TUNLAYA-ANUKIT, SERMSAWAT. Statistical and Gene Regulatory Network Analysis
of Monolignol Biosynthetic Genes in
Populus trichocarpa
. (Under the direction of Vincent
Chiang, and Fikret Isik).
Lignin is a heteropolymer of monolignol subunits that is in part covalently bound to
cellulose and hemicellulose in secondary cell walls. The difficulty of extraction of lignin is
an obstacle to the utilization of plant biomass. The manipulation of lignin composition and
structure is important for utilizing plant biomass in many applications such as paper
production, bioenergy, chemical feed stocks and animal forage.
The downregulated monolignol pathway transgenics are a great resource for studies
of genes that control the monolignol pathway. We examined transgenics downregulated in
the monolignol pathway using RNA-seq and proteomics to establish the relationships of
transcription factors and lignin content. First, we analyzed the full transcriptomes from
wildtype and transgenic
P. trichocarpa.
We observed direct target and non-direct target
effects of downregulated monolignol pathway genes. We identified xylem specific gene
expression based on the expression pattern of 4 tissues (xylem, phloem, leaf, and shoot). We
identified transcription factors that control monolignol biosynthesis from transcriptome data
of downregulated transgenics. We found new xylem specific candidate transcription factors
that may co-regulate monolignol genes.
by
Sermsawat Tunlaya-anukit
A thesis submitted to the Graduate Faculty of
North Carolina State University
in partial fulfillment of the
requirements for the degree of
Doctor of Philosophy
Forestry and Environmental Resources
Raleigh, North Carolina
2015
APPROVED BY:
_______________________________
______________________________
Vincent L. Chiang
Fikret Isik
Co-Chair of Advisory Committee
Co-Chair of Advisory Committee
________________________________
________________________________
Ronald R. Sederoff
Dahlia M. Nielsen
BIOGRAPHY
ACKNOWLEDGMENTS
TABLE OF CONTENTS
LIST OF TABLES
...
vii
LIST OF FIGURES
...
viii
Chapter 1 Introduction
...
1
Poplar genome ...
1
Transformation of poplar ...1
RNA-Seq ...2
The relationship between transcript abundance and protein abundance ...3
Plant secondary cell wall ...4
Lignin ...5
The monolignol biosynthetic enzymes ...6
Transcription factors regulate secondary cell wall biosynthesis...
9
NAC ...9
MYB
...10
LIM ...11
KNOX ...11
Conclusion ...12
References ...14
Chapter 2 A genetic regulatory network for monolignol biosynthesis in
Populus
trichocarpa
...
27
Abstract ...27
2.1 Introduction ...28
2.2 Materials and methods ...
31
Transgenic production ...31
Transcriptome analysis ...31
Differential gene expression profiles ...32
Identification of xylem specific transcription factors ...32
2.3 Results ...34
The expression of genes in each tissue ...34
Identification of the xylem specific genes ...34
Lignin content variation in transgenics downregulated for the lignin pathway ...47
Differential gene expression in downregulated monolignol biosynthesis
transgenics ...47
A Genetic regulatory network for monolignol biosynthesis ...50
2.4 Discussion ...56
References ...61
Chapter 3 Transcript and protein relationship of monolignol biosynthesis
genes in transgenic
Populus trichocarpa ...
71
Abstract
...71
3.1 Introduction ...71
3.2 Materials and methods ...74
Transgenic plants ...74
Quantitative estimation of transcript abundance by RNA-seq ...74
Protein quantification ...75
The statistical model ...76
3.3 Results ...77
Variation of transcript and protein abundance ...77
Statistical analysis and modeling ...78
3.4 Discussion ...86
References ...87
Chapter 4 Relationship Between Monolignol Biosynthetic Protein Abundance
and Lignin Content Using Transgenic
Populus trichocarpa ...
94
Abstract ...94
4.1 Introduction ...95
Production of Transgenic Trees ...103
Proteomic Analysis ...105
Lignin Quantification ...106
Statistical Analysis ...107
4.3 Results ...108
Production of Transgenic Trees Downregulated for Genes in Monolignol
Biosynthesis ...108
Absolute Quantitation of Protein Abundance ...110
Variation in Protein Abundance in Wildtype and Transgenic Constructs ...111
Variation in Lignin Content ...113
Relationship of Lignin Content and Protein Abundance ...113
4.4 Discussion ...116
References ...126
APPENDICES
...131
Appendix A The R code for analysis Chapter 2 A genetic regulatory network
for monolignol biosynthesis in
Populus trichocarpa
...132
Appendix B The R code for analysis Transcript and protein relationship of
monolignol biosynthesis genes in transgenic
Populus trichocarpa
...179
LIST OF TABLES
Chapter 2
Table 2.1
The expression of cell wall related genes in four types of tissue ...36
Table
2.2
The xylem specific transcription factors and their expression in four
tissues
...40
Table 2.3
The log 2 fold change compared to wildtype of monolignol genes
from 48 downregulated monolignol biosynthesis transgenic
Populus trichocarpa
...4
9
Table 2.4
Summary of transcription factors that can be identified in all 4 networks .60
Chapter 3
Table 3.1
Parameter estimates from linear regression model predicting
standardized protein abundance of monolignol biosynthesis in
P. trichocarpa
from standardized transcript abundance ...85
Chapter 4
Table 4.1
Core proteins of monolignol biosynthesis in
P. trichocarpa
...100
Table 4.2
Parameter estimates from segmented regression models
LIST OF FIGURES
Chapter 1
Figure 1.1
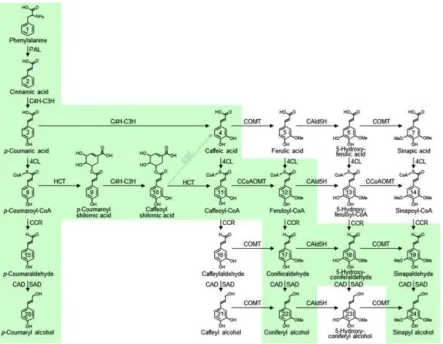
The monolignol biosynthetic pathway ...8
Chapter 2
Figure 2.1
The DEGs comparing the expression of xylem tissue with
other 3 tissues ...35
Figure 2.2
The Summary of GO analysis using xylem specific genes as input and
summarized by REVIGO ...38
Figure 2.3
The top 10 families of transcription factors in the poplar genome ...39
Figure 2.4
The gene regulatory network of monolignol biosynthesis
with all samples in transgenic
Populus trichocarpa
...51
Figure 2.5
The gene regulatory network of monolignol biosynthesis with a
10% lignin reduction in transgenic
Populus trichocarpa
...52
Figure 2.6
The gene regulatory network of monolignol biosynthesis with a
20% lignin reduction in transgenic
Populus trichocarpa
...53
Figure 2.7
. The gene regulatory network of monolignol biosynthesis with a
30% lignin reduction in transgenic
Populus trichocarpa
...54
Figure 2.8
. The merge gene regulatory network of monolignol biosynthesis for
all samples of transgenic
Populus trichocarpa
...55
Figure 2.9
The
numbers of 10 transcription factor groups found in all 4 networks ...58
Chapter 3
Figure 3.1
Scatter plots of monolignol biosynthesis standardized
transcript against standardized protein concentration ...80
Chapter 4
Figure 4.1
The monolignol biosynthetic pathway in
P. trichocarpa
...99
Figure 4.2
Monolignol biosynthetic protein abundance in wildtype
and transgenic
P. trichocarpa
...
112
Figure 4.3
Lignin content of wildtype and transgenic
P. trichocarpa
...115
Figure 4.4
Scatter plots of the monolignol biosynthetic protein family
Chapter 1 Introduction
Poplar genome
Poplar was the first tree to have a full genome sequence. Poplars have small genome
size and are a target species for bioenergy. The genome sequence was published in 2006
(Tuskan et al., 2006), which yielded 22,136 scaffolds and 45,555 genes in 473.1 Mbp. In year
2010 the second draft was released mapping 45,778 genes into 22,136 scaffolds. In the year
2011 the annotation of version 3 was released. This latest version 3.0 consists of 73,013
protein coding transcripts, which adds transcripts for alternative isoforms. The poplar is the
model species for tree as a result of the first sequence genome, there are 2,205 publications
(Nov 4, 2014) that cited the poplar genome paper, more than any other tree species. The
poplar genome was used to study gene families, transcription factors, alternative splicing
variation, and tissue specific gene expression, among other things.
Transformation of poplar
defining the function of genes. When knockout gene are created using post transcriptional
gene silencing by RNAi, they are able to knock down dominant genes. In combination with
tissue specific promoters it is possible to silence genes only in specific tissue and therefore
the plant shows less abnormal growth.
Agrobacterium tumefaciens
that is used for
transformation is derived from a soil pathogen of poplar trees (Keane et al., 1970; Sciaky et
al., 1978). Many Agrobacterium strains were isolated from the crown galls of infected
poplar. Consequently, there are more transgenic poplars than other tree species. The
transgenic poplars were generated for many purposes from abiotic resistance to enhancement
of biomass (Su et al., 2011). Transgenic poplars show a low rate of somaclonal variation and
have stable transformation (Li et al., 2008).
RNA-Seq
The gene expression measurement is an important goal of molecular biology to
understand cellular activity. There are early developments of mRNA measurement from low
throughput Quantitative Real Time Polymerase Chain Reaction or northern blotting to high
throughput oligo nucleotide microarray and RNA-seq.
RNA-seq is the most recent method
for the study of genome-wide expression at single base resolution by producing a quantitative
sample of the sequences. This technique requires ability of high throughput sequencing
platforms or second generation sequencing to sequence a large number of short DNA
fragments at low cost and high throughput. This allows each transcript to be detected a
sufficient number of times for the measurement to be quantitative. Most of the protocols for
RNA-seq begin with the conversion of the RNA into cDNA by reverse transcription,
of the library, and then sequencing the library. The sequencing results will have variation in
sequence length and number of reads (Lister et al., 2009). The sequences can be mapped to
the genome (Langmead et al., 2009b). The advantage of RNA-seq is that sequencing can be
done for any organism, even where no reference sequence or reference transcriptome exists.
In addition, sequencing can more readily reveal information about features such as
alternative splicing, novel transcribed regions, allele-specific expression, RNA-editing and
the discovery of fusion genes (Young et al 2012).
The RNA-seq from differentiating xylem provides a valuable resource for estimating
levels of gene expression. This kind of data can be used to examine the relationships between
transcription factors and their target genes. The RNA-seq can provide more information than
microarrays, such as information on alternative splicing, and identification of transcription
factors. This information has led to the identification of new transcription factors regulating
monolignol pathway genes.
The relationship between transcript abundance and protein abundance
et al., 1999). The translation efficiency of protein also relies on many factors such as amount
of tRNA and amino acids, ribosome binding and ribosome density. Measurement of protein
quantity is difficult due to the complexity of posttranslational processes, such as post
translational modification, compartmentation, and protein degradation (de Sousa Abreu et al.,
2009). Predicting protein or enzyme quantity may help predict the phenotype of an
individual. Early studies were first on bacteria, then yeast, and then on plants and animals
(Ghaemmaghami et al., 2003; Ishihama et al., 2005; Nie et al., 2006a; Schrimpf et al., 2009;
Vogel et al., 2010). Many of these studies focused on the potential to predict protein
abundance from the abundance of their transcripts. Such information should lead to greater
understanding of the regulatory mechanisms of protein synthesis and help to more accurately
predict phenotype from genotype.
Plant secondary cell wall
Lignin
Lignin exists as a three dimensional macromolecular complex, a high molecular
weight phenolic polymer that is essential for growth, morphology and adaptation (Sarkanen
and Ludwig, 1971a). The lignin polymer is bound covalently to cellulose and hemicelluloses.
Lignin aids plants in many situations, such as protection from pests and pathogens,
mechanical support, increasing root tensile strength, and creating water impermeable vessels
to transport water (EYNCK et al., 2012; Oda and Fukuda, 2012; Zhang et al., 2014). The
lignin polymer is highly resistance to chemical or enzymatic degradation. After death of the
plant, the polysaccharides and proteins are rapidly degraded by microorganisms (Jain et al.,
1979). The remaining lignin becomes the main component of humus in soils and is only
degraded slowly by specialized microbes such as the white rot fungi (Floudas et al., 2012;
Fernandez-Fueyo et al., 2012). The difficulty of degradation of lignin is also an obstacle to
human utilization of plants. High temperature and strong chemicals are needed to extract
lignin from cellulose and hemicelluloses, for example in the pulping process. The
manipulation lignin in plants is important to improve processing efficiency for production of
pulp and paper, liquid biofuels, chemical feedstock, and forage for livestock (Hu et al., 1999;
Chiang, 2002; Ragauskas et al., 2006; Chen and Dixon, 2007).
The monolignol biosynthetic enzymes
Transcription factors regulate secondary cell wall biosynthesis
The enzymes of the monolignol pathway come from multiple genes. Many more
transcription factors that may control their biosynthesis. These transcription factors that
control this complex pathway are interesting to study. In Arabidopsis, multiple transcription
factors regulate the monolignol pathway. NAC (NAM, ATAF1,2, CUC2) is the master
regulator of monolignol pathway and MYB46/58 have a central role (Ko et al., 2009; Zhong
et al., 2010b; Kim et al., 2012). The multiple steps of regulation may control monolignol
biosynthesis by fine tuning. Many transcription factors regulate lignin biosynthesis, such as
Squamosa promoter binding-like (SPL), knotted like homeobox (KNOX), and Cys3His Zn
finger (CCCH) (Fu et al., 2012; Li et al., 2012b; Kim et al., 2014). It is the important to find
the all transcription factors that regulate lignin biosynthesis. The major transcription factors
that regulate secondary cell wall biosynthesis are NAC, MYB, KNOX and LIM.
NAC
poplar TF, PtrSND1-B1 is known to regulate wood formation through alternative splicing
(Lin et al., 2013).
MYB
MYB was the first transcription factor identified in plants 27 years ago; COLORED1
that translate into MYB domain protein for control anthocyanin in maize (Paz-Ares et al.,
1987). After that time MYB transcription factors were characterized in many organisms.
MYB proteins are the conserved transcription factors found in animals and plants (Martin
and Paz-Ares, 1997). The structure of MYB is composed of a helix turn helix structure.
There are three helixes, which are imperfect repeats of their amino acid sequences. Each
helix can serve as a DNA recognition site. The MYB transcription factors can be classified
into four different groups based on the number of adjacent repeats of the protein structure in
the MYBs. In plants, R2R3 MYB is the major group of transcription factors (Martin and
Paz-Ares, 1997). In a comparative study of poplar and Arabidopsis, many poplar MYB genes
were not found in Arabidopsis and might have special functions to support the more complex
poplar genome (Zhao and Bartley, 2014).
downregulates 4CL and C4H. In poplar, MYB156 and MYB221 are highly similar homologs
of AtMYB4. In Arabidopsis MYB52 was reported to control the biosynthesis of the
secondary cell wall (Zhong et al., 2008). PtMYB90 and PtMYB167 are the closely related
homologs of AtMYB52. AtMYB52 can activate the promoters of CesA8, IRX9 and 4CL1.
LIM
LIM is a transcription factor with cysteine-rich domains, which is composed of two
zinc fingers separated by a two amino acid spacer. LIM transcription factors can be classified
into four groups based on phylogenetic analysis and tissue specific expression. LIM was
identified as a transcriptional regulator of some lignin biosynthetic genes (Kawaoka et al.,
2001). In tobacco (
Nicotiana tabacum
), NtLIM1 regulates lignin biosynthesis (Kawaoka et
al., 2001). When an antisense NtLIM1 construct is transformed into Eucalyptus, it reduces
the expression of PAL, C4H, and 4CL (Kawaoka et al., 2006). In Arabidopsis, LIM regulates
the actin cytoskeleton in male flowers (Papuga et al., 2010). In poplar, there are 12 LIM
transcription factors (Arnaud et al., 2012). The PtLIM1 and PtLIM2 transcription factors are
highly expressed in tension wood and normal xylem tissue (Arnaud et al., 2012).
KNOX
genes can be separated into 2 classes. First class is conserved in all plants and has a role in
meristem function and control of leaf shape and hormone homeostasis. For regulating the
compound leaf, KNOX activates cytokinins to increase cell division and repress gibberellin
and lignin biosynthesis and to repress cell differentiation (Hake et al., 2004). The second
class is only found in land plants, which suggests that a duplication occurred around 400 -500
MYA, which resulted in a gain-of-function change that affected leaf and petal shape (Li et
al., 2012b). KNOX can form heterodimers with other homeodomain protein in the TALE
superclass. In a cell fate study, KNOX genes are negative regulators of gibberellin and lignin
biosynthetic pathways. There are 8 KNOTTED ARABIDOPSIS THALIANA (KNAT)
KNOTTED1-like homeodomain (KNOX) genes in Arabidopsis. The KNAT7 is a class II
member of the KNOX family in Arabidopsis. KNAT7 has had many studies to explore its
function, because KNAT7 has strong co - expression with secondary cell wall biosynthesis
enzymes. KNATs are the direct targets of transcription factors SND1, VND6 and MYB46
that regulate the development of secondary cell wall. The
knat7
loss-of-function mutation
shows thicker interfascicular fiber cell walls and increased expression of cellulose, lignin and
xylan biosynthetic genes (Li et al., 2012b). In poplar, we found 14 KNOX genes, of which
KNAT7 (Potri.001G112200.1) is the only xylem specific gene. The suggest function of
KNOX in poplar is for negative feedback to fine tune the control of secondary cell wall
biosynthesis genes.
Conclusion
Metabolic Flux model (PKMF model) shows the complete enzyme kinetics for predicting the
pathway for the conversion of metabolites to each of the major monolignols (Wang et al.,
2014b). The model is built from studies of the synthesis of enzymes in bacteria and yeast,
and can predict lignin content using protein concentration. It is also interesting to find the
relationship between protein concentration of monolignol enzymes and lignin content of
transgenic plants downregulated for monolignol pathway enzymes. This relationship can be
used to identify the key enzymes that exert the most control of the lignin content, for future
work to modify lignin content.
References
Arnaud D., Déjardin A., Leplé J.C., Lesage-Descauses M.C., Boizot N., Villar M.,
Bénédetti H., and Pilate G.
(2012). Expression analysis of LIM gene family in poplar,
toward an updated phylogenetic classification.
5,
102.
Chen F. and Dixon R.A.
(2007). Lignin modification improves fermentable sugar yields for
biofuel production. Nat. Biotechnol.
25,
759-761, doi/10.1038/nbt1316.
Chen H., Song J., Williams C.M., Shuford C.M., Liu J., Wang J.P., Li Q., Shi R., Gokce
E., Ducoste J., Muddiman D.C., Sederoff R.R., and Chiang V.L.
(2013). Monolignol
Pathway 4-Coumaric Acid:Coenzyme A Ligases in Populus. trichocarpa: Novel Specificity,
Metabolic Regulation, and Simulation of Coenzyme A Ligation Fluxes. Plant Physiology
161,
1501-1516, doi/10.1104/pp.112.210971.
Chen H., Song J., Wang J.P., Lin Y., Ducoste J., Shuford C.M., Liu J., Li Q., Shi R.,
Nepomuceno A., Isik F., Muddiman D.C., Williams C., Sederoff R.R., and Chiang V.L.
(2014). Systems Biology of Lignin Biosynthesis in Populus trichocarpa: Heteromeric
4-Coumaric Acid:Coenzyme A Ligase Protein Complex Formation, Regulation, and Numerical
Modeling. The Plant Cell Online
26,
876-893, doi/10.1105/tpc.113.119685.
Chiang V.
(2002). From rags to riches. Nat. Biotechnol.
20,
557-558,
doi/10.1038/nbt0602-557.
Cseke L.J., Cseke S.B., and Podila G.K.
(2007). High efficiency poplar transformation.
Plant Cell Rep.
26,
1529-1538.
De Block M.
(July 1990). Factors Influencing the Tissue Culture and the Agrobacterium
tumefaciens-Mediated Transformation of Hybrid Aspen and Poplar Clones. Plant Physiology
93,
1110-1116, doi/10.1104/pp.93.3.1110.
de Sousa Abreu R., Penalva L.O., Marcotte E.M., and Vogel C.
(2009). Global signatures
of protein and mRNA expression levels.
5,
1512-1526.
EYNCK C., SÉGUIN-SWARTZ G., CLARKE W.E., and PARKIN I.A.P.
(2012).
Monolignol biosynthesis is associated with resistance to Sclerotinia sclerotiorum in Camelina
sativa.
13,
887-899, doi/10.1111/j.1364-3703.2012.00798.x.
Fernandez-Fueyo E., Ruiz-Dueñas F.J., Ferreira P., Floudas D., Hibbett D.S., Canessa
P., Larrondo L.F., James T.Y., Seelenfreund D., Lobos S., Polanco R., Tello M., Honda
Y., Watanabe T., Watanabe T., San R.J., Kubicek C.P., Schmoll M., Gaskell J.,
Hammel K.E., St. John F.J., Vanden Wymelenberg A., Sabat G., Splinter BonDurant
S., Syed K., Yadav J.S., Doddapaneni H., Subramanian V., Lavín J.L., Oguiza J.A.,
Perez G., Pisabarro A.G., Ramirez L., Santoyo F., Master E., Coutinho P.M., Henrissat
B., Lombard V., Magnuson J.K., Kües U., Hori C., Igarashi K., Samejima M., Held
B.W., Barry K.W., LaButti K.M., Lapidus A., Lindquist E.A., Lucas S.M., Riley R.,
Salamov A.A., Hoffmeister D., Schwenk D., Hadar Y., Yarden O., de Vries R.P.,
Kersten P., Martinez A.T., Vicuna R., and Cullen D.
(2012). Comparative genomics of
Ceriporiopsis subvermispora and Phanerochaete chrysosporium provide insight into selective
ligninolysis. Proceedings of the National Academy of Sciences
109,
5458-5463,
doi/10.1073/pnas.1119912109.
Fillatti J.J., Sellmer J., McCown B., Haissig B., and Comai L.
(1987). Agrobacterium
mediated transformation and regeneration of Populus.
206,
192-199,
doi/10.1007/BF00333574.
Floudas D., Binder M., Riley R., Barry K., Blanchette R.A., Henrissat B., Martínez
A.T., Otillar R., Spatafora J.W., Yadav J.S., Aerts A., Benoit I., Boyd A., Carlson A.,
Copeland A., Coutinho P.M., de Vries R.P., Ferreira P., Findley K., Foster B., Gaskell
J., Glotzer D., Górecki P., Heitman J., Hesse C., Hori C., Igarashi K., Jurgens J.A.,
Kallen N., Kersten P., Kohler A., Kües U., Kumar T.K.A., Kuo A., LaButti K.,
Larrondo L.F., Lindquist E., Ling A., Lombard V., Lucas S., Lundell T., Martin R.,
McLaughlin D.J., Morgenstern I., Morin E., Murat C., Nagy L.G., Nolan M., Ohm
R.A., Patyshakuliyeva A., Rokas A., Ruiz-Dueñas F.J., Sabat G., Salamov A., Samejima
M., Schmutz J., Slot J.C., St. John F., Stenlid J., Sun H., Sun S., Syed K., Tsang A.,
Wiebenga A., Young D., Pisabarro A., Eastwood D.C., Martin F., Cullen D., Grigoriev
I.V., and Hibbett D.S.
(2012). The Paleozoic Origin of Enzymatic Lignin Decomposition
Fu C., Sunkar R., Zhou C., Shen H., Zhang J., Matts J., Wolf J., Mann D.G.J., Stewart
C.N., Tang Y., and Wang Z.
(2012). Overexpression of miR156 in switchgrass (Panicum
virgatum L.) results in various morphological alterations and leads to improved biomass
production.
10,
443-452, doi/10.1111/j.1467-7652.2011.00677.x.
Ghaemmaghami S., Huh W., Bower K., Howson R.W., Belle A., Dephoure N., O'Shea
E.K., and Weissman J.S.
(2003). Global analysis of protein expression in yeast. Nature
425,
737-741.
Grant E.H., Fujino T., Beers E.P., and Brunner A.M.
(2010). Characterization of NAC
domain transcription factors implicated in control of vascular cell differentiation in
Arabidopsis and Populus. Planta
232,
337-52, doi/10.1007/s00425-010-1181-2.
Gygi S.P., Rochon Y., Franza B.R., and Aebersold R.
(1999). Correlation between protein
and mRNA abundance in yeast. Mol. Cell. Biol.
19,
1720-1730.
Hake S., Smith H.M., Holtan H., Magnani E., Mele G., and Ramirez J.
(2004). THE
ROLE OF KNOX GENES IN PLANT DEVELOPMENT#. Annu. Rev. Cell Dev. Biol.
20,
125-151.
Hay A. and Tsiantis M.
(2009). A knox family tale. Curr. Opin. Plant Biol.
12,
593-598.
Howe G.T., Goldfarb B., and Strauss S.H.
(1994). Agrobacterium-mediated transformation
Hu W.J., Harding S.A., Lung J., Popko J.L., Ralph J., Stokke D.D., Tsai C.J., and
Chiang V.L.
(1999). Repression of lignin biosynthesis promotes cellulose accumulation and
growth in transgenic trees.
Hu W., Kawaoka A., Tsai C., Lung J., Osakabe K., Ebinuma H., and Chiang V.
(1998).
Compartmentalized expression of two structurally and functionally distinct 4-coumarate :
CoA ligase genes in aspen (Populus tremuloides) RID C-2450-2009. Proc. Natl. Acad. Sci.
U. S. A.
95,
5407-5412, doi/10.1073/pnas.95.9.5407.
Ishihama Y., Oda Y., Tabata T., Sato T., Nagasu T., Rappsilber J., and Mann M.
(2005). Exponentially modified protein abundance index (emPAI) for estimation of absolute
protein amount in proteomics by the number of sequenced peptides per protein. Mol. Cell.
Proteomics
4,
1265-1272, doi/M500061-MCP200 [pii].
Jackson D., Veit B., and Hake S.
(1994). Expression of maize KNOTTED1 related
homeobox genes in the shoot apical meristem predicts patterns of morphogenesis in the
vegetative shoot. Development
120,
405-413.
Jain M., Kapoor K., and Mishra M.
(1979). Cellulase activity, degradation of cellulose and
lignin, and humus formation by thermophilic fungi.
73,
85-89.
Jin H., Cominelli E., Bailey P., Parr A., Mehrtens F., Jones J., Tonelli C., Weisshaar B.,
and Martin C.
(2000). Transcriptional repression by AtMYB4 controls production of
Kawaoka A., Kaothien P., Yoshida K., Endo S., Yamada K., and Ebinuma H.
(2001).
Functional analysis of tobacco LIM protein Ntlim1 involved in lignin biosynthesis.
22,
289-301.
Kawaoka A., Nanto K., Ishii K., and Ebinuma H.
(2006). Reduction of lignin content by
suppression of expression of the LIM domain transcription factor in Eucalyptus
camaldulensis. Silvae Genet.
55,
269-277.
Keane P., Kerr A., and New P.
(1970). Crown gall of stone fruit II. Identification and
nomenclature of Agrobacterium isolates. Aust. J. Biol. Sci.
23,
585-596.
Kim W., Ko J., Kim J., Kim J., Bae H., and Han K.
(2012). MYB46 directly regulates the
gene expression of secondary wall-associated cellulose synthases in Arabidopsis. no-no,
doi/10.1111/j.1365-313X.2012.05124.x.
Kim W., Kim J., Ko J., Kang H., Kim J., and Han K.
(2014). AtC3H14, a plant-specific
tandem CCCH zinc finger protein, binds to its target mRNAs in a sequence-specific manner
and affects cell elongation in Arabidopsis thaliana. n/a-n/a, doi/10.1111/tpj.12667.
Ko J., Kim W., and Han K.
(2009). Ectopic expression of MYB46 identifies transcriptional
regulatory genes involved in secondary wall biosynthesis in Arabidopsis.
60,
649-665,
doi/10.1111/j.1365-313X.2009.03989.x.
Koch G.W., Sillett S.C., Jennings G.M., and Davis S.D.
(2004). The limits to tree height.
Langmead B., Trapnell C., Pop M., and Salzberg S.L.
(2009). Ultrafast and
memory-efficient alignment of short DNA sequences to the human genome RID F-6162-2011.
Genome Biol.
10,
R25-R25, doi/10.1186/gb-2009-10-3-r25.
Li J., Brunner A.M., Shevchenko O., Meilan R., Ma C., Skinner J.S., and Strauss S.H.
(2008). Efficient and stable transgene suppression via RNAi in field-grown poplars.
Transgenic Res.
17,
679-694.
Li E., Bhargava A., Qiang W., Friedmann M.C., Forneris N., Savidge R.A., Johnson
L.A., Mansfield S.D., Ellis B.E., and Douglas C.J.
(2012). The Class II KNOX gene
KNAT7 negatively regulates secondary wall formation in Arabidopsis and is functionally
conserved in Populus. New Phytol.
194,
102-1, doi/10.1111/j.1469-8137.2011.04016.x.
Lin Y., Li W., Sun Y., Kumari S., Wei H., Li Q., Tunlaya-Anukit S., Sederoff R.R., and
Chiang V.L.
(2013). SND1 Transcription Factor–Directed Quantitative Functional
Hierarchical Genetic Regulatory Network in Wood Formation in Populus trichocarpa. Plant
Cell
25,
4324-4341.
Lister R., Gregory B.D., and Ecker J.R.
(2009). Next is now: new technologies for
Ma C., Strauss S., and Meilan R.
(2004). Agrobacterium-mediated transformation of the
genome-sequenced poplar clone, Nisqually-1 (Populus trichocarpa). Plant Mol. Biol. Rep.
22,
311a.
Martin C. and Paz-Ares J.
(1997). MYB transcription factors in plants.
13,
67-73.
Nie L., Wu G., and Zhang W.
(2006a). Correlation between mRNA and protein abundance
in Desulfovibrio vulgaris: a multiple regression to identify sources of variations. Biochem.
Biophys. Res. Commun.
339,
603-610, doi/10.1016/j.bbrc.2005.11.055.
Nie L., Wu G., and Zhang W.
(2006b). Correlation of mRNA Expression and Protein
Abundance Affected by Multiple Sequence Features Related to Translational Efficiency in
Desulfovibrio vulgaris: A Quantitative Analysis. Genetics
174,
2229-2243,
doi/10.1534/genetics.106.065862.
Oda Y. and Fukuda H.
(2012). Secondary cell wall patterning during xylem differentiation.
Curr. Opin. Plant Biol.
15,
38-44, doi/10.1016/j.pbi.2011.10.005.
Ohashi-Ito K., Oda Y., and Fukuda H.
(2010). Arabidopsis VASCULAR-RELATED
NAC-DOMAIN6 Directly Regulates the Genes That Govern Programmed Cell Death and
Secondary Wall Formation during Xylem Differentiation. The Plant Cell Online
22,
3461-3473, doi/10.1105/tpc.110.075036.
Ohtani M., Nishikubo N., Xu B., Yamaguchi M., Mitsuda N., Goue N., Shi F.,
regulation of wood formation in poplar RID F-2183-2011. Plant J.
67,
499-512,
doi/10.1111/j.1365-313X.2011.04614.x.
Papuga J., Hoffmann C., Dieterle M., Moes D., Moreau F., Tholl S., Steinmetz A., and
Thomas C.
(2010). Arabidopsis LIM proteins: a family of actin bundlers with distinct
expression patterns and modes of regulation. Plant Cell
22,
3034-3052,
doi/10.1105/tpc.110.075960 [doi].
Paz-Ares J., Ghosal D., Wienand U., Peterson P.A., and Saedler H.
(1987). The
regulatory c1 locus of Zea mays encodes a protein with homology to myb proto-oncogene
products and with structural similarities to transcriptional activators. EMBO J.
6,
3553-3558.
Ragauskas A.J., Williams C.K., Davison B.H., Britovsek G., Cairney J., Eckert C.A.,
Frederick W.J., Hallett J.P., Leak D.J., and Liotta C.L.
(2006). The Path Forward for
Biofuels and Biomaterials. Science
311,
484-489.
Sarkanen K.V. and Ludwig C.H.
(1971). Lignins: occurrence, formation, structure and
reactions.
Schrimpf S.P., Weiss M., Reiter L., Ahrens C.H., Jovanovic M., Malmström J., Brunner
E., Mohanty S., Lercher M.J., and Hunziker P.E.
(2009). Comparative functional analysis
of the Caenorhabditis elegans and Drosophila melanogaster proteomes.
7,
e1000048.
Shi R., Sun Y., Li Q., Heber S., Sederoff R., and Chiang V.L.
(2010). Towards a Systems
Approach for Lignin Biosynthesis in Populus trichocarpa: Transcript Abundance and
Specificity of the Monolignol Biosynthetic Genes. Plant and Cell Physiology
51,
144-163.
Shi R., Shuford C., Wang J., Sun Y., Yang Z., Chen H., Tunlaya-Anukit S., Li Q., Liu
J., Muddiman D., Sederoff R., and Chiang V.
(2013). Regulation of phenylalanine
ammonia-lyase (PAL) gene family in wood forming tissue of Populus trichocarpa. Planta
238,
487-497, doi/10.1007/s00425-013-1905-1.
Song J., Lu S., Chen Z., Lourenco R., and Chiang V.L.
(2006). Genetic transformation of
Populus trichocarpa genotype Nisqually-1: A functional genomic tool for woody plants.
47,
1582-1589, doi/10.1093/pcp/pcl018.
Su X., Chu Y., Li H., Hou Y., Zhang B., Huang Q., Hu Z., Huang R., and Tian Y.
(2011). Expression of multiple resistance genes enhances tolerance to environmental
stressors in transgenic poplar (Populus euramericana 'Guariento').
6,
e24614,
doi/10.1371/journal.pone.0024614.
Tsai C., Podila G., and Chiang V.
(1994). Agrobacterium-Mediated Transformation of
Quaking Aspen (Populus Tremuloides) and Regeneration of Transgenic Plants Rid
C-2450-2009. Plant Cell Rep.
14,
94-97.
Tuskan G.A., DiFazio S., Jansson S., Bohlmann J., Grigoriev I., Hellsten U., Putnam N.,
Bhalerao R.P., Blaudez D., Boerjan W., Brun A., Brunner A., Busov V., Campbell M.,
Carlson J., Chalot M., Chapman J., Chen G.-., Cooper D., Coutinho P.M., Couturier J.,
Covert S., Cronk Q., Cunningham R., Davis J., Degroeve S., Déjardin A., dePamphilis
C., Detter J., Dirks B., Dubchak I., Duplessis S., Ehlting J., Ellis B., Gendler K.,
Goodstein D., Gribskov M., Grimwood J., Groover A., Gunter L., Hamberger B.,
Heinze B., Helariutta Y., Henrissat B., Holligan D., Holt R., Huang W., Islam-Faridi N.,
Jones S., Jones-Rhoades M., Jorgensen R., Joshi C., Kangasjärvi J., Karlsson J.,
Kelleher C., Kirkpatrick R., Kirst M., Kohler A., Kalluri U., Larimer F., Leebens-Mack
J., Leplé J.-., Locascio P., Lou Y., Lucas S., Martin F., Montanini B., Napoli C., Nelson
D.R., Nelson C., Nieminen K., Nilsson O., Pereda V., Peter G., Philippe R., Pilate G.,
Poliakov A., Razumovskaya J., Richardson P., Rinaldi C., Ritland K., Rouzé P., Ryaboy
D., Schmutz J., Schrader J., Segerman B., Shin H., Siddiqui A., Sterky F., Terry A.,
Tsai C.-., Uberbacher E., Unneberg P., Vahala J., Wall K., Wessler S., Yang G., Yin T.,
Douglas C., Marra M., Sandberg G., Van de Peer Y., and Rokhsar D.
(2006). The
Genome of Black Cottonwood, Populus trichocarpa (Torr. & Gray). Science
313,
1596-1604,
doi/10.1126/science.1128691.
Vogel C., de Sousa Abreu R., Ko D., Le S., Shapiro B.A., Burns S.C., Sandhu D., Boutz
D.R., Marcotte E.M., and Penalva L.O.
(2010). Sequence signatures and mRNA
concentration can explain two
‐
thirds of protein abundance variation in a human cell line.
6,
.
Vollbrecht E., Veit B., Sinha N., and Hake S.
(1991). The developmental gene Knotted-1 is
Wang J.P., Naik P.P., Chen H.C., Shi R., Lin C.Y., Liu J., Shuford C.M., Li Q., Sun
Y.H., Tunlaya-Anukit S., Williams C.M., Muddiman D.C., Ducoste J.J., Sederoff R.R.,
and Chiang V.L.
(2014). Complete proteomic-based enzyme reaction and inhibition kinetics
reveal how monolignol biosynthetic enzyme families affect metabolic flux and lignin in
Populus trichocarpa. Plant Cell
26,
894-914, doi/10.1105/tpc.113.120881 [doi].
Yevtushenko D.P. and Misra S.
(2010). Efficient Agrobacterium-mediated transformation
of commercial hybrid poplar Populus nigra L.× P. maximowiczii A. Henry. Plant Cell Rep.
29,
211-221.
Zhang C., Chen L., and Jiang J.
(2014). Why fine tree roots are stronger than thicker roots:
The role of cellulose and lignin in relation to slope stability. Geomorphology
206,
196-202,
doi/
http://dx.doi.org/10.1016/j.geomorph.2013.09.024
.
Zhao K. and Bartley L.E.
(2014). Comparative genomic analysis of the R2R3 MYB
secondary cell wall regulators of Arabidopsis, poplar, rice, maize, and switchgrass.
14,
135.
Zhong R., Lee C., Zhou J., McCarthy R.L., and Ye Z.H.
(2008). A battery of transcription
factors involved in the regulation of secondary cell wall biosynthesis in Arabidopsis.
20,
2763-2782.
Zhong R., Lee C., and Ye Z.H.
(2010). Global analysis of direct targets of secondary wall
Zhou J., Zhong R., and Ye Z.
(2014). Arabidopsis NAC domain proteins, VND1 to VND5,
are transcriptional regulators of secondary wall biosynthesis in vessels.
9,
e105726.
Zhou J., Lee C., Zhong R., and Ye Z.H.
(2009). MYB58 and MYB63 are transcriptional
Chapter 2 A genetic regulatory network for monolignol
biosynthesis in
Populus trichocarpa
Abstract
Lignin is an important component of wood, which supports the structure of the
secondary cell wall by forming covalent and non-covalent bond to cellulose and
hemicelluloses. The binding to cellulose and hemicellulose makes lignin difficult to extract
and binders the utilization of biomass from the plant. The transcription factors that control
regulation of monolignol biosynthesis are key to understand and manipulate the monolignol
biosynthesis genes are important factors for the potential modification of the secondary cell
walls.
2.1 Introduction
Lignin is a high energy polymer and a potential source of renewable energy. Wood in
angiosperms generally contains, cellulose 42-50%, hemicellulose 25-30%, lignin 20-25% and
extractives 5-8% (Fengel and Wegener, 1984). Cellulose is synthesized by cellulose synthase
complexes located on the plasma membrane of secondary cell walls in wood forming tissues
(Doblin et al., 2002). Lignin and hemicelluloses are also major components of the secondary
cell wall. The components of lignin, cellulose and hemicellulose form the main carbon sink
in trees. In forest ecosystems wood and the residual humus in the soils, account for 20% of
terrestrial carbon storage (Schlesinger and Lichter, 2001). The amount of lignocellulose
offers an enormous, renewable polysaccharide feedstock for materials and biofuels
(Ragauskas et al., 2006). The US Department of Energy has designated trees as target energy
crops, because they do not compete with food crops for agricultural land (McAloon et al.,
2000).
a direct target of a NAC transcription factor controlling secondary cell wall biosynthesis
(Nakano et al., 2010).
LIM is a transcription factor that is characterized by having two LIM domains in its
protein (Arnaud et al., 2007). In tobacco (
Nicotiana tabacum
), NtLIM1 regulates monolignol
biosynthesis (Kawaoka et al., 2001). When antisense NtLIM1 is transformed into Eucalyptus,
it reduces the expression of PAL, C4H, and 4CL (Kawaoka et al., 2006). In Arabidopsis,
LIM regulates the organization and dynamics of the actin cytoskeleton in male flowers
(Papuga et al., 2010). In
P. trichocarpa
, there are 12 LIM transcription factors (Arnaud et al.,
2012). The PtLIM1 and PtLIM2 transcription factors are highly expressed in differentiating
xylem from tension wood and normal differentiating xylem tissue (Arnaud et al., 2012).
IRX9 and 4CL1. AtMYB58 and AtMYB63 are lignin-specific transcription factors regulated
by AtSND1 (Zhou et al., 2009).
The knowledge of monolignol biosynthesis and regulation is still incomplete. Most
experiments have only studied individual gene knockouts and provide a limited picture of
monolignol biosynthesis and regulation. More comprehensive information comes from a
combination of RNA-seq analysis of many transgenic perturbations and the integration of
data to establish a hierarchical genetic regulatory network. An understanding of the genetic
regulatory network for monolignol biosynthesis will provide greater insight into the
formation of the secondary cell wall.
monolignol pathway genes.
2.2 Materials and methods
Transgenic production
Populus trichocarpa
genotype Nisqually-1, was transformed to downregulate genes
for monolignol biosynthesis using Agrobacterium transformation and RNAi or amiRNA
constructs (Schwab et al., 2006; Song et al., 2006b). RNAi downregulates the target genes
with high efficiency and specificity. All transgenic plants were grown in a greenhouse with
nontransformed trees as control groups. The greenhouse experiments consisted of three
replicates of each line. The growth conditions of plants in greenhouse were as described in
(Shi et al., 2013).
Transcriptome analysis
M-values (TMM) method (Robinson and Oshlack, 2010b). The TMM normalization calculates
factors to normalize the different sequencing depth of each library. The counts per million
(CPM) or normalized counts are obtained by dividing the raw read counts by a normalization
factor for each library.
Differential gene expression profiles
Differentially expressed genes were identified using EdgeR: Bioconductor (Robinson
et al., 2010). EdgeR used raw read counts that were determined by BedTools as the input
and testing for differentially expression genes by an exact test based on a negative binomial
distribution (Robinson et al., 2010). The binomial distribution is like the Possion distribution
with a dispersion estimate for each gene. The exact test is like the Fisher test for computing a
P-value by testing if the observed count is different from the expected distribution of the null
hypothesis. Benjamini and Hochberg's algorithm is used to control for the false discovery
rate by correcting for multiple tests (Benjamini and Hochberg, 1995).
Identification of xylem specific transcription factors
Transcription factor genes were identified from TreeTFDB, a database that included
all transcription factors in six tree species based on Hidden Markov Model profiles of
DNA-binding domain families (Mochida et al., 2009).
P. trichocarpa
genome version 2, contains
3,106 transcription factors from a total of 45,033 genes. After screening for xylem specific
genes, we selected those that have annotated functions as a transcription factor.
Gene regulatory network construction by Gaussian Graphical Models
to build a hierarchical network until 6 layers of transcription factors are defined. The
resulting genetic regulatory network was visualized using Cytoscape (Shannon et al., 2003)
2.3 Results
The expression of genes in each tissue
RNA-seq reads from each tissue were mapped onto the
P.trichocarpa
genome version
2 (Tuskan et al., 2006). For all four tissues we mapped 81% of the mRNAs to the
P.trichocarpa
genome. The RNA-seq fragments from xylem, shoot, phloem, and leaf tissue
were mapped at 77%, 74%, 76% and 72% of the transcripts respectively.
Identification of the xylem specific genes
When comparing the expression from 4 tissues, we identified 5,015 xylem specific
genes (Figure 2.1). We selected those genes that have higher expression in xylem compared
to the other three tissues using a significance test for change in abundance (false discovery
rate of 0.05). The xylem specific genes are composed of many functional groups of genes
such as those involved in the biosynthesis of cellulose, hemicellulose, or monolignol
biosynthesis (Table 2.2). All of the following cellulose biosynthesis genes (PtrCesA4,
PtrCesA7, PtrCesA8, PtrCesA17, and PtrCesA18) are xylem specific genes. For
hemicellulose genes only three genes (PtrIRX10L-B1, PtrIRX9-L-A1, and PtrIRX9-L-B1)
are not xylem specific. They have very low expression in xylem and the other three tissues
too. Eighteen of 21 of the monolignol biosynthesis genes are xylem specific, in agreement
with our previous study (Shi et al., 2010).
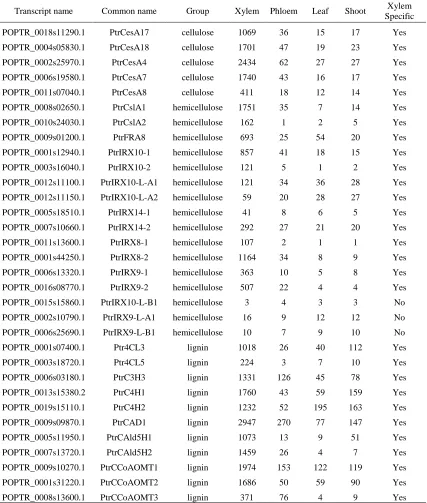
Table 2.1
The expression of cell wall related genes in four types of tissue. Each value is the
average of 3 biological replicates from each tissue. Expression unit is count per million
(cpm).
Transcript name Common name Group Xylem Phloem Leaf Shoot Xylem Specific
POPTR_0018s11290.1 PtrCesA17 cellulose 1069 36 15 17 Yes
POPTR_0004s05830.1 PtrCesA18 cellulose 1701 47 19 23 Yes POPTR_0002s25970.1 PtrCesA4 cellulose 2434 62 27 27 Yes
POPTR_0006s19580.1 PtrCesA7 cellulose 1740 43 16 17 Yes POPTR_0011s07040.1 PtrCesA8 cellulose 411 18 12 14 Yes
POPTR_0008s02650.1 PtrCslA1 hemicellulose 1751 35 7 14 Yes POPTR_0010s24030.1 PtrCslA2 hemicellulose 162 1 2 5 Yes
POPTR_0009s01200.1 PtrFRA8 hemicellulose 693 25 54 20 Yes POPTR_0001s12940.1 PtrIRX10-1 hemicellulose 857 41 18 15 Yes
POPTR_0003s16040.1 PtrIRX10-2 hemicellulose 121 5 1 2 Yes POPTR_0012s11100.1 PtrIRX10-L-A1 hemicellulose 121 34 36 28 Yes
POPTR_0012s11150.1 PtrIRX10-L-A2 hemicellulose 59 20 28 27 Yes POPTR_0005s18510.1 PtrIRX14-1 hemicellulose 41 8 6 5 Yes
POPTR_0007s10660.1 PtrIRX14-2 hemicellulose 292 27 21 20 Yes POPTR_0011s13600.1 PtrIRX8-1 hemicellulose 107 2 1 1 Yes
POPTR_0001s44250.1 PtrIRX8-2 hemicellulose 1164 34 8 9 Yes POPTR_0006s13320.1 PtrIRX9-1 hemicellulose 363 10 5 8 Yes
POPTR_0016s08770.1 PtrIRX9-2 hemicellulose 507 22 4 4 Yes POPTR_0015s15860.1 PtrIRX10-L-B1 hemicellulose 3 4 3 3 No
POPTR_0002s10790.1 PtrIRX9-L-A1 hemicellulose 16 9 12 12 No POPTR_0006s25690.1 PtrIRX9-L-B1 hemicellulose 10 7 9 10 No
POPTR_0001s07400.1 Ptr4CL3 lignin 1018 26 40 112 Yes POPTR_0003s18720.1 Ptr4CL5 lignin 224 3 7 10 Yes
POPTR_0006s03180.1 PtrC3H3 lignin 1331 126 45 78 Yes POPTR_0013s15380.2 PtrC4H1 lignin 1760 43 59 159 Yes
POPTR_0019s15110.1 PtrC4H2 lignin 1232 52 195 163 Yes POPTR_0009s09870.1 PtrCAD1 lignin 2947 270 77 147 Yes
POPTR_0005s11950.1 PtrCAld5H1 lignin 1073 13 9 51 Yes POPTR_0007s13720.1 PtrCAld5H2 lignin 1459 26 4 7 Yes
POPTR_0009s10270.1 PtrCCoAOMT1 lignin 1974 153 122 119 Yes POPTR_0001s31220.1 PtrCCoAOMT2 lignin 1686 50 59 90 Yes
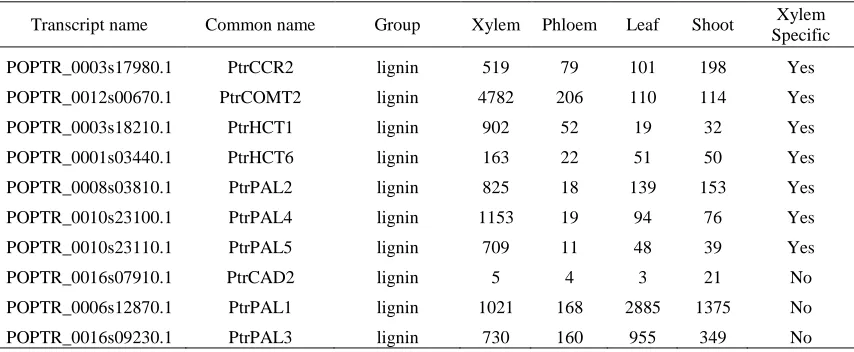
Table 2.1 Continued
Transcript name Common name Group Xylem Phloem Leaf Shoot Xylem Specific
POPTR_0003s17980.1 PtrCCR2 lignin 519 79 101 198 Yes POPTR_0012s00670.1 PtrCOMT2 lignin 4782 206 110 114 Yes
POPTR_0003s18210.1 PtrHCT1 lignin 902 52 19 32 Yes POPTR_0001s03440.1 PtrHCT6 lignin 163 22 51 50 Yes
POPTR_0008s03810.1 PtrPAL2 lignin 825 18 139 153 Yes POPTR_0010s23100.1 PtrPAL4 lignin 1153 19 94 76 Yes
POPTR_0010s23110.1 PtrPAL5 lignin 709 11 48 39 Yes POPTR_0016s07910.1 PtrCAD2 lignin 5 4 3 21 No
POPTR_0006s12870.1 PtrPAL1 lignin 1021 168 2885 1375 No POPTR_0016s09230.1 PtrPAL3 lignin 730 160 955 349 No
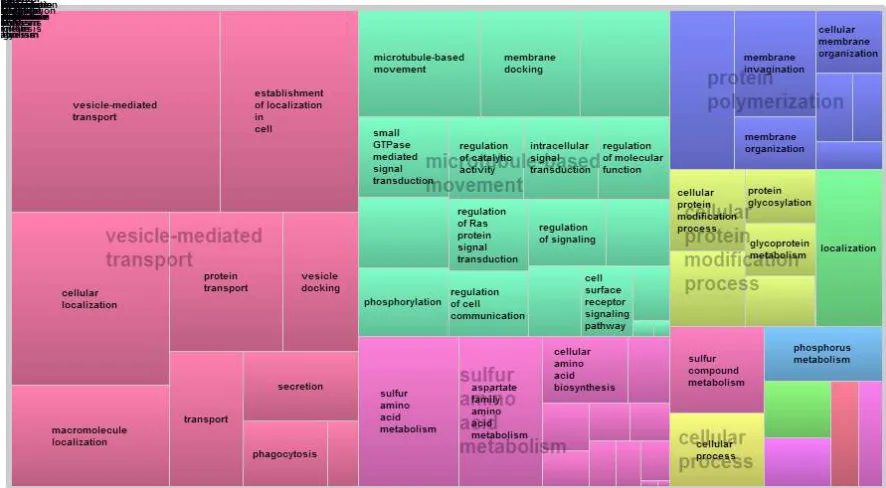
Figure 2.2
The summary of GO analysis using xylem specific genes as input and summarize
by REVIGO. The xylem specific genes have biological process relate to vesicle mediated
transport, microtubule based movement, sulfur amino acid metabolism, protein
polymerization, and cellular protein modification process
NACs previously identified as key regulators of secondary cell wall biosynthesis. NACs also
control MYBs as regulators of monolignol biosynthesis (Goicoechea et al., 2005; Ko et al.,
2007; Shen et al., 2009; Ohashi-Ito et al., 2010; Lin et al., 2013; Ng et al., 2013;
Craven-Bartle et al., 2013; Tian et al., 2013). The TF with the highest expression is WOX13, which
controls cell type differentiation (Haecker et al., 2004; Zhang et al., 2010; Hedman et al.,
2013). Another group of interest is the PHD TFs that function to control histone modification
(Sung et al., 2006). The xylem specific transcription factors were pooled and used to a build
a hierarchical genetic regulatory network that controls monolignol biosynthesis.
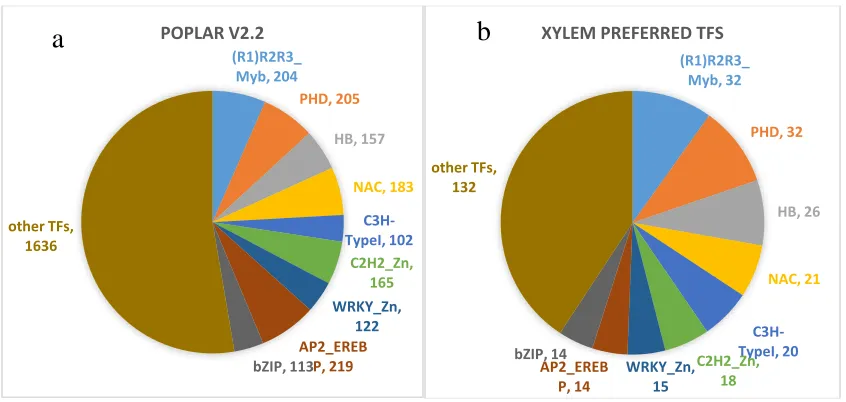
Figure 2.3
The top 10 families of transcription factors in the poplar genome (a) comparisons
to xylem specific transcription factors (b).
(R1)R2R3_ Myb, 204
PHD, 205
HB, 157
NAC, 183
C3H-TypeI, 102
C2H2_Zn, 165 WRKY_Zn,
122
AP2_EREB P, 219
bZIP, 113 other TFs,
1636
POPLAR V2.2
(R1)R2R3_ Myb, 32
PHD, 32
HB, 26
NAC, 21
C3H-TypeI, 20 C2H2_Zn,
18 WRKY_Zn,
15
AP2_EREB P, 14
bZIP, 14 other TFs,
132
XYLEM PREFERRED TFS
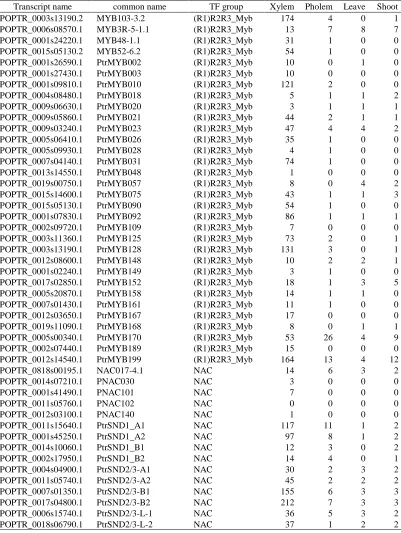
Table 2.2
The xylem specific transcription factors and their expression in four tissues. The
unit of expression is count per million (cpm)
Transcript name common name TF group Xylem Pholem Leave Shoot POPTR_0003s13190.2 MYB103-3.2 (R1)R2R3_Myb 174 4 0 1 POPTR_0006s08570.1 MYB3R-5-1.1 (R1)R2R3_Myb 13 7 8 7 POPTR_0001s24220.1 MYB48-1.1 (R1)R2R3_Myb 31 1 0 0 POPTR_0015s05130.2 MYB52-6.2 (R1)R2R3_Myb 54 1 0 0 POPTR_0001s26590.1 PtrMYB002 (R1)R2R3_Myb 10 0 1 0 POPTR_0001s27430.1 PtrMYB003 (R1)R2R3_Myb 10 0 0 0 POPTR_0001s09810.1 PtrMYB010 (R1)R2R3_Myb 121 2 0 0 POPTR_0004s08480.1 PtrMYB018 (R1)R2R3_Myb 5 1 1 2 POPTR_0009s06630.1 PtrMYB020 (R1)R2R3_Myb 3 1 1 1 POPTR_0009s05860.1 PtrMYB021 (R1)R2R3_Myb 44 2 1 1 POPTR_0009s03240.1 PtrMYB023 (R1)R2R3_Myb 47 4 4 2 POPTR_0005s06410.1 PtrMYB026 (R1)R2R3_Myb 35 1 0 0 POPTR_0005s09930.1 PtrMYB028 (R1)R2R3_Myb 4 1 0 0 POPTR_0007s04140.1 PtrMYB031 (R1)R2R3_Myb 74 1 0 0 POPTR_0013s14550.1 PtrMYB048 (R1)R2R3_Myb 1 0 0 0 POPTR_0019s00750.1 PtrMYB057 (R1)R2R3_Myb 8 0 4 2 POPTR_0015s14600.1 PtrMYB075 (R1)R2R3_Myb 43 1 1 3 POPTR_0015s05130.1 PtrMYB090 (R1)R2R3_Myb 54 1 0 0 POPTR_0001s07830.1 PtrMYB092 (R1)R2R3_Myb 86 1 1 1 POPTR_0002s09720.1 PtrMYB109 (R1)R2R3_Myb 7 0 0 0 POPTR_0003s11360.1 PtrMYB125 (R1)R2R3_Myb 73 2 0 1 POPTR_0003s13190.1 PtrMYB128 (R1)R2R3_Myb 131 3 0 1 POPTR_0012s08600.1 PtrMYB148 (R1)R2R3_Myb 10 2 2 1 POPTR_0001s02240.1 PtrMYB149 (R1)R2R3_Myb 3 1 0 0 POPTR_0017s02850.1 PtrMYB152 (R1)R2R3_Myb 18 1 3 5 POPTR_0005s20870.1 PtrMYB158 (R1)R2R3_Myb 14 1 1 0 POPTR_0007s01430.1 PtrMYB161 (R1)R2R3_Myb 11 1 0 0 POPTR_0012s03650.1 PtrMYB167 (R1)R2R3_Myb 17 0 0 0 POPTR_0019s11090.1 PtrMYB168 (R1)R2R3_Myb 8 0 1 1 POPTR_0005s00340.1 PtrMYB170 (R1)R2R3_Myb 53 26 4 9 POPTR_0002s07440.1 PtrMYB189 (R1)R2R3_Myb 15 0 0 0 POPTR_0012s14540.1 PtrMYB199 (R1)R2R3_Myb 164 13 4 12
POPTR_0818s00195.1 NAC017-4.1 NAC 14 6 3 2
POPTR_0014s07210.1 PNAC030 NAC 3 0 0 0
POPTR_0001s41490.1 PNAC101 NAC 7 0 0 0
POPTR_0011s05760.1 PNAC102 NAC 0 0 0 0
POPTR_0012s03100.1 PNAC140 NAC 1 0 0 0
POPTR_0011s15640.1 PtrSND1_A1 NAC 117 11 1 2
POPTR_0001s45250.1 PtrSND1_A2 NAC 97 8 1 2
POPTR_0014s10060.1 PtrSND1_B1 NAC 12 3 0 2
POPTR_0002s17950.1 PtrSND1_B2 NAC 14 4 0 1
POPTR_0004s04900.1 PtrSND2/3-A1 NAC 30 2 3 2
POPTR_0011s05740.1 PtrSND2/3-A2 NAC 45 2 2 2
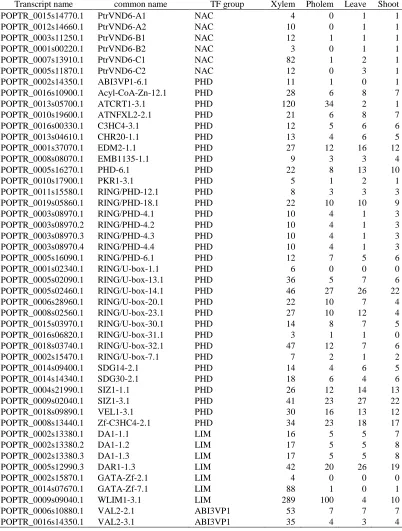
Table 2.2 Continued
Transcript name common name TF group Xylem Pholem Leave Shoot
POPTR_0015s14770.1 PtrVND6-A1 NAC 4 0 1 1
POPTR_0012s14660.1 PtrVND6-A2 NAC 10 0 1 1
POPTR_0003s11250.1 PtrVND6-B1 NAC 12 1 1 1
POPTR_0001s00220.1 PtrVND6-B2 NAC 3 0 1 1
POPTR_0007s13910.1 PtrVND6-C1 NAC 82 1 2 1
POPTR_0005s11870.1 PtrVND6-C2 NAC 12 0 3 1
POPTR_0002s14350.1 ABI3VP1-6.1 PHD 11 1 0 1
POPTR_0016s10900.1 Acyl-CoA-Zn-12.1 PHD 28 6 8 7
POPTR_0013s05700.1 ATCRT1-3.1 PHD 120 34 2 1
POPTR_0010s19600.1 ATNFXL2-2.1 PHD 21 6 8 7
POPTR_0016s00330.1 C3HC4-3.1 PHD 12 5 6 6
POPTR_0013s04610.1 CHR20-1.1 PHD 13 4 6 5
POPTR_0001s37070.1 EDM2-1.1 PHD 27 12 16 12
POPTR_0008s08070.1 EMB1135-1.1 PHD 9 3 3 4
POPTR_0005s16270.1 PHD-6.1 PHD 22 8 13 10
POPTR_0010s17900.1 PKR1-3.1 PHD 5 1 2 1
POPTR_0011s15580.1 RING/PHD-12.1 PHD 8 3 3 3
POPTR_0019s05860.1 RING/PHD-18.1 PHD 22 10 10 9
POPTR_0003s08970.1 RING/PHD-4.1 PHD 10 4 1 3
POPTR_0003s08970.2 RING/PHD-4.2 PHD 10 4 1 3
POPTR_0003s08970.3 RING/PHD-4.3 PHD 10 4 1 3
POPTR_0003s08970.4 RING/PHD-4.4 PHD 10 4 1 3
POPTR_0005s16090.1 RING/PHD-6.1 PHD 12 7 5 6
POPTR_0001s02340.1 RING/U-box-1.1 PHD 6 0 0 0 POPTR_0005s02090.1 RING/U-box-13.1 PHD 36 5 7 6 POPTR_0005s02460.1 RING/U-box-14.1 PHD 46 27 26 22 POPTR_0006s28960.1 RING/U-box-20.1 PHD 22 10 7 4 POPTR_0008s02560.1 RING/U-box-23.1 PHD 27 10 12 4 POPTR_0015s03970.1 RING/U-box-30.1 PHD 14 8 7 5 POPTR_0016s06820.1 RING/U-box-31.1 PHD 3 1 1 0 POPTR_0018s03740.1 RING/U-box-32.1 PHD 47 12 7 6 POPTR_0002s15470.1 RING/U-box-7.1 PHD 7 2 1 2
POPTR_0014s09400.1 SDG14-2.1 PHD 14 4 6 5
POPTR_0014s14340.1 SDG30-2.1 PHD 18 6 4 6
POPTR_0004s21990.1 SIZ1-1.1 PHD 26 12 14 13
POPTR_0009s02040.1 SIZ1-3.1 PHD 41 23 27 22
POPTR_0018s09890.1 VEL1-3.1 PHD 30 16 13 12
POPTR_0008s13440.1 Zf-C3HC4-2.1 PHD 34 23 18 17
POPTR_0002s13380.1 DA1-1.1 LIM 16 5 5 7
POPTR_0002s13380.2 DA1-1.2 LIM 17 5 5 8
POPTR_0002s13380.3 DA1-1.3 LIM 17 5 5 8
POPTR_0005s12990.3 DAR1-1.3 LIM 42 20 26 19
POPTR_0002s15870.1 GATA-Zf-2.1 LIM 4 0 0 0
POPTR_0014s07670.1 GATA-Zf-7.1 LIM 88 1 0 1
POPTR_0009s09040.1 WLIM1-3.1 LIM 289 100 4 10
POPTR_0006s10880.1 VAL2-2.1 ABI3VP1 53 7 7 7
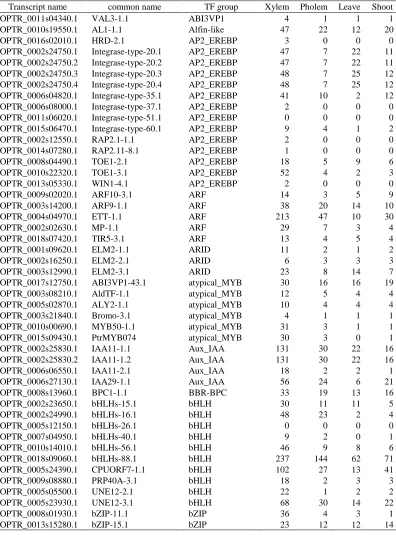
Table 2.2 Continued
Transcript name common name TF group Xylem Pholem Leave Shoot
POPTR_0011s04340.1 VAL3-1.1 ABI3VP1 4 1 1 1
POPTR_0010s19550.1 AL1-1.1 Alfin-like 47 22 12 20
POPTR_0016s02010.1 HRD-2.1 AP2_EREBP 3 0 0 0
POPTR_0002s24750.1 Integrase-type-20.1 AP2_EREBP 47 7 22 11 POPTR_0002s24750.2 Integrase-type-20.2 AP2_EREBP 47 7 22 11 POPTR_0002s24750.3 Integrase-type-20.3 AP2_EREBP 48 7 25 12 POPTR_0002s24750.4 Integrase-type-20.4 AP2_EREBP 48 7 25 12 POPTR_0006s04820.1 Integrase-type-35.1 AP2_EREBP 41 10 2 12 POPTR_0006s08000.1 Integrase-type-37.1 AP2_EREBP 2 0 0 0 POPTR_0011s06020.1 Integrase-type-51.1 AP2_EREBP 0 0 0 0 POPTR_0015s06470.1 Integrase-type-60.1 AP2_EREBP 9 4 1 2 POPTR_0002s12550.1 RAP2.1-1.1 AP2_EREBP 2 0 0 0 POPTR_0014s07280.1 RAP2.11-8.1 AP2_EREBP 1 0 0 0 POPTR_0008s04490.1 TOE1-2.1 AP2_EREBP 18 5 9 6 POPTR_0010s22320.1 TOE1-3.1 AP2_EREBP 52 4 2 3 POPTR_0013s05330.1 WIN1-4.1 AP2_EREBP 2 0 0 0
POPTR_0009s02020.1 ARF10-3.1 ARF 14 3 5 9
POPTR_0003s14200.1 ARF9-1.1 ARF 38 20 14 10
POPTR_0004s04970.1 ETT-1.1 ARF 213 47 10 30
POPTR_0002s02630.1 MP-1.1 ARF 29 7 3 4
POPTR_0018s07420.1 TIR5-3.1 ARF 13 4 5 4
POPTR_0001s09620.1 ELM2-1.1 ARID 11 2 1 2
POPTR_0002s16250.1 ELM2-2.1 ARID 6 3 3 3
POPTR_0003s12990.1 ELM2-3.1 ARID 23 8 14 7
POPTR_0017s12750.1 ABI3VP1-43.1 atypical_MYB 30 16 16 19 POPTR_0003s08210.1 AldTF-1.1 atypical_MYB 12 5 4 4 POPTR_0005s02870.1 ALY2-1.1 atypical_MYB 10 4 4 4 POPTR_0003s21840.1 Bromo-3.1 atypical_MYB 4 1 1 1 POPTR_0010s00690.1 MYB50-1.1 atypical_MYB 31 3 1 1 POPTR_0015s09430.1 PtrMYB074 atypical_MYB 30 3 0 1 POPTR_0002s25830.1 IAA11-1.1 Aux_IAA 131 30 22 16 POPTR_0002s25830.2 IAA11-1.2 Aux_IAA 131 30 22 16 POPTR_0006s06550.1 IAA11-2.1 Aux_IAA 18 2 2 1 POPTR_0006s27130.1 IAA29-1.1 Aux_IAA 56 24 6 21 POPTR_0008s13960.1 BPC1-1.1 BBR-BPC 33 19 13 16 POPTR_0002s23650.1 bHLHs-15.1 bHLH 30 11 11 5
POPTR_0002s24990.1 bHLHs-16.1 bHLH 48 23 2 4
POPTR_0005s12150.1 bHLHs-26.1 bHLH 0 0 0 0
POPTR_0007s04950.1 bHLHs-40.1 bHLH 9 2 0 1
POPTR_0010s14010.1 bHLHs-56.1 bHLH 46 9 8 6
POPTR_0018s09060.1 bHLHs-88.1 bHLH 237 144 62 71 POPTR_0005s24390.1 CPUORF7-1.1 bHLH 102 27 13 41
POPTR_0009s08880.1 PRP40A-3.1 bHLH 18 2 3 3
POPTR_0005s05500.1 UNE12-2.1 bHLH 22 1 2 2
POPTR_0005s23930.1 UNE12-3.1 bHLH 68 30 14 22
POPTR_0008s01930.1 bZIP-11.1 bZIP 36 4 3 1
Table 2.2 Continued
Transcript name common name TF group Xylem Pholem Leave Shoot
POPTR_0019s15040.1 bZIP-17.1 bZIP 23 11 9 11
POPTR_0001s01840.1 bZIP16-1.1 bZIP 17 11 11 10 POPTR_0003s09630.1 bZIP16-2.1 bZIP 25 13 14 13
POPTR_0003s20320.1 bZIP23-4.1 bZIP 27 8 11 5
POPTR_0017s14490.1 bZIP42-4.1 bZIP 4 0 0 1
POPTR_0002s04660.1 BZO2H1-1.1 bZIP 18 10 7 9
POPTR_0009s02360.1 DPBF3-4.1 bZIP 11 5 4 3
POPTR_0005s26480.1 FD-1-2.1 bZIP 16 5 0 5
POPTR_0014s08990.1 GBF3-2.1 bZIP 19 8 9 12
POPTR_0014s08990.2 GBF3-2.2 bZIP 19 8 9 12
POPTR_0004s21360.1 TGA9-1.1 bZIP 49 16 2 1
POPTR_0009s16590.1 TGA9-2.1 bZIP 12 3 0 1
POPTR_0007s02690.1 BtZf-6.1 C2C2_Zn-like 17 0 6 1 POPTR_0009s12730.1 BtZf-8.1 C2C2_Zn-like 36 20 22 25 POPTR_0002s22640.1 COL9-1.1 C2C2_Zn-like 23 10 3 4 POPTR_0004s16810.1 STH2-2.1 C2C2_Zn-like 5 0 0 0 POPTR_0004s22140.1 GATA11-1.1 C2C2_Zn-GATA 40 11 18 12 POPTR_0006s25410.1 GATA12-2.1 C2C2_Zn-GATA 12 3 3 3 POPTR_0010s23010.1 GATA9-3.1 C2C2_Zn-GATA 76 13 40 26 POPTR_0010s23010.2 GATA9-3.2 C2C2_Zn-GATA 76 14 41 26 POPTR_0014s06150.1 C2H2LZf-17.1 C2H2_Zn 18 3 1 1 POPTR_0001s15790.1 C2H2LZf-2.1 C2H2_Zn 11 4 2 4 POPTR_0017s12740.1 C2H2LZf-20.1 C2H2_Zn 8 0 0 0 POPTR_0003s07490.1 C2H2LZf-6.1 C2H2_Zn 15 7 7 8 POPTR_0004s11730.1 C2H2LZf-7.1 C2H2_Zn 4 0 0 0 POPTR_0010s21650.1 C2H2Zf-23.1 C2H2_Zn 12 0 0 0 POPTR_0014s06160.1 C2H2Zf-26.1 C2H2_Zn 12 0 0 0 POPTR_0002s14450.1 C2H2Zf-5.1 C2H2_Zn 6 0 0 0 POPTR_0018s05280.1 IDD7-1.1 C2H2_Zn 29 16 15 13
POPTR_0010s15060.1 KNU-2.1 C2H2_Zn 0 0 0 0
POPTR_0014s14320.1 PPR-1.1 C2H2_Zn 65 12 34 35 POPTR_0001s16080.1 SUF4-1.1 C2H2_Zn 40 13 17 18 POPTR_0001s16080.2 SUF4-1.2 C2H2_Zn 40 13 17 18
POPTR_0003s07180.1 SUF4-2.1 C2H2_Zn 21 5 6 7
POPTR_0003s07180.2 SUF4-2.2 C2H2_Zn 21 5 6 7
POPTR_0019s11040.1 TFIIIA-2.1 C2H2_Zn 7 4 3 4
POPTR_0002s04260.1 ZFP11-1.1 C2H2_Zn 3 0 0 0
POPTR_0005s24280.1 ZFP11-2.1 C2H2_Zn 1 0 0 0