0022-538X/78/0026-0001$02.00/0
Copyrighti 1978 AmericanSociety for Microbiology Printed in U.S.A.
Resolution
and
Characterization
of
Intracytoplasmic
Forms
of
Reverse
Transcriptase from Rauscher Leukemia
Virus-Producing
Cells
STUART L. MARCUS
MemorialSloan-Kettering Cancer Center, New York, New York10021
Received forpublication 9 November 1977
Themicrosomalsupernatantfraction obtained from a murinecellline
chroni-cally infected with and producing Rauscher leukemiavirus (JLSV-10)was found tocontaintwoforms of RNA-directed DNA polymerase (reverse transcriptase).
Thetwo enzyme forms, neither ofwhich is detectable in uninfected cells
(JLSV-9),wereinitiallypartiallypurified bypoly(C)-agarosechromatography, and their
separationwasachievedby phosphocellulose chromatography. Theenzymeform
elutingfirst fromphosphocellulose (0.3 MKCl),designated PCI, was found to be
identical in all parameters tested to that form isolated directly from purified
virions. The second enzymepeak, designated PC II, eluted from
phosphocellu-loseat0.5M KCI andwas notdetectable inpurified virions. The PC II enzyme
hasamolecular weight, determinedby velocitysedimentation, of approximately
109,000,ascomparedwith 70,000for the PC Ienzyme,andcouldnotbe further
dissociatedbyexposure tohighsaltornonionicdetergent. Mixing purifiedvirion
orPCI DNApolymerase with uninfected cells followed by fractionationdid not
produce the PCIIform, suggesting that it is neitheranartifactof purification nor
theresult of fortuitouscomplexingofreversetranscriptase with nornal cellular component(s). Both PCIandPCII enzymeforms appearedantigenically similar
tovirionDNApolymerase,demonstratedidentical divalent cationrequirements
forvarious template-primers, and were capableofcopying heteropolymeric
re-gions of rabbit globin mRNA. However, kinetic studies of heat inactivation
revealed that the PC II enzyme wasfarmore heat labile than the PC I form,
which appeared identical to the virion enzyme in this respect. Furthermore, whereas the PC I and virion-derived reverse transcriptase copied
poly(C)-(dG)12-18mostefficientlyat atemplate-to-primer molar nucleotide ratio of 25:1,
the PCII enzymepreferredaratio of5:1foroptimalratesofpoly(dG) synthesis.
Therefore, by these criteria, there appear to exist two intracellular fonms of
reversetranscriptaseinthe JLSV-10 Rauscher leukemiavirus-producingmurine
cell line.
RNA-directed DNA polymerase (reverse onthe
purification
ofreversetranscriptase
fromtranscriptase)has been showntoberequiredfor cells chronically infected with retrovirus (1, 4,
the productive infection and transformation of 8). Such studiesareneededto
identify
andchar-cellsbyavian retroviruses(17,31).Reversetran- acterize the form(s) of virion-related DNA
po-scriptase appears to be
required
for the initia- lymeraseinsuchcellsand also forthedevelop-tion,
although
notthemaintenance,
ofthe vir- mentofprocedures
toidentify
theseenzymesinally transformed cell state (13, 29). Antibiotics human tumors, where retroviral involvement that inhibit the reverse transcriptase of has beensuggested (10).In thispaper,thepartial
Rauscher leukemia virus have been shown to purification and biochemical characterization of
preventtheleukemogenic activity of thatvirus twoforms ofreverse
transcriptase
fromthecy-(34), suggestingthat this enzymeplaysadirect toplasm ofRauscher leukemia
virus-producing
role in the induction of murine leukemia by cellsaredescribed.
exogenous virus. Anumberofreportsare
avail-ablein theliteratureconcerningthepurification MATERIALS AND METHODS
andbiochemical characterizationofmammalian Reagents.Allradioactivedeoxyribonucleoside tri-retroviralDNApolymerase (6,11, 12, 14, 23,26, phosphateswerepurchasedfrom Amersham/Searle.
30,32). Littleinformation, however, isavailable Unlabeled triphosphatesand template-primerswere 1
on November 10, 2019 by guest
http://jvi.asm.org/
2
obtained from P-L Biochemicals. The molar ratio of volumes of ice-cold disruption buffer consisting of template to primer, unless otherwise indicated, was Tris-hydrochloride (pH 7.8), 1 mM dithiothreitol, 1 1:1 in the case ofpoly(A) (dT)12-,8and 20:1 in the case mMMgCl2,and 15%glycerol.Disruption was carried ofpoly(C) *(dG)12.18.Activated DNA was prepared as out with 40 strokes of a loose-fitting Dounce homog-describedbyAposhianandKornberg (2).Rabbitglo- enizer. Nuclei were removed by a 1,000 x g centrifu-bin mRNA was purchased from Amersham/Searle gation step carried out at4°C, and the resultingpellet and annealed tooligo(dT)12inthemolar ratio of 20:1 was discarded. The 1,000 x gsupernatant was sub-in 0.05 M Tris-hydrochloride, pH 7.8 (20). Starter jected to a10,000xgcentrifugationat40Cfor20min cultures of the non-virus-producing, continuous, to removemitochondria. Thesupernatantfrom that mouse bone marrow-derived fibroblastcelliine JLSV- step was then spun in an SW41 rotor for 1 h at 100,000 9 (3), as well as one chronically infected with and xg. Thepelletfraction thus obtained was suspended
actively producing Rauscher leukemia virus (JLSV- in1ml ofpoly(C)-agarosecolumnequilibrationbuffer,
10), were thegiftof Peter J.Gomatos of thisinstitute. and disruption buffer was added as previously de-Purified Rauscher leukemia virus and ahomogenous scribed forpurifiedRauscher leukemia virus(23). Pu-preparation ofEscherichia coli DNA polymerase I rification ofreverse transcriptase from this fraction
wereprovided byM. J.Modak of this institute. Puri- wasalsoaccomplishedaspreviouslydescribed(23)by
fied immunoglobulin G (IgG) preparations prepared using poly(C)-agarose affinity chromatography. The
frompreimmunized rabbit serum and fromantisera 100,000xgsupernatantfraction obtained from
JLSV-directedagainstpartiallypurifiedRauscherleukemia 9 or JLSV-10 cells was placed either directly onto
virus DNApolymerasewerethe kindgiftofCharles poly(C)-agaroseorafterthefollowingadditionswere
J. Sherr of the National Cancer Institute. DEAE- madetothat fraction: 0.05% (vol/vol)NonidetP-40, cellulose and phosphocellulose were obtained from 0.02%(wt/vol) sodiumdeoxycholate,and 0.04 MKCI. Whatman, Inc. Afterthe columnswerewashed with 10mil ofbuffer,
Preparationofpoly(C)-agarose.Poly(C)wasob- enzymewaseluted withalinear saltgradient(0to0.6
tained from P-L Biochemicals and was covalently M KCI)in wash buffer. Trasylol (Aprotinin), a
pro-coupledto agarosebeads (Sepharose 4B, Pharmacia tease inhibitor obtained fromMobay Chemical Co.,
Fine Chemicals, Inc.) througha modificationof the when usedwaspresent in the celldisruptionbufferat
original procedure (19).Cyanogenbromidewasadded aconcentration of 100U/ml. All columnswere run at
toSepharoseforactivation after firstbeingdissolved aflowrateof18ml/hat 4°C unlessotherwise
indi-inN,N-dimethylformamide (2ml ofsolvent for1g of cated. Fractions of 1-ml volumewerecollected.
BrCN).When thisprocedurerather than the addition Enzyme fractions that were obtained by elution
offinely dividedBrCN isused, the activationtime is from poly(C)-agarose weremade 0.5 M inKCI and
decreasedto 10min.All other stepsarethe same as passedthrough2-ml-bedvolumeDEAE-cellulose
col-previously described for coupling of poly(C)(19).After umns to remove any poly(C) that may have been
coupling,glycinewasadded to a final concentration of degraded during chromatography. The enzyme
frac-0.1 M tothepoly(C)-agarosesuspension,andstirring tions were thendiluted fivefold withpoly(C)-agarose wascontinued at4°Cfor2to4htoallow thecomplete column wash buffer andappliedto phosphocellulose substitution of activated groups thatmightbeleft on columns (0.9 by10cm) previously equilibrated with the agarose. Without suchblocking ofunsubstituted column wash buffercontaining0.1MKCI. Elution of
sites, someenzyme protein together with other pro- enzyme activity from phosphocellulose was
accom-teins may beinadvertentlycovalently coupledtothese plished (after columns were washedwith 10 ml of sitesduringthe course ofpurification, resultingin a buffer)with alinear saltgradient(0.1 M KCI to 0.8 M
loweringofyields. It isalso important to notethat KCI)inatotal volume of 60 ml. Peak enzyme fractions
during long periods of storage of poly(C)-agarose, wereusedimmediatelyorstored at0°Cinthe presence
traces of RNase may act to produce breaks in the of 100 ug ofbovineserum albumin monomer (Miles
polynucleotidestrands. Toremovefreepoly(C)from Laboratories, Inc.)per ml.
thematrix,columns should first be washed with 1M DNApolymerase assays.DNApolymerase
activ-KCI beforeequilibrationwith buffer.Poly(C)-agarose itywasassayedaspreviouslydescribed(23).Annealing columnswerepreparedwith1.5-to2-mlbedvolumes of varioustemplateandprimer combinations used in in Pasteurpipettesandequilibratedat4°C with buffer this study, both synthetic and natural in origin, was consisting of50 mM Tris-hydrochloride (pH 7.8), 1 carried out as described by Modak and Marcus (23).
mM dithiothreitol, and 10% (vol/vol) glycerol. We Molecular weight estimation. Sedimentation
haveshown thatpoly(C)-agaroseisaneffectiveaffin- coefficients were determined by centrifugation
ity matrix for the one-step purification of reverse through preformed 10 to 30%(vol/vol) linearglycerol
transcriptase from avian myeloblastosis virus (19) and gradienta in 0.05 MTris-hydrochloride buffer (pH 7.8) Rauscher leukemia virus(23) and for the partial pu- containing 1 mM dithiothreitol and 0.4 MKCI.DNA rification of murinemammary tumor virus DNA po- polymerase activity from peak poly(C)-agarose or lymerase (21). phosphocellulosecolumn fractions was diluted 1:1 in
Cell fractionation and enzyme purification. the same buffer used to prepare the glycerolgradients
CelLswere grown at 370C in plastic (Corning Glass and layered over5-ml gradients. Centrifugation was
Works) orglassrollerbottles in Eagle minimal essen- carried out in anSW50.1 rotor at 48,000 rpm for 16 h tial medium (5) supplemented with 10% fetal calf at4°C.Fractions were thencollectedfrom the bottom serum.Cellswereharvested inmid-logphase by scrap- of the tube and assayed for DNA polymerase activity ing and washed twice in phosphate-buffered saline. with poly(A) (dT)1218. Parallel gradients were run
Thecells(5 g[wetweight]) were then suspended in 5 with E. coli DNA polymerase I and purified
on November 10, 2019 by guest
http://jvi.asm.org/
derived Rauscher leukemia virus DNA polymeraseas 9 cells contained no poly(rCm)
(dG)12.18-di-molecular weight markers. The marker gradientswere rected synthesizing activity and were not
frac-assayedaspreviouslydescribed (19, 23). tionated
further
(data notshown).
The relative Inhibition of DNA polymeraseactivitybyIgG recoveries of DNA polymerase activity from addition.Enzymefractions (10pl)
wereincubatedatviral
andcytoplasmicsourcesare
listed
inTable
00Cfor1h in atotal reaction volume of 35pl
contain-ing 85 mM Tris-hydrochloride (pH 7.8), 100 mMKCI, 1. 1.7 mM dithiothreitol, 0.85 mM MnCl2, 0.085%
(vol/vol) Triton X-100, and the desired quantity of 5 immune or preimmuneIgG. Poly(A) *(dT)1218(0.5 ug) 40 _ A
and 5,uM[3H]dTTP (4,000 cpm/pmol) were added in
A30
0.6
a
25Spl
volume after preincubation, and polymerization Iwas initiated by placing the reaction mixtures in a v 20 0.4
370Cbathfor 60 min. Thequantitationofincorporated 10-
'02
[3H]dTMP was carried out as described above for ____
DNApolymerase assays. x
RESULTS x
Purification of virion and microsomal 2 8
membrane pellet reverse transcriptase. Y?
Figure1shows theelutionprofilesof virion and e Y 6_ -0.6
JLSV-10 microsomal membrane pellet fraction , 0.5
DNA polymerase from poly(C)-agarose col- i 4 -0.4 T
umna. The template-primer poly(rCm).
(dG)12.
- 0.4is,whichhas been reported specificforthe de- -> 0.3
tectionof reversetranscriptase activity (4),was 2 0.2
included in assays of column eluate fractions V
/._
from the cellular preparations. The
poly(A) *(dT)12.18-utilizingactivitycoeluted with 4 8 12 16 20 24 2 32 36 40
poly(rCm)*
(dG)12.i8-utilizing
activity from the 12 numbercytoplasmic pelletfraction at the same salt
con-Fracton
numbr
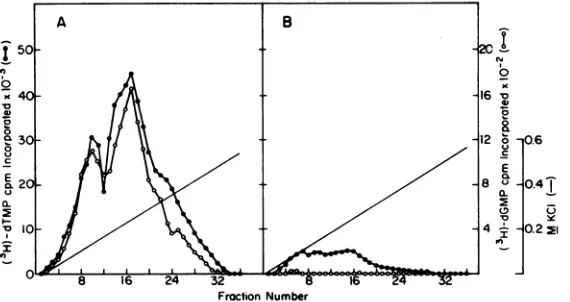
centration (0.24 M KCl) as did virion reverse FIG. 1. Elution from poly(C)-agarose columns of
transcrnptase. No terinal deoxynucleotidyl DNApolymeraseactivities from(A)
Rauscher
leuke-transferase activity was observed. F urther tud- mia
virions
and(B)
the microsomalmembranepeUet
ies (see below) revealed that the microsomal fraction derived fromprepared and columns
JLSV-10
were runcell& Fractionsas described inwerepellet DNA polymerase fromJLSV-10 cellspur-
the
text.Portions(20 fd)fromeachfraction
wereas-ified in thismannerpossessed propertiesidenti- sayed forpoly(A)
(dT)1248-directed
(0)orpoly(rCm) calto those of the virion enzyme. Pellets pre-(dG)i2-18directed
(0) or terminaldeoxynuclkotid-paredin a similar fashionfrom uninfectedJLSV- yltransferaseactivity(A)asdescribedinthe text. TABLE 1. Recovery ofpoly(A)*(dT)12i18andpoly(rCm) (dG)121s-directedDNAsyntheticactivityfromviral
and cellular sourcesafterpoly(C)-agarosechromatographya
Unitsof enzymeactivity'(x10-3)in:
Enzymesource Crude fractions Poly(C)-agarosecol- Poly(C)-agarosecol- Activityrecovered in
umnwash fractions umneluate fractions eluatefractions(%)
dTMP dGMP dTMP dGMP dTMP dGMP dTMP dGMP
Rauscher leukemia virus 160 10.1 15 0.9 115 7.5 72 75
(100Ag of viral protein)
JLSV-10 microsomal pellet 124 8.1 20.4 0.8 96 6.04 69 74
JLSV-10 microsomal super- 26.7 1.05 13.6 0.15 9.1 0.71 34 71
natant
JLSV-9 microsomal super- 6.0 NDC 5.6 ND 1.2 ND 20
natant
aApproximately5g(wetweight)ofeitherJLSV-9orJLSV-10celLswasharvested andprocessedas
described
inthe text, and the various fractionswerechromatographedonpoly(C)-agarose. Assaysfor DNAsynthetic activity directedbypoly(A)-(dT)12-1s (expressedasdTMPunits)orpoly(rCm)-(dG)12-18 (expressedasdGMP
unitsWcontained0.5mMMnCI2and50mMKCI,in additiontothe standardcomponentsdescribed in thetext. bOneunit ofactivityisdefinedasthat amountof enzymeincorporating1pmolof tritiated substrate into
acid-insolublematerialduring30 min of incubationat370C.
cND, Not detectable(<0.05pmolofdGMPincorporatedper
20-pd
portionunder assayconditions).on November 10, 2019 by guest
http://jvi.asm.org/
[image:3.504.257.447.140.367.2]4 MARCUS J. VIROL.
ResolutionofDNApolymerase activities phosphocellulose chromatograms of
poly(rC)-inJLSV-10high-speed cytoplasmic super- agarosecolumnpeaks corresponding to fractions
natant.Cellsweredisruptedasdescribed above 8to 12inclusive and15 to 19inclusive in Fig. 2A and in as gentleamanneras possible to avoid are shown in Fig. 3A and B, respectively. The
thepossibledisruption of enzyme complexes by secondarypeak of activity from poly(C)-agarose
detergent treatment(4). Nuclei were not used in (fractions 8 to 12) was resolved in this manner thefractionation procedure, since the supposed into two distinct peaks of roughly equivalent site of action andsynthesisof viral DNApolym- activity eluting from phosphocellulose at 0.3 M erase is thecytoplasm (12). In contrast to the and 0.5 M KCl. The primary peak of activity sharp, single-peak elution profiles shown for from poly(C)-agarose (fractions 15 to 19, Fig.
DNApolymerasefromRauscher leukemia virus 2A)elutedfromphosphocellulose as one major
and the JLSV-10 microsomal pellet
enzyme
(Fig. peak of activity at 0.3 M KCl (Fig. 3B), as did1),elutionprofiles showing twopeaksof DNA the poly(C)-agarose peak reverse-transcriptase
polymerase activitywerereproduciblyobtained activityderived from either Rauscher leukemia
fromcytoplasmichigh-speedsupernatant chro- virus orJLSV-10 microsomal membranepellet
matographedonpoly(C)-agarose (Fig. 2A). The (data notshown). Therefore, the high-speed
cy-additionofdetergent(seeabove)toapproximate toplasmicsupernatantfrom Rauscher leukemia
conditionsused forchromatographyofdisrupted virus-producing cells appeared to contain two virions did notsignificantly alter the observed chromatographically separable forms of reverse elution profile. The primary peak of transcriptase, whereas the virion and
micro-poly(A)*(dT)12 18- andpoly(rCm)*(dG)12.18-uti- somalpelletfractionsappearedtocontain only
lizingactivityelutedat0.24 MKCI(asdoes the oneform of the enzyme. To determine whether
virion enzyme), whereas the secondary peak the two peaks of activity resolved by phospho-elutedat0.12 to 0.15MKCl. Chromatography cellulose chromatography were due to
artifac-ofJLSV-9 cytoplasmic high-speedsupernatant tual complexing of the virion enzyme with a
on poly(C)-agarose yielded the elution profile solublecytoplasmicconstituent, virion or
micro-shown inFig.2B. Smallamounts (seeTable1) somal pellet-derived enzyme was mixed with
ofpoly(A)*(dT)12.18-utilizing
activity
wereseen virus-freeJLSV-9 cellhomogenates,whichweretoelute from 0.1 Mto0.22 MKCl,representing then fractionated in a manner identical to that
5 to10% of theactivityobserved in the JLSV-10 describedabove.Onlyonepeakof
reverse-tran-column eluates. No
poly(rCm)
*(dG)12
18activity scriptase activity, eluting at 0.3 M KCI, waswasobservedinJLSV-9column eluate fractions. resolved by phosphocellulose chromatography
The twopeaksofreverse-transcriptase activity in thismanner(datanotshown),suggesting that
obtainedfromvirus-producingcellswerepooled the0.5MKCIpeak activityfromJLSV-10cells
separately andfurther
analyzed by chromatog-
was indeedaform restricted tocellsproducingraphy on
phosphocellulose
after passage virus. Further biochemical analysis of the twothrough DEAE-cellulose to remove
possible
forms ofreversetranscriptase, designated PC Itraces of poly(C) (see above). Representative (0.3 M
KCl)
and PC II(0.5MKCl)correspond-A B
50 C
0 6
40- -~~~~~~~~~~~~~~~~~~~~16v
eo 30- 120 -0Q6
(J (~~~~~~~~~~1
~~~~~~~~~C
E
0-8 16 24 32 8 16 2 2
FractionNumber
FIG. 2. Elutionfrompoly(C)-agarose columns of DNApolymeraseactivityfrom high-speedmicrosomal
supernatantpreparationsderivedfrom (A)JLSV-1O cells and (B) JLSV-9 cells. High-speed cytoplasmic supernatantfractionswereprepared as described in the text.Portions(20
itl)
fromeach eluatefractionwereassayed forpoly(A)J (dT)12.IS(0)-andpoly(rCm) (dG)1218(0)-utilizingactivity.
on November 10, 2019 by guest
http://jvi.asm.org/
[image:4.504.119.404.471.622.2]VOL 26,1978 INTRACELLULAR R-MuLV DNA POLYMERASES 5
112
A Bo
OI
E -0~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~.8
IZS
T
EE
0. CL4
VO 2
2-~~~~~~~~~~~~~~~~~~~~~~
4 8 12 1610 24 28 23640 44 48 4 8 12 16 20 24 28 32 36 40 44 48
[image:5.504.108.397.60.210.2]FroctionNumber
FIG. 3. Chromatography of JLSV-10high-speedsupenatantpoly(rC)-agarosepooledeluatefractions8 to 12(A)and15to19(B) (seeFig. 2A)onphosphocellulose. Thepooled eluate fractionswereprocessedthrough
DEAE-cellulosecolumnsbefore phosphocelulose chromatography(see text). Phosphocellulosecolumneluate
fiactions
(2O-pdportions) upereassayedforpoIY(A)-(dT)12 8 (0)-andpoly(rCm)*(dG)1218 (0)-directedDNA synthesi8.ing to their order of elution fromphosphocellu- ,
lose,
was carried out to detectpossible
differ- 18 +encesin theirproperties. i
Analysis by
sedimentationvelocity.
The16-molecularweights ofthe PCIandPC II forms ; 14
ofreverse transcriptase derived from the high- X 12
speed cytoplasmic supernatant fraction of °4
JLSV-10 cells were determined bysedimenta-
-tionthroughlinear 10 to30% (vol/vol) glycerol 8
-gradients in the presence of 0.4 MKCI(Fig.4).
Rauscher leukemia virus reverse transcriptase j 6 (70,000 daltons), purified by a single passage 4
-throughpoly(C)-agarose (23), and E. coli DNA 2
polymerase I (109,000 daltons) were used as
molecular weight markers. The PC I form of
reverse transcriptase (derivedfrommicrosomal B 2 4 6 8 10 12 14 16 18 20
pellet or supematant) cosedimented with the Fraction number T
virion-derived enzyme, indicating asedimenta-
FiG.
4. Glycerol gradient analysis ofJLSV-10
in-tion coefficientof4.2S corresponding to a mo- traceUularforms of reversetranscriptase.
Phospho-lecular weight of 70,000. The PC II enzyme ceUulose peakPC I(0) or PC II (A)fractions (150
consitently sedimented faster than the PC I id) werediluted 1:1 with 0.5M This-hydrochloride
form,at 5.5 to5.68, conrespondingto amolecular buffer, pH 7.8, containing 0.4 MKCI and I nM
weightof 109,000. The sedimentation patterns dithiothreitoland thenlayered on 10to 30% glycerol
were not alteredbythe useofNonidetP-40 at
gradients
prepared with thesame buffer.Centrifu-0.2%(vol/vol)inaddition to 04 MKCI (datanot
gation
wascarried
out for 16hwith
anSW5O.1
rotor.shown. *Parallelgradients conaining
poly(C)-agarose-puri-shown). .
fled
Rauscher
lukemia
viru DNA polymerase(0)
Template-primerutilization. Since the PC andE. coliDNA polymerase I (arrow indicates peak
I andPC II activities differed in apparent mo- fractionposition) were also runasmolecular
uwight
lecularweight, their respective abilitiestoutilize markers. FractionscoUectedfromthebottom of the
various template-primers (synthetic and natu-
cenrifuge
tubes were assayedfor DNApolymeraseral)weretested andcomparedwiththose of the activitywithpoly(A)*(d-)-2-Ie
virion-derived
enzyme (23). Theresults of thisstudyare shownin Table2. Both forms ofre- formswereableto copyheteropolymericregions
versetransciptasewereabletoutilize thesame ofrabbitglobin mRNA into DNA, thereby
con-spectrum of
template-primers
as has beenpre- firmingtheiridentityas truereversetanscrip-viouslydescribed for the Rauscher leukemia vi- tases.Optimal divalent cation concentrationsfor
rusenzyme(23), withnosignificantdifferences theuseofthevarioustemplate-primersfor both
observedbetween the PCIand PCI forms. It PCIand PC I enzymes werealso foundtobe
should be noted that both the PC I and PC II identical to those previously reported for the
on November 10, 2019 by guest
http://jvi.asm.org/
[image:5.504.256.447.272.438.2]6 MARCUS J. VIROL.
TABLE 2. Utilizationofvarioustemplate-primersby PC I andPC IIJLSV-lO DNApolymerasesand Rauscher leukemia virus reverse transcrptase"
Substrate incorporationby:
Template-primer Divalent cation substrate PClen- PCIlen- Virion-de-substateP Ien PC I en-rived
en-zyme zyme zyme Synthetic
Poly(A) *(dT)1218 Mn2+ TTP 10.1 9.2 26
Poly(dA) (dT)1218 Mn2+ TTP <0.02 <0.02 <0.02
Poly(C) (dG)1218 Mg2+ dGTP 1.5 0.8 3.1
Poly(rCm) (dG)12-18 Mn2+ dGTP 0.8 0.7 1.0
Poly(dC) (dG)1218 Mg2+ dGTP 6.1 5.2 20.2
Natural
Activated calfthymus DNA Mg2+ dGTP 0.3 0.28 1.55
Globin mRNA+(dT)12 Mn2+ dGTP 0.15 0.13 0.21
GlobinmRNA+(dT)12 Mn2+-dATP and dCTP dGTP <0.01 <0.01 <0.01
(dT)12
Mg2e
orMn2+ dGTP <0.01 <0.01 <0.01a
Assay
conditions and standard components used in reaction mixturesaregiveninthetext.In additiontostandardcomponentsandtemplate-primers,0.5mMMnCl2or2mMMgCl2was used whereverindicated,and
100 mMKCIwaspresent in all reactions.Approximately 5ngof RauscherleukemiavirusDNApolymerase enzymewasusedperreaction.Peakphosphocellulosefractions
(20-ptl
portions)ofPC I and PC IIenzymeswereusedforeach reaction.Synthetictemplate-primer-directed reactionswereincubatedat37°Cfor30min.For
globin-mRNA-directedreactions,assaymixturesweredoubled insize,andthe incubation timewasincreasedto
60min.
Rauscher leukemia virus DNApolymerase (23).
optimal
ratio ofpoly(C)
to(dG)12.18
revealedWe have
recently
reported that Pi, at low differences between thetwointracellularforms concentrations(<10mM),effectivelyandselec- ofreversetranscriptase (Fig.
5). The PC Ien-tively inhibits mammalian type C viral DNA zyme and the virion-derived reverse
transcrip-polymerases (23). The PCI, PCII, and virion- tasepreferred a template-to-primer ratio of 25:1, derivedreverse-transcriptase activitieswerein- whereas under identical conditions the PC II hibited80 to90%bytheaddition of2mMPito enzyme showed a marked preference for the reaction mixtures and to an equivalent degree 5:1ratio. Thisdifferencein therateof utiization
by 10mMN-ethylmaleimide (datanotshown). also serves to
explain
thevarying
responsesThese results serve to identify the phosphate of the PC I and PC II enzymes to
sensitivity ofthe PC I and PC II enzymes and
poly(C)
-(dG)12.18
showninTable2,
inwhichaalso suggest that these reverse transcriptase 20:1template-to-primerratiowasused.
preparationsare notcontaminated with signifi- ApparentKmvalueswere also determined for
cantquantitiesof DNApolymerasesfOory,the substrate DNA precursors for thePC I,PC II,
formerbeingrelativelyresistanttoinhibitionby andvirion-derivedforms ofreversetranscriptase
N-ethylmaleimide (33)andthe latterbeingstim- with thesetwo
template-primers.
Allthreeformsulated rather than inhibitedbythe addition of ofreversetranscriptase yieldedanapparentKm Pitoreaction mixtures (16, 23). value for dTTP and dGTP of 20 ,uM. This value
In adetailed studyof thecatalytic properties did not change, regardless of the
template-to-of theRauscher leukemia virus DNApolymer- primer nucleotide ratio that was used. Apparent
ase,we have shownthatchangingthetemplate- Kmvaluesfor thesetwotemplate-primers for
all
to-primer nucleotide ratio can significantly af- threeforms ofreversetranscriptase were
iden-fect the rate of substrate polymerization (23). ticalat0.5 to0.7jug/ml.The pH optima for PC
The effect ofchanging the template-to-primer I, PC II, andthe virion-derived or microsomal
ratio on the rate of DNAsynthesis by PCI,PC pellet-derived reverse transcriptase were
deter-II,orthe Rauscherleukemiavirus-derived DNA mined by using Tris-hydrochloride buffers of polymerase wasdetermined, using afixed con- varying pH in the reaction mixture. For centration of
poly(A)looo
or poly(C)soo and an- poly(A).(dT)12-1s-directed
synthesis, pH 7.8 tonealing various amounts ofappropriate DNA 8.1 was the range found
optimal
for enzymeoligomersasprimers (30).All three forms of the activity from all four sources. Therefore, al-enzymepreferredanequimolarratio ofpoly(A) thoughapparent kinetic constants for PC I and
to
(dT)o
nucleotides (datanotshown) for opti- PC II reverse transcriptase are identical withmal rates of synthesis. Determination of the respect to substrateand template-primer
on November 10, 2019 by guest
http://jvi.asm.org/
VOL. 26, 1978 INTRACELLULAR R-MuLV DNA POLYMERASES 7
|1. 003' ,' \' z 2
~~~~~~~10
80
.E l 60 4 8 l 6 2 42
E~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
Ig^G
o20_ _ ~~~~~~~~~~FIG.6. Inhibition of PCI1(O)and PCII1(0)
intra-C cellular JLSV-10 reverse transcriptase and JLSV-9
t;
~~~~~~~~~~DNA
polymerase(A\)
by IgG directed againstX I I l I
~~~~~~~Rauscher
leukemia virus DNA polymerase. Enzyme10 20 3 0 5 niiinsuiswr are out as described in
theextTheJLS-9 DAplymeaseactivityused Ratioottemplate to primer in this study was that derived frompoly(C)-agarose FIG. 5. Effect ofvarioustemplate-to-primernolar columneluatefractions (see Fig. 2) corresponding to
nucleotide ratiosofpoly(C) (dG)12.18 on theactivity differentareas of the elutionprofile, whichall gave
reversetranscrwptaseand Rauscher leukemia virus- (as picomoles offH]dTMP incorporated per hour) derived (0) reverse transcrptase. Poly(C) and~ were: PC I, 18; PC II, 13; JLSV-9 DNA polymerase,
(dO)12.18 were annealed in thedesiredratios as de- 4
scribed in the text. Enzyme activity inallcases was
KCI. Values corresponding to lOO'% activity in this DNAsynthesis by enzoymai acivtedluting8at
experiment
(aspicomoles ofpHJdGMP
ingcorporated
DAsnhssb nyai ciiyeuigaper 30min)Jwere:PC Ipolymerase,2.5;-PClpolym- various salt concentrations after
chromatogra-erase, 2.0; Rauscherleukemia virus DNApolymnerase, phy of uninfected JLSV-9 extracts on
poly(C)-12.5. agarose (Fig. 2B) is also shown (Fig. 6). No
significantinhibitionof the normal cellular DNA
ties, as well as to pH optima, these enzymes may polymerase activity was observed. These results
bedistinguiishedby the template-to-primer nu- suggest that there are no significant antigenic
cleotide ratio forpoly(C)* (dG)1218requiredfor differences between the intracellular forms of
optimal rates of poly(dG) synthesis. reversetranscriptasethat wereresolvedby
phos-Susceptibility to IgG inhibition. The stud- phocellulose chromatography.
ies described above on the catalytic properties Enzyme stability and heat inactivation.
Of the PC I and PC II enzymes derived from Enzymefractions that were obtained by
phos-JLSV-10 Rauscher leukemia virus-producing phocellulose chromatography were either used
cells suggested that they were two forms of immediately or stored at000 after the addition
virion-coded reverse transcriptase. The immu- of bovineserumalbumin monomer (Miles
Lab-nological relatedness of the intracellular forms oratories, Inc.) to a final concentration of 100
of reverse transcriptase was tested by examining ,ug/ml. Long-term storage of PC I and PC II
the susceptibility of PC I and PC II toinhibition enzymefractionswas notattempted.During the
by IgG purified from rabbit antiserum directed course of the above studies, it was noted that,
againstpartiallypurifiedRauscherleukemiavi- even with the addition ofexogenous proteins,
rus DNApolymerase. The IgGpreparationthat
PCOII
activity declined at000more rapidly thanwas used has been shown to be capable of de- did PC I activity. To determine whether this
tecting differences in mammalian type C viral decline in activity was intrinsic to a particular
DNA polymerases from various sources (28). enzymeform, the PC I and PC II enzyme
frac-The results of this study are shown in Fig. 6. tions, as well as poly(C)-agarose-purified
The ability of increasing quantities of anti- Rauscher leukemia virus DNA polymerase, were
Rauscher leukemia virusDNApolymeraseIgG heated for various periods of time at4500in the
to inhibit PC I and PC II enzyme forms is absence of
template-prmner,
substrate, ordiva-qualitatively
siilai
and equivalent to that de- lent cation (30). The enzymefractionswere thentermned withvirion-derived enzyme (data not cooled in an ice bath and tested for the loss of
shown). In addition to this study, the abilityof ability to direct DNA synthesis with
on November 10, 2019 by guest
http://jvi.asm.org/
8 MARCUS J. VIROL.
poly(rCm) (dG)1218as template-primer. There- identical to that found in the virion orin the sults of this study are shown in Fig. 7. The microsomal membranepellet fraction.
virion-purified reverse transcriptase and the in- DISCUSSION
tracellular PC I peak enzyme fraction
showed,
dIscUiON
underidenticalconditions, identical rates of heat Thisreport has described the presence of two inactivation. The PC II enzyme was far more forms ofRauscher leukemia virus DNA polym-sensitive to heatinactivation thanwasthe PCI erase in avirus-producingcell line.Fractionation enzyme or virion-derived polymerase. The ap- of the cytoplasmrevealed that the
majority
of proximate To.5 (one-half thermal inactivation reverse-transcriptase activity (>85%) in JLSV-time) for the PC I enzyme andRauscherleuke- 10 cells is found in the microsomal membrane mia virus-derived enzyme at this temperature, pellet fraction and that it is identical to the and measured with the template-primer, is 65virion
form of the enzyme in such parameters asman, whereas the T.6 for the PC II form Of molecular weight, optimal conditions for tem-reverse transcriptase is 11 mi. From this as well
plate-primer
utilization, and heat inactivation as the other data described above, the PC Ikinetics.
The high-speed cytoplasmicsuperna-intracellular form of reverse transcriptase I tant fraction was found to contain only 10 to 20% of the total reverse-transcriptase activity,
100 andapproximately 20% of this soluble enzyme
90 activity was present in a form that differed from
the virion enzyme. The two forms that were
70 \ - t found in this fraction differ in their
position
of70 _ elutionfromphosphocellulose columns. The PC
60 \ _ I
form
(eluting
first fromphosphocellulose)
ap-pears identicaltothevirion-derived enzyme in
so _ those
properties
thatwe havedescribedprevi-ously (23). The PC II enzyme form appears
40 higher in molecular weight (109,000 as opposed
to70,000[Fig.3]) than the PC I form, prefers a lowertemplate-to-primer ratio for optimal
copy-; \ming ofpoly(C) templates (Fig. 4), and is more
E
30 \ _ thermolabilethan PC I
(Fig.
7).
Both the PCIand PC II forms ofreverse
transcriptase,
how-ever, appearantigenically similar in the degree
to which they are inhibited by IgG directed
20 _ agat the
partially purified
virion enzyme(Fig.
6).
Although the quantity of PC II appears low compared with that of the PC Iformintotal cell fractions, it appears to be ofsignificant quantity since the majority of viral DNA polymerase
activityobserved is mostlikely due to the
pres-10.
sC
, ,I0
I5
, ence(in
the microsomal membranepellet)
of20 30
budding
virusparticles
aswell
as"core"parti-Minutes
[image:8.504.65.256.250.518.2]o5
incubation451
at cles. Because no treatment of cytoplasm wasFIG. 7. Kinetics ofheat inactivation of PCI(0) undertaken other than fractionation by
differ-andPCH(A) JLSV-10
intracelular
DNApolymer- ential centrifugation, the soluble enzyme frac-asesandRauscherleukemia virus reverse transcrip- tion should reflect the status ofreverse-tran-tase (0) at
450C.
Thepurified DNApolymerasesin scnptase formspriortopackaging in viral cores. buffer containing 0.05 MTris-hydrochloride, pH7.8, Processing ofthe uninfected cell line (JLSV-9) ImMdithiothreitol, 0.5MKCI, 10%glycerol, and 100 in anidentical manner produced no detectable pg of bovineserumalbumin monomer per ml, in 25- quantity ofreverse-transcriptaseactivity, asde-ytl
portions, wereplaced in a45C waterbathfor termined by the ability of a DNA polymerase toincreasing period'softime,after which thete were copy themodified template poly(rCm)
(Fig.
2B).
cooled in anice bath.Ice-cold reaction mixtures were Experiments in which purified viral or
mi-then added to the enzyme fractions, and crosomal J n whic
purived
versetran-poly(rCm) (dG)128-directed synthesis wasinitiated crosomal JLSV-lO pellet-derived reverse
tran-byplacing the tubes in a 37°C water bath. Thepercent scriptase was mixed withJLSV-9 cells, which activity remaining was calculated byusing as con- werethen processedtodetect thepossible
for-trolsenzymefractions that had not been exposedto mation of PC II, yielded negative results (only
heat. the PC Iformwas observed). Therefore, PC II
on November 10, 2019 by guest
http://jvi.asm.org/
does not appear to arise from the fortuitous gent on the PC II intracellular form of reverse complexing of viral DNA polymerase with a transcriptase also failed to resolve high- and low-normal cellularcomponent or as a minor polym- molecular-weight components. Therefore, the erase ofuninfected cells. Theseexperiments also PC II enzyme
forn
that we havedescribed
does demonstrate for the first time the ability of not appear similar to the enzyme complex re-poly(C)-agarose to select for reverse transcrip- ported earlier.tase in cytoplasmic preparations of virus-pro- The PC II enzyme form that we have de-ducing cells and to discriminate against cellular scribed appears similar in molecular weight to
polymerase. Such discrimination may be ob- anintracellular form of RD-114 virus DNA
po-served in the disproportionate loss of lymerase described by Gerwin et al. in infected
poly(A) (dT)128-directed DNA synthesis as cells (8). Although the high- and
low-molecular-compared with poly(rCm)-(dG)12.s-directed weight forms of the intracellular RD-114 virus
synthesis (specific forreverse transcriptase) in enzymes appeared identical with respect to
di-the JLSV-10 soluble cellular fraction poly(C)- valent cationoptima and apparent kinetic con-agarose column flow through (Table 1). The stants for substrates and template-primers,
identification of PC I and PC HIas"true" reverse other biochemical parameters, including the
transcriptases was made on the basis of their abilitytodirect naturalRNA-directed synthesis,
abilitytocopyheteropolymeric regions of rabbit were nottested (8). The increasedthermolability globin mRNA (Table 2) as well as on their of the PC II Rauscher leukemia virus intracel-responsetospecificantibodies (Fig.6). lular enzyme form, along with the increase in The existence of multiple forms of reverse apparent molecular weight, suggests an
altera-transcriptase withdiffering properties in intra- tion in structure as compared with the
virion-cellular particles obtained from cells infected derived form of reversetranscriptase. The
find-with gibbon ape leukemia virus has been re- ing that the PC II enzyme formcopiespoly(rC)
portedbyGillespieetal. (11). However, the two optimallyat atemplate-to-primer ratio different
forms studied by this group appeared to arise from that which is optimal for the virion enzyme via monomer-dimer interconversion mediated alsosuggests apossiblestructural difference
be-by salt and/or detergent concentrations. tweenthe twoenzymes. It istemptingto
spec-Rauscher leukemia virus DNApolymerasehas ulate, asothers have (8), that the
high-molecu-also beenreportedtoexistas aseries of aggre- lar-weight form serves as a precursor to the gate forms(26),althoughwehave shownthat in virion-packagedform inasystem similarto
that.
the presence of0.4MKCInosignificant degree of the/Bsubunit of avian retroviral reverse
tran-ofaggregationis observed(23). Bandyopadhyay scriptase(9). Our findings (i) that the PC II form
fractionatedand characterized the DNApolym- wasobservedonlyin soluble rather than
partic-erasesfrom JLSV-9(3)and JLSV-10(4)cellsby ulate fractions and (ii) thatmixing experiments
using classical purification procedures. He ob- failed togenerate the PC II form frompurified
servedauniqueDNApolymerasewithamolec- polymerase and uninfected cell components
ularweightof110,000in thecytoplasmofJLSV- would seem to lend support to that concept.
10cells,whichappearedtorepresentacomplex However,additionalpossibilitiesthat thePCII
formnedbetween allcytoplasmicviralpolymerase enzyme form(i)represents the DNApolymerase
and a 35,000-dalton cellular DNA polymerase. of a variant strain ofRauscher leukemia virus
Thecomplexcouldbeseparated by detergentor presentinsmall amounts or (ii) isproduced
by
phospholipasetreatmentandwasalsoobserved a related endogenous viral genome present in
by the author to be present in purified viral JLSV-9 cells that is activatedduring viral
infec-preparations. Attempts inourlaboratoryto re- tioncannot,atpresent, becompletelyruledout.
producetheseresultsby followingtheidentical Futurestudiesontheenzyme formPCIIwill
fractionationprocedurewereunsuccessful. How- concentrate onthequestionof whether it indeed
ever, it was found that the ammonium sulfate representsanenzymatically active intennediate
fractionationstep resulted in enrichment for the in thecleavage of the polyproteinprecursor to
PCI(virion) enzymeformn byprecipitatingpar- DNA
polymerase
(15) or a noncovalently butticle-bound
polymerase.
Furthermore, sedimen- tightlylinkedprotein-enzymecomplex.
tation velocity profiles of Rauscher leukemia
ACKNOWLEDGMENTS
virus reverse transcriptase from crude or puri- I thank Mukund J. Modak for helpful discussions and fiedenzymefractions,in thepresenceorabsence purified Rauscherleukemia virusand E.coliDNApolymerase ofdetergent, wereconsistentlyfree ofanysuch IandCharles J. Sherr foranti-reverse-tanscriptaseIgG.The 35,000-dalton polymerase activity (23)whenas- continuedinterestandencouragement of Nurul H. Sarkaris
sayedunderconditions reportedoptimalfor its appreciated.for experttechnical assistance.Thanksarealsoexpressed to Steven W.Smith
detection (4). Sedimentation velocity analysis Thisstudy wassupportedby Public Health Servicegrants
performed in thepresenceor absence of deter- CA-08748and CA-18369from the National CancerInstitute.
on November 10, 2019 by guest
http://jvi.asm.org/
LITERATURE CITED Virology53:258-273.
1.Allaudeen, H. S., M. G. Sarg haran, and R. C. 18. Marcus, S.L., and M. J.Modak.1976. Observations on Gallo. 1976. Acomparative evaluation of methods for the template specific conditions for DNA synthesisby isolation ofRNA-directedpolymerasein a reconstituted avianmyeloblastosis virus DNA polymerase. Nucleic system. Biochim.Biophys. Acta 435:45-62. AcidRes.3:1473-1486.
2. Aposhian, H. V., and A. Kornberg. 1972. Enzymatic 19. Marcus, S. L,M. J. Modak, and L F.Cavalieri.1974. synthesis ofdeoxyribonucleicacid. IX. Thepolymerase Purification of avianmyeloblastosisvirus DNA polym-formedafterT2bacteriophageinfectionofEscherichia eraseby affinity chromatography on polycytidylate-coli:a newenzyme.J. Biol. Chem.237:519-525. agarose. J.Virol.14:853-859.
3. Bandyopadhyay,A. K. 1975. Purificationand properties 20. Marcus,S.L, M. J.Modak,andL F.Cavalleri.1974. of nuclear and cytoplasmic DNA polymerases from Evidence fortemplate-specificsites on DNA polymer-JLS-V9cells. Arch.Biochem. Biophys.1":72-82. ases.Biochem.Biophys. Res.Commun.56:516-621. 4. Bandyopadhyay, A. K. 1975. Partialpurificationand 21. Marcus, S. L., N.H. Sarkar,and M. J. Modak. 1976.
properties of DNApolymerases from JLSV-10 cells. Purification andpropertiesofmurinemammary tumor Arch.Biochem.Biophys. 166:83-93. virus DNApolymerase.Virology71:242-254. 5. Eagle,H. 1959. Amino acidmetabolism in mammalian 22. Marcus,S.L,S. W.Smith,C. L.Jarowski,and M.J.
cell cultures. Science310:432-437. Modak. 1976. Terminaldeoxyribonucleotidyl transfer-6. Gerard, G. F.,and D. P.Grandgenett.1975.Purifica- an activity in acute undifferentiated leukemia.
Bio-tion and characterizaBio-tion of the DNA polymerase and chem.Biophys.Res. Commun. 70:37-44.
RNase HactivitiesinMoloneymurine sarcoma-leuke- 23. Modak, M. J., and S. L Marcus. 1977. Purificationand miavirus. J. Virol. 15:785-797. propertiesof Rauscher leukemia virus DNApolymerase 7. Gerard, G. F.,F.Rottman, andML Green.1974.Poly and selective inhibition ofmammalian virus reverse (2'TO-methylcytidylate) oligodeoxyguanylateas atem- transcriptase by inorganicphosphate. J.Biol. Chem. plateforthe ribonucleicacid directed deoxyribonucleic 252:11-19.
acidpolymeraseinribonucleic andtumorvirus particles 24. Modak,M.J.,S. LMarcus,and L F. Cavalieri. 1973. andaspecific probefor theribonucleic acid directed DNAcomplementaryto rabbitglobinmRNA madeby enzyme in transformed murine cells. Biochemistry E. coli DNApolymeraseI. Biochem. Biophys. Res.
13:1632-1640. Common.55:1-7.
8. Gerwin, B. I., S. G.Smith,and P. T.Peebles. 1975. 25. Moelling,K. 1974. Characterization ofreverse transcrip-Twoactive forms ofRD-114virus DNA polymerase in taseand RNase H from Friend murine leukemia virus. infected cells. Cell6:45-52. Virology62:46-59.
9.Gibson, W., and I. M. Verma. 1974.Studies on the 26. Nakajima,K.,K.Ohno,and Y. Ito. 1974. Interconver-reversetranscriptaseofRNAtumorviruses. I.Struc. sion of molecular size of the DNA polymerasefrom tural relatednessof the two subunits of avian RNA Rauscher leukemia virus.Intervirology3:332-341. tumor viruses. Proc. Natl. Acad. Sci. U.S.A. 27. Ritzi, E., D. S.Martin,R. LStolfi, andS.Spiegelman. 71:49914994. 1976. Plasma levels of a viral protein as adiagnostic 10.Gillespie, D., R. E.Gallagher, R.G. Smith, W. C. signalforthe presence of tumor: themurinemammary Saxinger,and R. C.Gallo.1975.On theevidencefor tumor model. Proc. Natl. Acad. Sci. U.S.A. Type C RNAtumor virusinformationand virus-related 73:4190-4194.
reversetranscriptaseinanimalsand inhuman leukemic 28. Sherr, C.F., M. M.Lieber,R. E.Benveniste,and G. cells,p. 1-27. In A.Gottlieb,0.J.Plescia,and D. H.L. J. Todaro. 1974. Endogenous baboon type C virus
Bishop (ed.), Fundamentalaspects ofneoplasia.Sprin- (M7):biochemical and immunologic characterization.
ger-Verlag,New York. Virology58:492-503.
11. Gillespie,D., W. C.Saxinger,andR.C. Gallo. 1975. 29. Ting,R.C.,S.S. Yang, and R. C. Gallo. 1972.Reverse Informationtransferincellsinfected by RNA tumor transcriptase RNA tumor virustransformationand de-virusesand extension to humanneoplasia. Prog. Nucleic rivativesofrifamycinSV.Nature (London) NewBiol. AcidRHs.Mol. Biol.15:1-108. 236:163-166.
12. Green,M., and G. F.Gerard.1974.RNA directed DNA 30. Verma,L.M. 1975. Studies onreversetranscriptase of polymerase-properties and functions in oncogenic RNA tumorviruses. IH.Properties of purified Moloney RNAviruses and cells. Prog. Nucleic AcidRes.Mol. murineleukemia virus DNA polymerase andassociated Biol. 14:187-334. RNase H.J. Virol. 15:843-854.
13. Hana ILH.,and T.Hanafusa. 1971. Non-infectious 31. Verma, I.M., G. F. Temple, H. Fan, and D. Baltimore. RSV deficient in DNA polymerase. Virology 1972. In vitro synthesis of DNA complementary to
43:313-320. rabbitreticulocyte RNA. Nature (London) NewBiol.
14. Hurwitz, J.,and J. P.Leis.1972.RNA-dependent DNA 235:163-167.
polymerase activityof RNA tumorviruses.I. Directing 32. Waters, L. C., and W.-K. Yang. 1974. Comparative influence of DNA in the reaction. J.Virol. 9:116-129. biochemicalproperties ofRNA-directed DNA polym-15.Jamjoom, G.A.,R.B. Naso, and R. B.Ariinghaus. erasesfrom Rauscher murine leukemiavirusandavian
1977.Furthercharacterization of intracellularprecursor myeloblastosisvirus. CancerRes.34:2585-2593. polypeptides of Rauscher leukemia virus. Virology 33. Weissbach, A. 1977. Eukaryotic DNA polymerases.
78:11-34. Annu.Rev.Biochem.46:25-49.
16.Knopf, K.-W., MLYamada, and A.Weissbach.1976. 34. Wu, A. M., M. G. Sarugadharan, andR.C. Gallo. HeLacellDNApolymerasey,furtherpurification and 1974.Separation of ribonuclease H and RNA-directed propertiesof theenzyme. Biochemistry15:4540-4548. DNApolymerase (reverse transcriptase) of murine type 17. Linial,K,andW. S.Mason. 1973.Characterization of C RNAtumor viruses. Proc.Natl.Acad. Sci. U.S.A.
two conditionalearly mutants of Rous sarcoma virus. 71:1871-1876.
on November 10, 2019 by guest
http://jvi.asm.org/