organic papers
Acta Cryst.(2005). E61, o1173–o1175 doi:10.1107/S1600536805009281 Zhang, Shan and Xu C
9H9N3O2S2
o1173
Acta Crystallographica Section EStructure Reports Online
ISSN 1600-5368
Methyl
b
-
N
-(3-nitrophenylmethylene)-dithiocarbazate
Yan-Ling Zhang,aShang Shana* and Duan-Jun Xub
a
College of Chemical and Materials Engineering, Zhejiang University of Technology, People’s Republic of China, andbDepartment of
Chemistry, Zhejiang University, People’s Republic of China
Correspondence e-mail: shanshang@mail.hz.zj.cn
Key indicators
Single-crystal X-ray study
T= 295 K
Mean(C–C) = 0.002 A˚
Rfactor = 0.032
wRfactor = 0.079
Data-to-parameter ratio = 17.9
For details of how these key indicators were automatically derived from the article, see http://journals.iucr.org/e.
#2005 International Union of Crystallography Printed in Great Britain – all rights reserved
Crystals of the title compound, C9H9N3O2S2, were obtained from a condensation reaction ofS-methyl dithiocarbazate and 3-nitrobenzaldehyde. The planar dithiocarbazate moiety subtends an angle of 10.54 (8) with respect to the plane of the nitrophenyl ring. Electron delocalization occurs between the imino and dithiocarboxyl groups. The partially overlapped arrangement of parallel benzene rings of neighboring mol-ecules, with a face-to-face distance of 3.343 (8) A˚ , suggests the existence of–stacking.
Comment
Phenylhydrazone and its derivatives have attracted our attention as they show potential applications in the biological field (Okabe et al., 1993; Hu et al., 2001). As part of our ongoing investigation into the anticancer properties of phenylhydrazone, the title compound, (I), has been prepared and its structure is presented here.
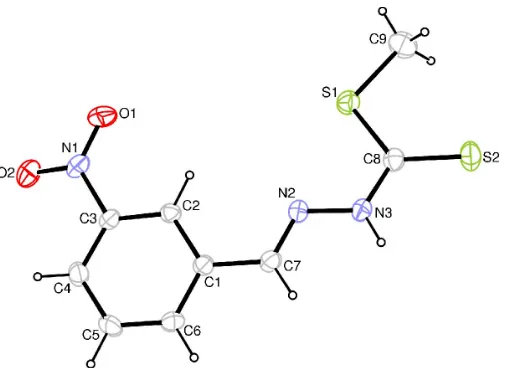
The molecular structure of (I) is shown in Fig. 1. The di-thiocarbazate moiety is planar; its mean plane subtends an
[image:1.610.208.461.539.723.2]Received 18 March 2005 Accepted 23 March 2005 Online 31 March 2005
Figure 1
angle of 10.54 (8) with respect to the plane of the benzene ring. This differs from the situation in the related 4-nitro-phenyl isomer, in which both sections of the molecule are coplanar (Duanet al., 1997). The relatively short C8—N3 bond distance (Table 1) suggests a degree of electron delocalization between the imino and dithiocarboxyl groups. The nitro group is tilted out of the benzene plane, with a dihedral angle of 10.6 (2), which may be due to hydrogen bonding between atom O2 of the nitro substituent and the imino group of a neighboring molecule (Table 2 and Fig. 2).
A partially overlapped arrangement of parallel benzene rings [symmetry code: (ii) 1x,y, 1z] is observed in the crystal structur (Fig. 3). The face-to-face distance of 3.343 (8) A˚ clearly suggests the existence ofe – stacking between benzene rings.
Experimental
Methyl dithiocarbazate was synthesized in the manner reported previously (Huet al., 2001). Methyl dithiocarbazate (1.24 g, 10 mmol) and 3-nitrobenzaldehyde (1.50 g, 10 mmol) were dissolved in ethanol (10 ml) and refluxed for 4 h. Fine yellow crystals appeared on cooling. They were separated and washed with cold water three times. Single crystals of (I) were obtained by recrystallization from absolute ethanol.
Crystal data
C9H9N3O2S2 Mr= 255.31 Monoclinic,P21=c a= 8.7176 (4) A˚
b= 7.5589 (3) A˚
c= 17.5879 (7) A˚
= 100.666 (2)
V= 1138.94 (8) A˚3 Z= 4
Dx= 1.489 Mg m3 MoKradiation Cell parameters from 4542
reflections
= 2.4–27.0
= 0.46 mm1 T= 295 (2) K Needle, yellow 0.500.130.10 mm
Data collection
Rigaku R-AXIS RAPID diffractometer
!scans
Absorption correction: multi-scan (ABSCOR; Higashi, 1995)
Tmin= 0.770,Tmax= 0.952
4646 measured reflections
2611 independent reflections 1703 reflections withI> 2(I)
Rint= 0.021
max= 27.5 h=11!11
k=9!9
l=22!22
Refinement
Refinement onF2 R[F2> 2(F2)] = 0.032 wR(F2) = 0.079 S= 0.88 2611 reflections 146 parameters
H-atom parameters constrained
w= 1/[2(Fo2) + (0.0448P)2]
whereP= (Fo2+ 2Fc2)/3
(/)max< 0.001
max= 0.21 e A˚ 3
[image:2.610.310.567.65.269.2]min=0.19 e A˚ 3
Table 1
Selected geometric parameters (A˚ ,).
S1—C8 1.7457 (17)
S1—C9 1.791 (2)
S2—C8 1.6559 (17)
N2—C7 1.270 (2)
N2—N3 1.3707 (18)
N3—C8 1.340 (2)
C1—C7 1.462 (2)
C8—S1—C9 102.10 (9) S1—C8—S2 125.87 (11)
Table 2
Hydrogen-bonding geometry (A˚ ,).
D—H A D—H H A D A D—H A
N3—H3 O2i
0.86 2.29 3.125 (2) 163
Symmetry code: (i)x;1 2y;z
1 2.
organic papers
o1174
Zhang, Shan and Xu C [image:2.610.58.281.74.414.2]9H9N3O2S2 Acta Cryst.(2005). E61, o1173–o1175
Figure 2
The crystal packing, showing the hydrogen bonding (dashed lines) and parallel arrangement of benzene rings of neighboring molecules.
Figure 3
[image:2.610.313.565.596.647.2]Methyl H atoms were placed in calculated positions (C—H = 0.96 A˚ ) and torsion angle was refined to fit the electron density, with
Uiso(H) = 1.5Ueq(C). Other H atoms were placed in calculated
posi-tions (C—H = 0.93 A˚ and N—H = 0.86 A˚) and included in the final cycles of refinement as riding, withUiso(H) = 1.2Ueq(C,N).
Data collection: PROCESS-AUTO (Rigaku, 1998); cell refine-ment:PROCESS-AUTO; data reduction:CrystalStructure (Rigaku/ MSC, 2002); program(s) used to solve structure:SIR92(Altomareet al., 1993); program(s) used to refine structure: SHELXL97 (Shel-drick, 1997); molecular graphics:ORTEP-3 for Windows(Farrugia, 1997) andXP(Siemens, 1994); software used to prepare material for publication:WinGX(Farrugia, 1999).
The work was supported by the Natural Science Foundation of Zhejiang Province of China (grant No. M203027) and the
National Natural Science Foundation of China (grant No. 20443003).
References
Altomare, A., Cascarano, G., Giacovazzo, C. & Guagliardi, A. (1993).J. Appl. Cryst.26, 343–350.
Duan, C.-Y., Tian, Y.-P., You, X.-Z. & Mak, T. C. W. (1997).Polyhedron,16, 4097–4103.
Farrugia, L. J. (1997).J. Appl. Cryst.30, 565. Farrugia, L. J. (1999).J. Appl. Cryst.32, 837–838.
Higashi,, T. (1995).ABSCOR.Rigaku Corporation, Tokyo, Japan. Hu, W., Sun, N. & Yang, Z. (2001).Chem. J. Chin. Univ.22, 2014–2017. Okabe, N., Nakamura, T. & Fukuda, H. (1993).Acta Cryst.C49, 1678–1680. Rigaku (1998).PROCESS-AUTO. Rigaku Corporation, Tokyo, Japan. Rigaku/MSC (2002).CrystalStructure.Version 3.00. Rigaku/MSC, 9009 New
Trails Drive, The Woodlands, TX 77381-5209, USA.
Sheldrick, G. M. (1997).SHELXL97. University of Go¨ttingen, Germany. Siemens (1994).XP.Version 5.03. Siemens Analytical X-ray Instruments Inc.,
Madison, Wisconsin, USA.
organic papers
Acta Cryst.(2005). E61, o1173–o1175 Zhang, Shan and Xu C
supporting information
sup-1
Acta Cryst. (2005). E61, o1173–o1175
supporting information
Acta Cryst. (2005). E61, o1173–o1175 [https://doi.org/10.1107/S1600536805009281]
Methyl
β
-N-(3-nitrophenylmethylene)dithiocarbazate
Yan-Ling Zhang, Shang Shan and Duan-Jun Xu
Methyl β-N-(3-nitrophenylmethylene)dithiocarbazate
Crystal data C9H9N3O2S2
Mr = 255.31
Monoclinic, P21/c
Hall symbol: -p 2ybc a = 8.7176 (4) Å b = 7.5589 (3) Å c = 17.5879 (7) Å β = 100.666 (2)° V = 1138.94 (8) Å3
Z = 4
F(000) = 528 Dx = 1.489 Mg m−3
Mo Kα radiation, λ = 0.71073 Å Cell parameters from 4542 reflections θ = 2.4–27.0°
µ = 0.46 mm−1
T = 295 K Needle, yellow 0.50 × 0.13 × 0.10 mm
Data collection
Rigaku R-AXIS RAPID diffractometer
Radiation source: fine-focus sealed tube Graphite monochromator
Detector resolution: 10.00 pixels mm-1
ω scans
Absorption correction: multi-scan (ABSCOR; Higashi, 1995) Tmin = 0.770, Tmax = 0.952
4646 measured reflections 2611 independent reflections 1703 reflections with I > 2σ(I) Rint = 0.021
θmax = 27.5°, θmin = 2.4°
h = −11→11 k = −9→9 l = −22→22
Refinement Refinement on F2
Least-squares matrix: full R[F2 > 2σ(F2)] = 0.032
wR(F2) = 0.079
S = 0.88 2611 reflections 146 parameters 0 restraints
Primary atom site location: structure-invariant direct methods
Secondary atom site location: difference Fourier map
Hydrogen site location: inferred from neighbouring sites
H-atom parameters constrained w = 1/[σ2(F
o2) + (0.0448P)2]
where P = (Fo2 + 2Fc2)/3
(Δ/σ)max < 0.001
Δρmax = 0.21 e Å−3
Δρmin = −0.19 e Å−3
Special details
supporting information
sup-2
Acta Cryst. (2005). E61, o1173–o1175
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2,
conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used
only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2
are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger.
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
x y z Uiso*/Ueq
S1 0.09695 (6) 0.48574 (7) 0.39342 (3) 0.04758 (15) S2 0.10856 (6) 0.62236 (8) 0.23266 (3) 0.05584 (17) N1 0.53123 (18) 0.1284 (2) 0.66293 (8) 0.0434 (4) N2 0.39713 (16) 0.37137 (19) 0.39098 (7) 0.0377 (3) N3 0.33002 (17) 0.44926 (19) 0.32250 (7) 0.0421 (4) H3 0.3809 0.4545 0.2851 0.050* O1 0.42125 (18) 0.2272 (2) 0.66359 (8) 0.0672 (4) O2 0.57772 (17) 0.0236 (2) 0.71532 (7) 0.0628 (4) C1 0.60959 (18) 0.2208 (2) 0.46639 (8) 0.0341 (4) C2 0.54034 (18) 0.2238 (2) 0.53179 (8) 0.0338 (4) H2 0.4491 0.2874 0.5319 0.041* C3 0.60982 (19) 0.1308 (2) 0.59610 (8) 0.0344 (4) C4 0.7453 (2) 0.0358 (3) 0.59985 (9) 0.0447 (4) H4 0.7896 −0.0251 0.6445 0.054* C5 0.8132 (2) 0.0342 (3) 0.53498 (11) 0.0519 (5) H5 0.9047 −0.0294 0.5355 0.062* C6 0.7467 (2) 0.1262 (3) 0.46917 (10) 0.0455 (5) H6 0.7947 0.1246 0.4261 0.055* C7 0.5337 (2) 0.3088 (2) 0.39532 (9) 0.0382 (4) H7 0.5850 0.3189 0.3536 0.046* C8 0.1858 (2) 0.5173 (2) 0.31329 (9) 0.0383 (4) C9 −0.0876 (2) 0.5930 (3) 0.36350 (13) 0.0679 (6) H9A −0.0709 0.7141 0.3507 0.102* H9B −0.1457 0.5886 0.4048 0.102* H9C −0.1451 0.5336 0.3189 0.102*
Atomic displacement parameters (Å2)
U11 U22 U33 U12 U13 U23
supporting information
sup-3
Acta Cryst. (2005). E61, o1173–o1175
C5 0.0310 (10) 0.0730 (14) 0.0536 (10) 0.0146 (10) 0.0127 (8) 0.0107 (10) C6 0.0358 (10) 0.0641 (13) 0.0401 (9) 0.0041 (9) 0.0161 (7) 0.0021 (9) C7 0.0391 (10) 0.0430 (10) 0.0337 (8) −0.0017 (8) 0.0102 (7) −0.0001 (8) C8 0.0404 (10) 0.0370 (10) 0.0357 (8) −0.0033 (8) 0.0022 (7) −0.0028 (8) C9 0.0408 (12) 0.0903 (17) 0.0733 (14) 0.0088 (11) 0.0122 (10) 0.0031 (13)
Geometric parameters (Å, º)
S1—C8 1.7457 (17) C2—C3 1.373 (2) S1—C9 1.791 (2) C2—H2 0.9300 S2—C8 1.6559 (17) C3—C4 1.373 (2) N1—O1 1.2171 (19) C4—C5 1.379 (2) N1—O2 1.2258 (18) C4—H4 0.9300 N1—C3 1.4661 (19) C5—C6 1.382 (2) N2—C7 1.270 (2) C5—H5 0.9300 N2—N3 1.3707 (18) C6—H6 0.9300 N3—C8 1.340 (2) C7—H7 0.9300 N3—H3 0.8600 C9—H9A 0.9600 C1—C6 1.386 (2) C9—H9B 0.9600 C1—C2 1.395 (2) C9—H9C 0.9600 C1—C7 1.462 (2)
C8—S1—C9 102.10 (9) C5—C4—H4 121.3 O1—N1—O2 122.74 (15) C4—C5—C6 120.72 (17) O1—N1—C3 119.06 (15) C4—C5—H5 119.6 O2—N1—C3 118.18 (15) C6—C5—H5 119.6 C7—N2—N3 117.10 (13) C5—C6—C1 120.94 (15) C8—N3—N2 120.62 (13) C5—C6—H6 119.5 C8—N3—H3 119.7 C1—C6—H6 119.5 N2—N3—H3 119.7 N2—C7—C1 119.44 (14) C6—C1—C2 118.78 (15) N2—C7—H7 120.3 C6—C1—C7 121.04 (14) C1—C7—H7 120.3 C2—C1—C7 120.11 (15) N3—C8—S2 120.76 (13) C3—C2—C1 118.60 (15) N3—C8—S1 113.36 (12) C3—C2—H2 120.7 S1—C8—S2 125.87 (11) C1—C2—H2 120.7 S1—C9—H9A 109.5 C2—C3—C4 123.47 (15) S1—C9—H9B 109.5 C2—C3—N1 117.79 (15) H9A—C9—H9B 109.5 C4—C3—N1 118.70 (15) S1—C9—H9C 109.5 C3—C4—C5 117.49 (16) H9A—C9—H9C 109.5 C3—C4—H4 121.3 H9B—C9—H9C 109.5
Hydrogen-bond geometry (Å, º)
D—H···A D—H H···A D···A D—H···A N3—H3···O2i 0.86 2.29 3.125 (2) 163