….. 169
Figure 6. Equilibrium binding of the hmtASLMetCAU-Ψ27;f5C34 or the hmtASLMetCAU-Ψ27 to the unconventional Methionine codons AUU, AUC, and AUA……… 170
Figure 7. The molecular dynamics simulation of the A:f5C base pair……….………. 171
CHAPTER 6
Figure 1. Accommodation of tRNA on the ribosome………... 174
Figure 2. Modifications of the anticodon stem and loop (ASL) of tRNA order the loop, prevent frameshifting and allow accurate codon selection……… 176
CHAPTER 1
Post-transcriptional modifications are a trademark of tRNA
1.1 The roles of transfer ribonucleic acid (tRNA) and the significance of
post-transcriptional modifications to tRNA functions.
Transfer ribonucleic acid (tRNA) is a crucial participant in protein biosynthesis. The
tRNA’s primary role occurs on the ribosome during translation of genetic information, where
the tRNA reads the codes for amino acids imbedded in messenger RNA (mRNA) and places
the appropriate amino acid into a growing protein (Figure 1). The Genetic Code is the
language of protein synthesis; it is stored in deoxyribonucleic acid (DNA), transcribed into
mRNA, and translated into proteins in the ribosome by the tRNA. Therefore, the tRNA is a
critical translator of the Genetic Code in gene expression. tRNA is the faithful keeper of the
Genetic Code, keeping fidelity from the start of protein synthesis with aminoacylation to
completion with the production of viable proteins. Inherent in its primary, secondary, and
tertiary structure is information that gives the tRNA its ability to function (Figure 2). The
tRNA’s function is dependent on its interactions with a variety of cellular molecules such as
aminoacyl-tRNA synthetases (aaRSs), translation initiation factors, translation elongation
factors, mRNAs, ribosomes, and peptidylhydrolases. Post-transcriptional modifications of
Figure 1. Bacterial tRNAs on the ribosome during translation. During initiation, the 30S
ribosomal subunit (transparent grey circle) binds the mRNA (light blue strip) and the initiator tRNA
at the ribosomal peptidyl (P)-site with the aid of initiation factors (not shown). The 50S subunit (dark
grey) then joins the complex to form the complete 70S ribosome. Amino acids are added to the
nascent protein during elongation, where an elongator aminoacyl-tRNA binds its codon on the mRNA
at the the aminoacyl (A)-site with the aid of elongation factors. A peptide bond is formed between
the amino acid bound to the tRNA at the P-site and the amino acid of the tRNA at the A-site, and the
entire protein chain is transferred to the tRNA at the A-site. The P-site tRNA then moves to the exit
(E)-site (not shown), and the tRNA with the growing protein at the A-site is translocated to the P-site.
Another elongator aminoacyl-tRNA binds its codon at the A-site until the termination stage, where
release factors terminate protein synthesis. Accuracy and efficiency are achieved when the
appropriate tRNAs bind their corresponding codon.
A
P
Tyr
Met
Lys
Ser
UCG AUG AGC UAC
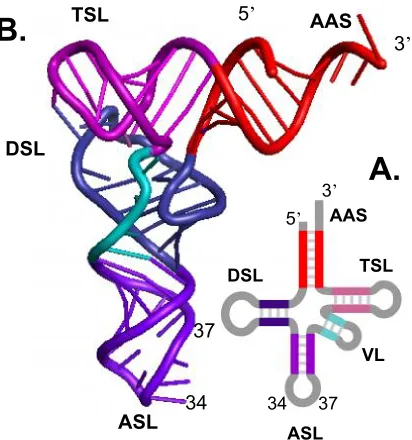
Figure 2. The tRNA. A. The secondary cloverleaf structure of tRNA. B. The tertiary L-shaped
structure of tRNA. The anticodon triplet consists of positions 34, 35, and 36. The tRNA consists of
four main domains: the dihydrouridine stem and loop (DSL), the anticodon stem and loop (ASL), the
ribothymidine (T), pseudouridine (Ψ), cytidine (C) stem and loop (TSL). The tRNA also consists of a
segment of a variable number of nucleosides (VL). Positions 34 and 37 are often
post-transcriptionally modified.
TSL
ASL
DSL
AAS
5
’
3
’
ASL
DSL
TSL
AAS
VL
3
’
5
’
34
37
34
37
Approximately 100 different modified nucleosides have been found in RNA [5].
tRNA is the most highly modified nucleic acid in the cell and consists of the greatest variety
of modification chemistries [6]. Post-transcriptional modifications decorate nucleosides all
along the tRNA and are integral to its purpose [1-4]. Up to thirty percent of tRNA
nucleosides at more than sixty different positions are modified [6]. Modified nucleosides are
derivatives of the four ribonucleic acid monomers Adenosine (A), Guanosine (G), Uridine
(U), and Cytidine (C), and therefore add additional chemistries to the limited number of only
four main nucleosides. The significance of modified nucleosides in the tRNA is evident in
their conservation among species and the amount of genetic material assigned to their
synthesis. In bacteria, approximately one percent of the genome codes for tRNA modifying
enzymes - this is four times more than the approximately 0.25 percent of bacterial genes
designated to tRNA [7]. Therefore, more genetic information is allocated to the modifying of
tRNA than to the tRNA themselves [7]. Modifications help the tRNA fold into its secondary
cloverleaf structure and tertiary L-shaped conformation mandatory for function and may aid
in the tRNA’s many interactions with proteins [2, 8-12]. Modifications are important to the
stability of tRNA and thus, affect its half-life [13-14].
Aminoacylation of the tRNA is the first step of protein synthesis. Aminoacyl-tRNA
synthetase charges the tRNA with the correct amino acid. Fidelity of translation begins at
aminoacylation. Modifications on the tRNA are specific identity determinants by some
aminoacyl-tRNA synthetases and therefore important to the aminoacylation of these tRNAs
[15-19]. Modifications may also affect aminoacylation via the modifications’ contributions
unmodified) can be aminoacylated, there are small differences in kinetics, suggesting that
modified nucleosides can be of some influence [11]. This is likely due to the inability of
tRNA to fold into its functional conformation. Unmodified tRNAs also have a greater
probability of being mischarged by a noncognate aminoacyl-tRNA synthetase [21, 22].
In the succeeding steps to the production of functional proteins, tRNA interacts with
many proteins such as inititation factors or elongation factors, and the proteins of the
ribosome. In the ribosome, it decodes the sequence of mRNAs into sequences of protein.
The tRNA is not a passive adaptor in translation but an active player during its function by
undergoing conformational changes [23-25]. Being the linker between genetic language and
end product (protein), the tRNA’s accuracy is necessary if the cell is to produce functional
proteins. And this fidelity must be balanced with the high speed of translation (over 20
peptide bonds per second). Post-transcriptional modifications at or near the anticodon of
tRNA are often required for accurate and efficient translation of the codons of mRNA [1, 11,
25-35].
The tRNA is an ancient molecule that is a remnant of the RNA World, where cellular
processes were carried out solely by RNA [36]. Thus, the tRNA has evolved into a
multifunctional molecule in the cell and is involved in a variety of processes in the cell.
While its main job is the adaptor molecule in protein synthesis, the tRNA has been found to
have other roles outside of translation on the ribosome [37]. The tRNA has been found to
function in viral proliferation [38-40], gene expression regulation [41, 42], cell division/DNA
chlorophyll synthesis [50]. Post-transcriptional modifications of tRNA may be directly or
indirectly involved in the functions of tRNA [11, 51].
The tRNA has been shown to be an important player in retroviral replication [38-40].
A specific tRNA isoacceptor, tRNALys3, is recruited by infected cells as the primer for the
reverse transcription of human immunodeficiency virus (HIV-1) [39, 52-55]. This has placed
specific tRNAs in study as potential drug targets [38]. The tRNALys3 consists of specific
modifications at the anticodon loop [30]. These modifications play a role in HIV-1’s
specificity for tRNALys3 and stabilize the tRNA Lys3’s interaction with the HIV-1 genome
[56-58].
The tRNA is also involved in DNA replication and/or cell division. Specific Serine
and Arginine tRNA isoacceptors have been shown to affect the cell cycle [59, 60]. The
modifications of tRNA may be involved in cell cycle regulation in that certain tRNA
isoacceptors with specific modifications are apparent in certain cell cycle stages [61-63].
The degree of tRNA modifications’ importance in many cellular processes has yet to
be fully appreciated. tRNA modifications are most studied in the context of tRNA’s central
role of translation. The modifications at tRNA’s anticodon stem and loop (ASL) are
especially important, as they have been shown to be significant factors in codon: anticodon
pairing, reading frame maintenance, translocation of tRNA from the ribosomal aminoacyl
(A)-site to the peptidyl (P)-site, and the balance between decoding accuracy and speed
[64-66]. The tRNA position 34 (wobble position) and position 37 at the ASL are often modified
Modifications at these two specific positions affect tRNA’s main function of reading codons
on the ribosome.
1.2 The Genetic Code
Proteins are produced by the cell using instructions compiled in the Genetic Code
(Figure 3). The Genetic Code is stored in DNA, transcribed into mRNA, and translated on
the ribosome by tRNA. The elucidation of the Genetic Code is arguably the greatest
accomplishment in the biomedical research of the 1960s. Upon its discovery, the Genetic
Code was considered universal in that there appeared to be little variation among organisms.
The Genetic Code consists of 64 codons: 61 sense codons that code for any of the 20 amino
acids and three termination codons. In the universal Genetic Code, most amino acids have
two to six codons. The different codons that code for the same amino acid are synonymous
codons. Only Tryptophan and Methionine have no synonymous codons, as these two amino
acids only have one codon each.
The Genetic Code is arranged into sixteen sets of four codons, portrayed in a table of
sixteen codon boxes (Figure 3). The four codes of a codon box are similar to each other in
the first two letters and differ only in the third letter. Some amino acids are encoded in
synonymous codons all residing in the same codon box. Other amino acids share the codes
of a codon box; these codons are termed to be in a “split box.” For example, Valine is
have two codons that share a codon box. Thus, the codons of Asparagine and Lysine are
very similar and differ only in the third position, the wobble position of the codon.
Figure 3. The universal Genetic Code. There are 61 sense codons that code for the 20
amino acids and 3 stop codons. The Genetic Code is arraigned into 16 codon boxes. Each
codon box consists of four codons that have the same first two letters. The four codons of a
box differ only in their third letter (the wobble position).
UUU UUC UUA UUG Phe Phe Leu UCU UCC UCA UCG Ser Ser Ser Ser UAU UAC UAA UAG Tyr Tyr Stop Stop UGU UGC UGG Cys Cys Trp CUU CUC CUA CUG Leu Leu Leu Leu CCU CCC CCA CCG Pro Pro Pro Pro CAU CAC CAA CAG His His Gln Gln CGU CGC CGA CGG Arg Arg Arg Arg AUU AUC AUA AUG Ile Ile Ile Met ACU ACC ACA ACG Thr Thr Thr Thr AAU AAC AAA AAG Asn Asn Lys Lys AGU AGC AGA AGG GUU GUC GUA GUG Val Val Val Val GCU GCC GCA GCG Ala Ala Ala Ala GAU GAC GAA GAG Asp Asp Glu Glu GGC GGA GGG Gly Gly Gly
U
C
A
G
The Genetic Code was once thought to be “frozen” in that it was considered universal
among all protein-producing systems [67]. However, the sequencing of the human and
bovine mitochondrial DNA in 1981 and 1982, respectively, has proven that the Genetic Code
can deviate from the Code established from Escherichia coli in the 1960s [68, 69]. Soon
after the sequencing of the mammalian mitochondrial DNA, other deviations from the
universal Genetic Code were found [70-72].
In the ribosome, the tRNA must decipher the Genetic Code with great accuracy and
efficiency. tRNA modifications at the anticodon loop have been shown to facilitate accurate
and efficient reading of codons. ASL modifications aid in the recognition of synonymous
codons by restricting codon recognition of split boxes or expanding codon recognition of
whole boxes [1]. Certain ASL modifications also allow the reading of unconventional
codons in the alteration of the universal Genetic Code [73]. Thus, modifications at the ASL
modulate the tRNA’s decoding capacity.
1.3 The effects of tRNA post-transcriptional modifications on synonymous-codon
recognition
The Genetic Code is redundant or degenerate in that there are 61 codons for only 20
amino acids. Codons also outnumber the tRNA species that decode them. In E. coli, there
are 45 tRNA species [74] that must translate the 61 codes; therefore, on average, there are
only some three tRNAs for four codons. Francis Crick’s Wobble Hypothesis explained that a
position (the wobble position) of codon:anticodon binding [75]. Crick hypothesized that the
first two positions of codon:anticodon pairings followed Watson-Crick nucleoside base
pairing rules of Adenosine (A) pairing with Uridine (U) and Guanosine (G) pairing with
Cytosine (C). However, the third position (wobble position) is more flexible in that a G can
pair with its cognate C or “wobble” to a U. Also, Crick explained that Inosine (I), a modified
A, can exhibit a great deal of flexibility by pairing with U, C, or A.
The forty-plus years of research since Crick’s hypothesis has revealed that, although
Crick was fundamentally correct in his assumption of pairing flexibility,
wobble-pairing can be more complicated than just G:U and Inosine wobble-pairings. Oftentimes, the wobble
position of tRNA (position 34, first position of the anticodon) is modified, and these
modifications are significant to tRNA’s ability to wobble to a particular codon (Table 1) [76].
tRNA position 37 is also often modified and aid in codon recognition (Table 1).
Modified nucleosides at the anticodon stem and loop (ASL) of tRNA often give the
tRNA its ability to recognize synonymous codons. Some modifications at the ASL restrict
codon recognition to a few synonymous codons while other modifications serve to extend
codon recognition to include synonymous codons (Table 2). The ability of ASL
modifications to restrict or expand codon reading is due to the extent of degeneracy of the
codons. For example, Lysine (Lys) is two-fold degenerate in that it has two codons, AAA
and AAG. The two Lys codons share a codon box with Asparagine (Asn). There are two
Asn codons – AAU and AAC. Thus, Lys and Asn codons are similar, with the only
The human tRNALys3UUU is modified with 5-methoxycarbonylmethyl-2-thiouridine at
position 34 (wobble position of tRNA) and 2-methylthio-N6-threonylcarbamoyladenosine at
position 37 (tRNALys3UUU-mcm5s2U34, ms2t6A37). Chapter 2 describes how the specific
modifications at the ASL of tRNALys3UUU (ASLLys3UUU-mcm5s2U34, ms2t6A37) serve to limit
codon recognition to only the two Lys codons, AAA and AAG.
On the other hand, Valine (Val) is encoded by four codons. Thus, Val is four-fold
degenerate. All four Val codons reside in the same codon box; therefore, Val codons are
unique from the codons of the other amino acids. Chapter 3 explains how a 5-oxyacetic acid
modification at the wobble Uridine and N6-methyladenosine at position 37 of the anticodon
stem and loop of tRNAValUAC (ASLValUAC-cmo5U34, m6A37) are not required to bind the
tRNA’s cognate codon GUA but required to read the other Val codons of the GUN codon
box. Thus, the modifications of ASLValUAC-cmo5U34, m6A37 allow expansion of codon
recognition. The tRNAValUAC-cmo5U34, m6A37 is an example of one ASL-modified tRNA
that can read the synonymous codons from the same codon box.
tRNA modifications can also serve to expand codon recognition to include
unconventional codons. The Genetic Code varies to a certain extent among certain
organisms and semiautomous organelles. Various codon reassignments can be found in the
translational systems of the semiautomous organelles, mitochondria and chloroplasts, of
different organisms. The mammalian mitochondrial Genetic Code deviates from the
universal Genetic Code in that the universal termination codon UGA codes for Tryptophan;
the two universal Arginine codons AGA and AGG are stop codons, and the universal
In the universal Genetic Code, AUG is the start codon and the sole codon for
Methionine. Genetic translation occurs in three phases: initiation, elongation, and
termination. Initiation is when the mRNA binds the small ribosomal subunit and the initiator
tRNA reads the start codon at the ribosome’s peptidyl (P)-site. Initiation is aided by
initiation factors. After initiation, elongation causes the growth of the polypeptide chain until
completion with the termination step. In the cytoplasm, two tRNA species recognize AUG in
accordance to the codon’s placement on the ribosome. At initiation, AUG is at the P-site of
the ribosome and is recognized by the initiator tRNAMetCAU with the aid of initiation factors.
During elongation, elongator tRNAMetCAU reads AUG at the aminoacyl (A)- site of the
ribosome, facilitated by elongation factors. Mammalian mitochondrial translation differs
from cytoplasmic translation in that 1) there are two Met codons AUG and AUA, 2) only one
tRNAMetCAU recognizes both codons AUG and AUA, and 3) this one tRNAMetCAU acts in both
initiation and elongation. Interestingly, this one mitochondrial tRNAMetCAU is modified at the
wobble position with a 5-formyl group (mtRNAMetCAU-f5C34) [73, 78, 79]. Cytoplasmic
initiator tRNAMetCAU is unmodified at the wobble position. Cytoplasmic elongator
tRNAMet
CAU is modified at the wobble position with a 2’O-methyl group at the ribose moiety. Bacterial initiator tRNAMetCAU is unmodified at the wobble position, while the bacterial
elongator tRNAMetCAU is wobble-modified with N4-acetylcytidine (tRNAMetCAU-ac4C34).
Chapter 4 reports the first chemical synthesis of modified human mitochondrial ASLMet
(hmtASLMetCAU-f5C34) and the significance of the 5-formyl modification in reading the
For some mRNAs in the mitochondria, the universal Isoleucine codons AUU and
AUC are used as initiation codons. Chapter 5 shows how the 5-formyl group enhances
binding to the unconventional Met codon AUA at the ribosomal P- and A-sites and expands
codon recognition to include the unconventional start codons AUU and AUC at the P-site.
Thus, the 5-formyl modification of hmtRNAMetCAU-f5C34 allows codon-reading extension to
include the entire codon box AUN. Furthermore, these results demonstrate how a
modification at the ASL enhances the reading of unconventional codons in the reassignment
Table 2. The recognition of tRNA wobble position-34 for the wobble position (N3) of
the codon. xm5U includes all 5-methylene uridine derivatives, excluding 2-thio; s2U
includes all 2-thiouridine derivatives, and xo5U consists of all 5-oxyuridine derivatives.
a f5C refers to the wobble-modified Cytidine found in mitochondrial tRNAMet.
b Recognition of codon N3 at the P-site.
Anticodon N34 Codon N3
G C, U
C G
I U, C, A
xm5U G
s2U A, G
xo5U A, G, U, C
REFERENCES
1. Agris, P.F., Vendeix, F.A., Graham, W.D. (2007) tRNA’s wobble decoding of the genome: 40 years of modification. J Mol Biol., 366, 1-13.
2. Helm, M. (2006) Post-transcriptional nucleotide modification and alternative folding of RNA. Nucleic Acids Res., 34, 721-33.
3. Nakanishi, K. and Nureki, O. (2005) Recent progress of structural biology of tRNA processing and modification. Mol Cells., 19, 157-66.
4. Björk, G.R., Ericson, J.U., Gustafsson, C.E., Hagervall, T.G., Jönsson, Y.H., Wikström, P.M. (1987) Transfer RNA modification. Annu Rev Biochem., 56, 263-87.
5. Rozenski, J., Crain, P.F., McCloskey, J.A. (1999) The RNA modification database: 1999 update. Nucleic Acids Res., 27, 196-197.
6. Jühling, F., Mörl, M., Hartmann, R.K., Sprinzl, M., Stadler, P.F., Pütz, J. (2009) tRNAdb 2009: compilation of tRNA sequences and tRNA genes. Nucleic Acids Res., 37, D159-62.
7. Björk, G.R. (1995) Biosynthesis and function of modified nucleosides. tRNA: Structure, Biosynthesis, and Function. ASM Press, Washington D.C.
8. Jones, C.I., Spencer, A.C., Hsu, J.L., Spremulli, L.L., Martinis, S.A., DeRider, M., Agris, P.F. (2006) A counterintuitive Mg2+-dependent and modification-assisted functional folding of mitochondrial tRNAs. J. Mol. Biol., 362, 771-86.
9. Nobles, K.N., Yarian, C.S., Guenther, R.H., Agris, P.F. (2002) Highly conserved modified nucleosides influence Mg2+_dependent tRNA folding. Nucl Acids Res., 30, 4751-60.
10. Agris, P.F. (1996) The importance of being modified: roles of modified nucleosides and Mg2+ in RNA structure and function. Prog Nucleic Acid Res Mol Biol., 53, 79-129.
12. Hopper, A.K. and Phizicky, E.M. (2003) tRNA transfers to the limelight. Genes Dev., 17, 162-80.
13. Engelke, D.R. and Hopper, A.K. (2006) Modified view of tRNA: stability amid sequence diversity. Mol Cell., 21, 144-5.
14. Alexandrov, A., Chernyakov, I., Gu, W., Hiley, S.L., Hughes, T.R., Grayhack, E.J., Phizicky, E.M. (2006) Rapid tRNA decay can result from lack of nonessential modifications. Mol Cell, 21, 87-96.
15. Madore, E., Florentz, C., Giegé, R., Sekine, S., Yokoyama, S., Lapointe, J. (1999) Effect of modified nucleotides on Escherichia coli tRNAGlu structure and on its aminoacylation by glutamyl-tRNA synthetase. Predominant and distinct roles of the mnm5 and s2 modifications of U34. Eur J Biochem., 266, 1128-35.
16. Gustilo EM, Dubois DY, Lapointe J, Agris PF. (2007) E. coli glutamyl-tRNA synthetase is inhibited by anticodon stem-loop domains and a minihelix. RNA Biol., 4, 85-92.
17. Sylvers, L. A., Rogers, K. C., Shimizu, M., Ohtsuka, E., Söll, D. (1993) Prevention of mis-aminoacylation of a dual-specificity aminoacyl-tRNA synthetase. Biochemistry, 32:3836–3841.
18. Muramatsu, T., Nishikawa, K.; Nemoto, F.; Kuchino, Y.; Nishimura, S.; Miyazawa, T.; Yokoyama, S. (1988) Codon and amino-acid specificities of a transfer RNA are both converted by a single post-transcriptional modification. Nature, 336, 179–181.
19. Tamura, K., Himeno, H., Asahara, H., Hasegawa, T., Shimizu, M. (1992) In vitro study of E.coli tRNA(Arg) and tRNA(Lys) identity elements. Nucleic Acids Res., 20, 2335–2339.
20. Perret, V., Garcia, A., Puglisi, J., Grosjean, H., Ebel, J.P., Florentz, C., Giegé, R. (1990) Conformation in solution of yeast tRNA(Asp) transcripts deprived of modified nucleotides. Biochimie., 72, 735-43.
21. Lipman, R.S., Wang, J., Sowers, K.R., Hou, Y.M. (2002) Prevention of mis-aminoacylation of a dual-specificity aminoacyl-tRNA synthetase. J Mol Biol., 315, 943-9.
aminoacyl-tRNA: a common molecular mechanism revealed by cryo-EM. EMBO J., 27, 3322-31.
23. Frank, J., Sangupta, J., Gao, H., Li, W., Valle, M., Zavialov, A., Ehrenberg, M. (2005) The role of tRNA as a molecular spring in decoding, accommodation, and peptidyl transfer. FEBS Lett., 579, 959–962.
24. Stark, H., Rodnina, M.V., Wieden, H.J., Zemlin, F., Wintermeyer, W., van Heel, M. (2002) Ribosome interactions of aminoacyl-tRNA and elongation factor Tu in the codon-recognition complex. Nat Struct Biol., 9, 849-54.
25. Agris, P.F. (2008) Bringing order to translation: the contributions of transfer RNA anticodon-domain modifications. EMBO Rep., 9, 629-35.
26. Kurata, S., Weixlbaumer, A., Ohtsuki, T., Shimazaki, T., Wada, T., Kirino, Y., Takai, K., Watanabe, K., Ramakrishnan, V., Suzuki, T. (2008) Modified uridines with C5-methylene substituents at the first position of the tRNA anticodon stabilize U.G wobble pairing during decoding. J Biol Chem., 283, 18801-11.
27. Johansson, M.J., Esberg, A., Huang, B., Björk, G.R., Byström, A.S. (2008) Eukaryotic wobble uridine modifications promote a functionally redundant decoding system. Mol Cell Biol., 28, 3301-12.
28. Yasukawa, T., Kirino, Y., Ishii, N., Holt, I.J., Jacobs, H.T., Makifuchi, T., Fukuhara, N., Ohta, S., Suzuki, T., Watanabe, K. (2005) Wobble modification deficiency in mutant tRNAs in patients with mitochondrial diseases. FEBS Lett., 579, 2948-52.
29. Phelps, S.S., Malkiewicz, A., Agris, P.F., Joseph, S. (2004) Modified nucleotides in tRNA(Lys) and tRNA(Val) are important for translocation. J Mol Biol., 338, 439-44.
30. Agris, P.F. (2004) Decoding the genome: a modified view. Nucleic Acids Res., 32, 223-38.
31. Krüger, M.K., Pedersen, S., Hagervall, T.G., Sørensen, M.A. (1998) The modification of the wobble base of tRNAGlu modulates the translation rate of glutamic acid codons in vivo. J Mol Biol., 284, 621-31.
33. Harrington, K.M., Nazarenko, I.A., Dix, D.B., Thompson, R.C., Uhlenbeck, O.C. (1993) In vitro analysis of translational rate and accuracy with an unmodified tRNA. Biochemistry, 32, 7617-22.
34. Wilson, R.K. and Roe, B.A. (1989) Presence of the hypermodified nucleotide N6-(delta 2-isopentenyl)-2-methylthioadenosine prevents codon misreading by Escherichia coli phenylalanyl-transfer RNA. Proc Natl Acad Sci U S A., 86, 409-13.
35. Smith, D.W. and Hatfield, D.L. (1986) Effects of post-transcriptional base modifications on the site-specific function of transfer RNA in eukaryote translation. J Mol Biol., 189, 663-71.
36. Gilbert, W. (1986) The RNA World. Nature, 319, 618.
37. Wêgrzyn, G. and Wêgrzyn, A. (2008) Is tRNA only a translation factor or also a regulator of other processes? J. Appl. Genet., 49, 115–122.
38. Abbink, T.E. and Berkhout, B. (2008) HIV-1 reverse transcription initiation: a potential target for novel antivirals? Virus Res., 134, 4-18.
39. Raba, M., Limburg, K., Burghagen, M., Katze, J.R., Simsek, M., Heckman, J.E., Rajbhandary, U.L., Gross, H.J. (1979) Nucleotide sequence of three isoaccepting lysine tRNAs from rabbit liver and SV40-transformed mouse fibroblasts. Eur J Biochem., 97, 305-18.
40. Waters, L.C. and Mullin, B.C. (1977) Transfer RNA in RNA tumor viruses. Prog. Nucleic Acid Res. Mol. Biol., 20, 131-160.
41. Gustilo, E.M., Vendeix FA, Agris PF. (2008) tRNA's modifications bring order to gene expression. Curr Opin Microbiol., 11, 134-40.
42. Sakamoto, K., Ishimaru, S., Kobayashi, T., Walker, J.R., Yokoyama, S. (2004) The Escherichia coli argU10(Ts) phenotype is caused by a reduction in the cellular level of the argU tRNA for the rare codons AGA and AGG. J Bacteriol., 186, 5899-905.
43. Thorbjarnardóttir, S., Björnsson, A., Amundadóttir, L., Eggertsson, G. (1991) Temperature sensitivity caused by missense suppressor supH and amber suppressor supP in Escherichia coli. J Bacteriol., 173, 412-6.
45. Garcia, G.M., Mar, P.K., Mullin, D.A., Walker, J.R., Prather, N.E. (1986) The E. coli dnaY gene encodes an arginine transfer RNA. Cell, 45,453-9.
46. RajBhandary, U.L. and Söll, D. (2008) Aminoacyl-tRNAs, the bacterial cell envelope, and antibiotics. Proc Natl Acad Sci U S A., 105, 5285-6.
47. Stewart, T.S., Roberts, R.J., Strominger, J.L. (1971) Novel species of tRNA. Nature, 230, 36-8.
48. Ferber, S. and Ciechanover, A. (1986) Transfer RNA is required for conjugation of ubiquitin to selective substrates of the ubiquitin- and ATP-dependent proteolytic system. J Biol Chem., 261, 3128-34.
49. Scornik, O.A., Ledbetter, M.L., Malter, J.S. (1980) Role of aminoacylation of histidyl-tRNA in the regulation of protein degradation in Chinese hamster ovary cells. J Biol Chem.,255, 6322-9.
50. Schön, A., Krupp, G., Gough, S., Berry-Lowe, S., Kannangara, C.G., Söll, D. (1986) The RNA required in the first step of chlorophyll biosynthesis is a chloroplast glutamate tRNA. Nature, 322, 281-4.
51. Persson, B.C. (1993) Modification of tRNA as a regulatory device. Mol Microbiol., 8, 1011-1016.
52. Kleiman, L., Halwani, R., Javanbakht, H. (2004) The selective packaging and annealing of primer tRNALys3 in HIV-1. Curr HIV Res., 2, 163-75.
53. Kleiman, L., Caudry, S., Boulerice, F., Wainberg, M.A., Parniak, M.A. (1991) Incorporation of tRNA into normal and mutant HIV-1. Biochem Biophys Res Commun., 174, 1272-80.
54. Barat, C., Le Grice, S.F., Darlix, J.L. (1991) Interaction of HIV-1 reverse transcriptase with a synthetic form of its replication primer, tRNA(Lys,3). Nucleic Acids Res., 19, 751-7.
55. Wain-Hobson, S., Sonigo, P., Danos, O., Cole, S., Alizon, M. (1985) Nucleotide sequence of the AIDS virus, LAV. Cell, 40, 9-17.
57. Isel, C., Marquet, R., Keith, G., Ehresmann, C., Ehresmann, B. (1993) Modified nucleotides of tRNA(3Lys) modulate primer/template loop-loop interaction in the initiation complex of HIV-1 reverse transcription. J. Biol. Chem., 268, 25269– 25272.
58. Barat, C., Lullien, V., Schatz, O., Keith, G., Nugeyre, M.T., Grüninger-Leitch, F., Barré-Sinoussi, F., LeGrice, S.F., Darlix, J.L. (1989) HIV-1 reverse transcriptase specifically interacts with the anticodon domain of its cognate primer tRNA. EMBO J., 8, 3279-85.
59. Tamura, F., Nishimura, S., Ohki, M. (1984) The E. coli divE mutation, which differentially inhibits synthesis of certain proteins, is in tRNASer1. EMBO J., 3, 1103-7
60. Garcia, G.M., Mar, P.K., Mullin, D.A., Walker, J.R., Prather, N.E. (1986) The E. coli dnaY gene encodes an arginine transfer RNA. Cell, 45, 453-9
61. Ortwerth, B.J., Lin, V.K., Lewis, J., Wang, R.J. (1984) Lysine tRNA and cell division: a G1 cell cycle mutant is temperature sensitive for the modification of tRNA5Lys to tRNA4Lys. Nucleic Acids Res., 12, 9009-23.
62. Ortwerth, B.J. and Liu, L.P. (1973) Correlation between a specific isoaccepting lysyl transfer ribonucleic acid and cell division in mammalian tissues. Biochemistry, 12, 3978-84.
63. Ortwerth, B.J., Yonuschot, G.R., Carlson, J.V. (1973) Properties of tRNALys4 from various tissues. Biochemistry, 12, 3985-91.
64. Yarian, C., Townsend, H., Czestkowski, W., Sochacka, E., Malkiewicz, A.J., Guenther, R., Miskiewicz, A., Agris, P.F. (2002) Accurate translation of the genetic code depends on tRNA modified nucleosides. J Biol Chem., 277, 16391-5.
65. Krüger, M.K., Pedersen, S., Hagervall, T.G., Sørensen, M.A. (1998) The modification of the wobble base of tRNAGlu modulates the translation rate of glutamic acid codons in vivo. J Mol Biol., 284, 621-31.
66. Urbonavicius, J., Qian, Q., Durand, J.M., Hagervall, T.G., Björk, G.R. (2001) Improvement of reading frame maintenance is a common function for several tRNA modifications. EMBO J., 20,4863-73.
68. Anderson, S., Bankier, A.T., Barrell, B.G., de Bruijn, M.H., Coulson, A.R., Drouin, J., Eperon, I.C., Nierlich, D.P., Roe, B.A., Sanger, F. et al. (1981) Sequence and organization of the human mitochondrial genome. Nature, 290, 457–465.
69. Anderson, S., de Bruijn, M.H., Coulson, A.R., Eperon, I.C., Sanger, F., Young, I.G. (1982) Complete sequence of bovine mitochondrial DNA. Conserved features of the mammalian mitochondrial genome. J Mol Biol., 156, 683-717
70. Ohama, T., Inagaki, Y., Bessho, Y., Osawa, S. (2008) Evolving genetic code. Proc Jpn Acad Ser B Phys Biol Sci., 84, 58-74.
71. Santos, M.A., Moura, G., Massey, S.E., Tuite, M.F. (2004) Driving change: the evolution of alternative genetic codes. Trends Genet., 20, 95-102.
72. Ninio, J. (1990) The revised genetic code. Orig Life Evol Biosph., 20, 167-71.
73. Takemoto, C., Koike, T., Yokogawa, T., Benkowski, L.,Spremulli, L.L., Ueda, T.A., Nishikawa, K., Watanabe, K. (1995) The ability of bovine mitochondrial transfer RNAMet to decode AUG and AUA codons. Biochimie, 77, 104–108.
74. Komine, Y., Adachi, T., Inokuchi, H., Ozeki, H. (1990) Genomic organization and physical mapping of the transfer RNA genes in Escherichia coli K12. J Mol Biol., 212, 579-98.
75. Crick, F.H. (1966) Codon--anticodon pairing: the wobble hypothesis. J Mol Biol., 19, 548-55.
76. Agris, P.F. (1991) Wobble position modified nucleosides evolved to select transfer RNA codon recognition: a modified-wobble hypothesis. Biochimie., 73, 1345-9
77. Jukes, T.H. (1983) Changes in the amino acid code. Adv Space Res., 3, 107-11.
78. Moriya, J., Yokogawa, T., Wakita, K., Ueda, T., Nishikawa, K., Crain, P.F., Hashizume, T., Pomerantz, S.C., McCloskey, J.A., Kawai, G. et al. (1994) A novel modified nucleoside found at the first position of the anticodon of methionine tRNA from bovine liver mitochondria. Biochemistry, 33, 2234–2239.
mitochondrial transfer RNAMet to decode AUG and AUA codons. Biochimie, 77, 104–108.
CHAPTER 2
Modifications of the anticodon stem and loop of human tRNALys (ASLLysUUU) restrict
codon recognition to Lysine codons AAA and AAG.
2.1 Introduction
The universal Genetic Code consists of 64 codons arranged into sixteen codon boxes.
Each codon box is composed of four codons that are similar in their first two letters and
differ only in their third letter. In the ribosome, this third letter of the codon binds to the first
letter of the tRNA anticodon to form the wobble pair. The two modified nucleosides
5-methylaminomethyl-2-thiouridine or a derivative at the wobble position-34 (mnm5s2U34) and
N6-threonylcarbamoyladenosine or a derivative at position 37 (t6A37) are found in almost all
tRNAs that read A or G in the wobble (third) letter of Glutamine (Gln), Lysine (Lys), and
Glutamate (Glu) codons [1-5]. Modifications at the wobble position and position 37 of
tRNAGln, tRNALys and tRNAGlu are also required for efficient aminoacylation by the tRNAs’
cognate synthetases [6-10]. In bacteria, these two modifications within the anticodon loop of
tRNALys (ASLLysUUU- mnm5s2U34, t6A37 or derivatives) have been found to be crucial to the
structure of the anticodon loop, and therefore credited to the tRNA’s ability to perform its
function on the ribosome [4]. The 5-methylaminomethyl modification of tRNALys’s wobble
position 34 (tRNALysUUU- mnm5U34) has been implicated in the recognition of the Lys codons
AAA and AAG, the maintenance of the reading frame, and translocation from the aminoacyl
significantly to the stability of the U:A base pairs, whereas the mnm5-group enhances
recognition of G. E. coli has but one gene for tRNALys and it has the mnm5s2U34UUt6A37
anticodon loop modification. The codon preference is for AAA.
The human tRNALys3 consists of the modifications
5-methoxycarbonylmethyl-2-thiouridine at the wobble position and 2-methylthio-N6-threonylcarbamoyladenosine at
position 37 (tRNALys3UUU-mcm5s2U34, ms2t6A37). Humans have two other Lys tRNAs,
isoacceptors 1 and 2. The anticodon sequence and modifications for tRNALys1, 2 are
C34UUt6A37, which decode the codon AAG. Codon preference in human cells is for AAG.
However, tRNALys3has come under scrutiny for its ability to act as the primer for reverse
transcription of HIV-1 and HIV-2 genomes [13-16]. The details to why tRNALys3UUU
-mcm5s2U34, ms2t6A37 is specifically chosen by the virus are not entirely clear. The
modifications of tRNALys3UUU-mcm5s2U34 are important in HIV’s ability to utilize tRNA Lys3
for annealing to the viral genome and for reverse transcription [17-18].
Derivatives of the 5-methylaminomethyl-2-thio modification at the wobble position
(xm5s2U34) and N6-threonylcabamoyladenosine (xt6U37) at position 37 of tRNA are highly
conserved among organisms of Archea, Bacteria, and Eukarya [1]. Most of our current
understanding of the functions of tRNA modifications is a result of original studies with
bacterial translational systems. Far fewer studies have focused on the effects of
modifications on the functions of tRNAs of higher organisms. Recently, the
5-methoxycarbonylmethyl-2-thiouridine at the wobble position of tRNALys3 (tRNALys3UUU
-mcm5s2U
ms2t6A37 (ASL Lys3UUU-mcm5s2U34, ms2t6A37; Figure 1). As in bacteria, the modifications at
the anticodon loop of tRNA restrict codon binding to the Lys codons AAA and AAG. This
function prevents of binding of the tRNALys3 to the similar Asn codons in the split codon
box.
2.2 Materials and Methods
Ribosomal Binding Assay
The ribosomal binding assays consisted of reaction mixtures of purified E. coli (MRE
600) 70S ribosomes, chemically synthesized mRNAs, and chemically synthesized ASLs.
The primary sequences of the 27-nt mRNA oligo nucleotides were derived from T4 gp32
mRNA [19] and were purchased from Dharmacon (ThermoFisher, Lafayette, CO). In order
to study binding at the ribosomal A-site, the P-site needs to be blocked or occupied.
Therefore, we designed the mRNA with the Methionine (Met) codon AUG at the P-site. The
E. coli tRNAfMet was then used to saturate the P-site. The mRNA sequences were tested for a
low probability of any secondary structure using the program RNA Structure 4.2 [20]. The
mRNA sequences synthesized for our studies were as follows (Lys codons AAA and AAG
are in bold):
1) 5’-GGCAAGGAGGUAAAAAUGAAAGCACGU-3’;
2) 5’-GGCAAGGAGGUAAAAAUGAAGGCACGU-3’.
The 70S ribosomal subunits were isolated as previously described [21]. The ASLs were
5’-end-labeled using [γ-32P] ATP (MP Biomedicals). Unlabeled ASLs in a range of
detectable (2,000-5,000 CPM) of 5’-end, 32P-labeled ASLs in a fixed ratio of unlabeled ASL
to labeled ASL, in order to maintain radiochemical-specific activity. The assay was
performed in ribosomal binding buffer [50 mM HEPES, pH 7.0; 30 mM KCl; 70 mM
NH4Cl; 1 mM DTT; 100 μM EDTA; 20 mM MgCl2). Ribosomes (0.25 μM) were activated
by heating to 42 °C, incubating for 10 minutes and then slowly cooled to 37 °C. The
ribosomes were programmed with 2.5 μM mRNA for 15 minutes at 37 °C. To experiment
with the A-site, the P-site was saturated with E. coli tRNAfMet (Sigma-Aldrich) for 15
minutes at 37 °C. tRNAfMet binds to the Met codon AUG; see underlined codons of the
mRNA sequences above. Binding of ASLLys to the A-site was allowed to proceed for 30 min
at 37 °C. The reaction mixtures (20μL each) were then placed on ice for 20 minutes, diluted
with 100 μL buffer per reaction mixture, and filtered through nitrocellulose in a modified
Whatman Schleicher and Schuell (Brentford, U.K.) 96-well filtration apparatus [22]. Prior to
filtration of experimental samples, the nitrocellulose filter was equilibrated in binding buffer
at 4 °C for at least 20 min and each well of the filtration apparatus was washed with 100 μL
of cold binding buffer. After filtration of reaction samples, each well was then washed twice
with 100 μL of cold ribosomal binding buffer. The nitrocellulose was blotted dry with kim
wipes, and the radioactivity was measured using a phosphorimager (Molecular Dynamics,
GE Healthcare). Data were measured for radioactive intensity using the program ImageQuant
(Amersham). Nonspecific binding was determined by the binding of ASLs to ribosomes
without mRNA and subtracted from the experimental data. The final data is a result of at
least two separate experiments, each done with samples in triplicate, i.e. at least six results
2.3 Results
We have previously assessed the significance of modifications at the anticodon stem
and loop of the bacterial tRNALys (ASLLysUUU-mnm5U34, t6A37) in binding the Lys codons
AAA and AAG [4-5]. While either modification at the wobble position or position 37
enhanced codon binding over the unmodified ASLLys approximately seven-fold, the
modifications in combination increased binding by over ten-fold that of the unmodified
ASLLys. Our experiments with the anticodon stem and loop of human tRNALys3 (ASLLys3UUU
-mcm5s2U34, ms2t6A37) revealed similar results. In comparison to the modified ASLLys3UUU
-mcm5s2U34, t6A37 in in vitro ribosomal binding assays, the unmodified ASLLys3UUU showed
very poor binding to the Lys codons AAA and AAG (Figure 2). The binding of the
unmodified ASLLys to either Lys codon AAA or AAG was nearly undetectable unless
modifications at positions 34 and 37 were present (Figure 2). The modified ASLLys3UUU
-mcm5s2U34, ms2t6A37 bound AAA with a Kd of 3.1 ± 0.4 μM and AAG with a Kd of 3.9 ± 0.8
μM.
2.4 Conclusions
tRNA modifications are highly conserved among all three kingdoms. In split boxes,
the difference between codons of two different amino acids resides in the wobble position
(third letter) of the codon. Because codons in the same box are very similar, recognition of
split boxes must be stringent. tRNAs that decode A or G at the wobble position of split
boxes tend to have the xm5s2U
t6A37. In humans, the tRNA’s ASL is modified with derivatives of the bacterial
modifications: mcm5s2U34 and ms2t6A37. These modifications are been found to be highly
significant to this tRNA’s ability to decode the Lys codons AAA and AAG. Here we report
that the modifications at the anticodon stem and loop of human tRNALys3 (ASLLys3UUU
-mcm5s2U34, ms2t6A37) are required for the anticodon to bind to the Lys codons AAA and
AAG. According to Crick’s Wobble Hypothesis, the unmodified ASLLys3UUU should bind
AAA and AAG because the anticodon UUU is a cognate or Watson-Crick (W-C) pair for
codon AAA and the wobble pair for codon AAG [23]. However, we found that the
unmodified ASLLys showed very low levels of binding in in vitro ribosomal binding assay.
Only with the addition of the naturally-occurring modifications
5-methoxycarbonylmethyl-2-thiouridine to the wobble position and 2-methylthio-N6-threonylcarbamoyladenosine to
position 37 (ASLLys3UUU-mcm5s2U34, ms2t6A37), does the ASL bind both codons AAA and
AAG. Considering their significance to tRNALys3’s central role of decoding, modifications at
the ASL of human tRNALys3 may be have implications on other functions of the tRNALys,
such as its involvement in retroviral replication.
In addition to the quantitative results of codon binding reported here we now have the
NMR solution structure of the ASLLys3 and the X-ray crystallographic structures of the
ASLLys3UUU-mcm5s2U34, ms2t6A37 bound to both Lys codons AAA and AAG. A comparison
of the solution structure to that of the structure of the ribosomal X-ray indicates that the
modifications have restricted motional dynamics that directs the structure toward that
required for codon binding. These results are comparable to structures of bacterial ASLLys
Figure 1. The anticodon stem and loop of human tRNALys3 (ASLLys3UUU-mcm5s2U34,
ms2t6A
37). The ASL is modified at the wobble position-34 with
5-methoxycarbonylmethyl-2-thiouridine, at position 37 with 2-methylthio-N6-threonylcarbamoyladenosine, and at
position 39 with a pseudouridine.
U
Ψ A
C G A U G C A Ψ
A C
U
U U C G A G
A Ψ39
C
U msms22tt66A37
mcm 5s2U
34 U U
G C
Pseudouridine,Ψ
5‐methoxycarbonylymethyl‐2‐thiouridine,
mcm5s2U
2‐methylthio‐N6‐threonylcarbamoyladenosine, ms2t6A A A A
G A A 5’
5’ 3’
Figure 2. Ribosomal, equilibrium binding curves of the fully-modified human
ASLLys3UUU- mcm5s2U34, ms2t6A37 (▲) or the unmodified ASLLys3UUU (■). E. coli 70S
ribosomes were programmed with the Lysine codon A. AAA or B. AAG at the A-site. The
P-site was saturated with E. coli tRNAfMet, which binds to its cognate Methionine codon
AUG.
0
1
2
0
1
2
0
25
50
75
100
Bindi
n
g (
%
)
ASL
Lys(
μ
M)
REFERENCES
1. Jühling, F., Mörl, M., Hartmann, R.K., Sprinzl, M., Stadler, P.F., Pütz, J. (2009) tRNAdb 2009: compilation of tRNA sequences and tRNA genes. Nucleic Acids Res. 37:D159-62.
2. Agris, P.F., Vendeix, F.A., Graham, W.D. (2007) tRNA’s wobble decoding of the genome: 40 years of modification. J Mol Biol., 366(1), 1-13.
3. Björk, G.R., Huang, B., Persson, O.P., Byström, A.S. (2007) A conserved modified wobble nucleoside (mcm5s2U) in lysyl-tRNA is required for viability in yeast. RNA, 13(8):1245-55.
4. Murphy, F.V. 4th, Ramakrishnan, V., Malkiewicz, A., Agris, P.F. (2004) The role of modifications in codon discrimination by tRNA(Lys)UUU. Nat Struct Mol Biol., 11(12):1186-91.
5. Yarian, C., Marszalek, M., Sochacka, E., Malkiewicz, A., Guenther, R., Miskiewicz, A., Agris, P.F. (2000) Modified nucleoside dependent Watson-Crick and wobble codon binding by tRNALysUUU species. Biochemistry, 39(44):13390-5.
6. Gustilo, E.M., Dubois, D.Y., Lapointe, J., Agris, P.F. (2007) E. coli glutamyl-tRNA synthetase is inhibited by anticodon stem-loop domains and a minihelix. RNA Biol., 4(2):85-92.
7. Madore, E., Florentz, C., Giegé, R., Sekine, S., Yokoyama, S., Lapointe, J. (1999) Effect of modified nucleotides on Escherichia coli tRNAGlu structure and on its aminoacylation by glutamyl-tRNA synthetase. Predominant and distinct roles of the mnm5 and s2 modifications of U34. Eur J Biochem., 266(3):1128-35.
8. Sylvers, L. A., Rogers, K. C., Shimizu, M., Ohtsuka, E., Söll, D. (1993) Prevention of mis-aminoacylation of a dual-specificity aminoacyl-tRNA synthetase. Biochemistry, 32:3836–3841.
9. Muramatsu, T., Nishikawa, K.; Nemoto, F.; Kuchino, Y.; Nishimura, S.; Miyazawa, T.; Yokoyama, S. (1988) Codon and amino-acid specificities of a transfer RNA are both converted by a single post-transcriptional modification. Nature, 336:179–181.
11. Phelps, S.S, Malkiewicz, A., Agris, P.F, Joseph, S. 2004. Modified nucleotides in tRNA(Lys) and tRNA(Val) are important for translocation. J Mol Biol., 338(3):439-44.
12. Urbonavicius, J., Qian, Q., Durand, J.M., Hagervall, T.G., Björk, G.R. (2001) Improvement of reading frame maintenance is a common function for several tRNA modifications. EMBO J., 20(17):4863-73.
13. Kleiman L, Halwani R, Javanbakht H. (2004) The selective packaging and annealing of primer tRNALys3 in HIV-1. Curr HIV Res., 2(2):163-75.
14. Kleiman L, Caudry S, Boulerice F, Wainberg MA, Parniak MA. (1991) Incorporation of tRNA into normal and mutant HIV-1. Biochem Biophys Res Commun., 174(3):1272-80.
15. Barat C, Le Grice SF, Darlix JL. (1991) Interaction of HIV-1 reverse transcriptase with a synthetic form of its replication primer, tRNA(Lys,3). Nucleic Acids Res., 19(4):751-7.
16. Wain-Hobson S, Sonigo P, Danos O, Cole S, Alizon M. (1985) Nucleotide sequence of the AIDS virus, LAV. Cell, 40(1):9-17.
17. Bilbille, Y., Vendeix, F.A., Guenther, R., Malkiewicz, A., Ariza, X., Vilarrasa, J., Agris, P.F. (2009) The structure of the human tRNALys3 anticodon bound to the HIV genome is stabilized by modified nucleosides and adjacent mismatch base pairs. Nucleic Acids Res., In press.
18. Isel, C., Marquet, R., Keith, G., Ehresmann, C., Ehresmann, B. (1993) Modified nucleotides of tRNA(3Lys) modulate primer/template loop-loop interaction in the initiation complex of HIV-1 reverse transcription. J Biol Chem., 268(34):25269-72.
19. Fahlman, R.P., Dale, T., and Uhlenbeck, O.C. (2004) Uniform binding of aminoacylated transfer RNAs to the ribosomal A and P sites. Mol. Cell, 16: 799– 805.
20. Mathews, D.H., Turner, D.H., and Zuker, M. (2007) RNA secondary structure prediction. Curr. Protoc. Nucleic Acid Chem., Chapter 11: Unit 11 2.
22. Wong, I. and Lohman, T.M. (1993) A double-filter method for nitrocellulose-filter binding: application to protein-nucleic acid interactions. Proc. Natl Acad. Sci. USA, 90:5428–5432.
Chapter 3
Anticodon Domain Modifications Contribute Order to tRNA for Ribosome-Mediated Codon Binding
Franck A. P. Vendeix, Agnieszka Dziergowska, Estella M. Gustilo, William D. Graham, Brian Sproat, Andrzej Malkiewicz, and Paul F. Agris.
Department of Molecular and Structural Biochemistry, North Carolina State University, 128 Polk Hall, Raleigh, North Carolina 27695-7622, Institute of Organic Chemistry, Technical University, Zeromskiego 116, 90-924 Lŏdű, Poland, and Integrated DNA Technologies BVBA, Provisorium 2, Minderbroedersstraat 17-19, B-3000 Leuven, Belgium
Anticodon Domain Modifications Contribute Order to tRNA for
Ribosome-Mediated Codon Binding†,‡
Franck A. P. Vendeix‡, Agnieszka Dziergowska§, Estella M. Gustilo‡, William D. Graham‡,
Brian Sproat║, Andrzej Malkiewicz§, and Paul F. Agris*,‡
Department of Molecular and Structural Biochemistry, North Carolina State University, 128
Polk Hall, Raleigh, North Carolina, USA 27695-7622, Institute of Organic Chemistry,
Technical University, Żeromskiego 116, 90-924, Łódź, Poland, and ║Integrated DNA
Technologies BVBA, Provisorium 2, Minderbroedersstraat 17-19, B-3000 Leuven, Belgium
Running Title: Wobble Modifications Order tRNA's Anticodon
†This work was supported by grants from the National Institutes of Health (Grant
2-RO1-GM23037 to PFA), the National Science Foundation (MCB-0548602 and 2-RO1-2-RO1-GM23037
to PFA), RNA-TEC NV, and the Polish Ministry of Science and Education (to AM).
*To whom correspondence should be addressed.
Phone: 1-919-515-6188.
E.Mail: Paul_Agris@ncsu.edu
FAX: 1-919-515-2047.
‡North Carolina State University
║Integrated DNA Technologies
1Abbreviations: ASL, anticodon stem and loop domain; ASLVal3
UAC, ASL of E. coli's valine tRNA isoaccepting species 3 with anticodon UAC; CD, circular dichroism
spectropolarimetry; cmo5U34, uridine-5-oxyacetic acid; HPLC, high performance liquid
chromatography; m6A37, N6-methyladenosine-37; NOE, nuclear Overhauser effect; Tm,
ABSTRACT
The accuracy and efficiency with which tRNA decodes genomic information into
proteins require posttranscriptional modifications in or adjacent to the anticodon. The
modification uridine-5-oxyacetic acid (cmo5U34) is found at wobble position 34 in a single
isoaccepting tRNA species for six amino acids, alanine, leucine, proline, serine, threonine,
and valine, each having 4-fold degenerate codons. cmo5U34 makes possible the decoding of
24 codons by just six tRNAs. The contributions of this important modification to the
structures and codon binding affinities of the unmodified and fully modified anticodon stem
and loop domains of tRNAVal3UAC (ASLVal3UAC) were elucidated. The stems of the
unmodified ASLVal3UAC and that with cmo5U34 and N6-methyladenosine, m6A37, adopted an
A-form RNA conformation (rmsd 0.6 Å) as determined with NMR spectroscopy and
torsion-angle molecular dynamics. However, the UV hyperchromicity, circular dichroism
ellipticity, and structural analyses indicated that the anticodon modifications enhanced order
in the loop. ASLVal3UAC-cmo5U34;m6A37 exhibited high affinities for its cognate and wobble
codons GUA and GUG, and for GUU in the A-site of the programmed 30S ribosomal
subunit,whereas the unmodified ASLVal3UAC bound less strongly to GUA and not at all to
GUG and GUU. Together with recent crystal structures of ASLVal3UAC-cmo5U34;m6A37 bound
to all four of the valine codons in the A-site of the ribosome’s 30S subunit, these results
clearly demonstrate that the xo5U34 -type modifications order the anticodon loop prior to
A-site codon binding for an expanded codon reading, possibly reducing an entropic energy
INTRODUCTION
Transfer RNA is one of the most understood biological macromolecules. The
relationship of its nucleoside chemistry and oligonucleotide structure to its functions in
protein synthesis has been studied extensively (1, 2). Those studies have led to the
identification of more than 70 different posttranscriptional modifications present in tRNAs
(3). tRNA modifications increase stability (4), enhance decoding (5), restore ribosomal
binding (6), and influence reading frame maintenance (7, 8). In the course of translation,
anticodon domain modifications in particular play important roles in the accuracy and
efficiency of protein synthesis. Therefore, the modifications that occur at anticodon wobble
position 34, and at the conserved purine 37, 3′-adjacent to the anticodon, are of obvious
interest in tRNA’s decoding of mRNA (Figure 1). These modifications play critical and
distinctive roles in tRNA’s accurate and efficient binding of cognate and wobble codons
within the ribosome’s A-site (9).
Some 40 years ago, Francis Crick explained how a limited number of tRNAs could
decode the 61 amino acid codons (10). Our view of codon recognition by tRNA was then
altered in the modified wobble hypothesis to accommodate new information about
modifications (11). The limited number of tRNAs requires most tRNAs to read more than
one codon. Some tRNAs respond to codons in “mixed” codon boxes where distinction of the
third codon base (the most degenerate of coding positions) is important for discriminating
between the correct cognate or wobble codons and the incorrect but near-cognate codons. For
example, a wobble position, 2-thiouridine 34 (s2U
12). In contrast, other modification chemistries expand wobble codon recognition, such as
tRNA’s wobble position inosine 34 (I34) that will bind codons ending in A3, C3, or U3 (12).
Modifications expand tRNA’s codon recognition beyond that envisioned in the wobble
hypothesis. Crick suggested that a U34 would pair with an A and wobble to a G in the third
position of the codon (A3 or G3) but would not base pair with a U3 or C3. He argued that a
U34 paired with a U3 or C3 would markedly distort the anticodon-codon minihelix (10).
However, wobble position U34 of some tRNA species will base pair with a U3 and even a
C3 in recognizing all of the synonymous codons of a 4-fold degenerate codon box (9, 13–
16). For tRNAs to recognize codons ending with a U3 or C3, as well as codons ending with
A3 and G3, modification of wobble position U34 is essential (9, 13–16). The modified
nucleoside capable of binding to A, G, U, and C is uridine-5-oxyacetic acid, cmo5U341
(Figure 1). Each of the six amino acids, alanine, leucine, proline, serine, threonine, and
valine, having 4-fold degenerate codons, has a single isoaccepting tRNA species containing
the modification cmo5U34 (17), but would a tRNA species with cmo5U34 be sufficient for cell
viability in the absence of all other isoaccepting tRNAs in vivo? The function of cmo5U34 in
tRNAPro was analyzed by introducing null copies of the two genes (cmoA and cmoB)
identified as part of the cmo5U34 synthetic pathway into a strain of Salmonella having only
one of three tRNAPro isoacceptors, that with the wobble cmo5U34 modification (18). Growth
of this and other mutant strains demonstrated that all four proline codons (CCU/C/A/G) were
read by the cmo5U-containing tRNAPro, and that the complete modification was critical for
reading codons ending in U and C (18). However, the physicochemical contributions of
![Table 1. Distribution of modified nucleosides (from Auffinger, P. and Westhof, E.[80])](https://thumb-us.123doks.com/thumbv2/123dok_us/1645236.1205778/23.612.96.496.98.563/table-distribution-modified-nucleosides-auffinger-p-westhof-e.webp)